research use only
Copanlisib (BAY 80-6946) PI3K inhibitor
Cat.No.S2802
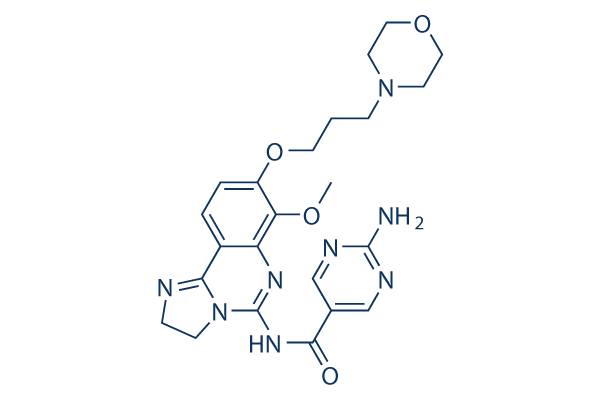
Chemical Structure
Molecular Weight: 480.52
Quality Control
Products Often Used Together with Copanlisib (BAY 80-6946)
| Related Targets | Akt mTOR GSK-3 ATM/ATR DNA-PK AMPK PDPK1 PTEN PP2A PDK |
|---|---|
| Other PI3K Inhibitors | GDC-0077 (Inavolisib) SAR405 Quercetin (Sophoretin) LY294002 XL147 analogue Tersolisib (STX-478) Buparlisib (BKM120) 740 Y-P (PDGFR 740Y-P) GO-203 TFA Eganelisib (IPI-549) |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| Huh7 | Growth inhibiton assay | 100 nM | Copanlisib dose-dependently inhibited cell growth in vitro. IC50=47.9 nM. | 30962952 | ||
| HepG2 | Growth inhibiton assay | 100 nM | Copanlisib dose-dependently inhibited cell growth in vitro. IC50=31.6 nM. | 30962952 | ||
| Hep3B | Growth inhibiton assay | IC50=72.4 nM | 30962952 | |||
| PLCPRF5 | Growth inhibiton assay | IC50=283 nM | 30962952 | |||
| Chang | Growth inhibiton assay | IC50=442 nM | 30962952 | |||
| JVM-3 | Function assay | 48 h | inhibits metabolic activity with an IC50 of 2 μM in the XTT assay | 25912635 | ||
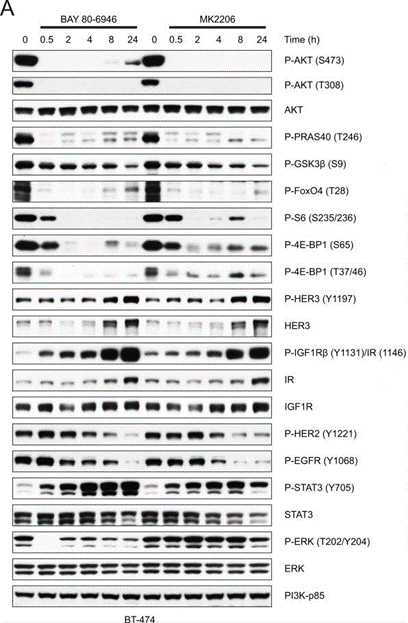
| BT-474 | Function assay | 50 nM | 0.5, 2, 4, 8, 24 h | rapidly inhibits the phosphorylation of AKT (S473, T308) as well as its direct substrates PRAS40 (T246) and GSK3β (S9), and inhibition was sustained for up to 24 hours | 24436048 | |
| SK-BR-3 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| UACC-893 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| HCC-1954 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| MDA-MB-453 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| MDA-MB-361 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| BT-20 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| MCF-7 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| T-47D | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| HCC1806 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| NCI-H292 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| NCI-H1650 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| CCRF-SB | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| U937 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| SU-DHL-4 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| SU-DHL-5 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| HCT116 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| A549 cells | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| SK-MEL-30 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| SK-MEL-2 cells | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| NCI-H1703 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| NCI-H661 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| PC9 | Function assay | 0, 1, 2, 4 h | downregulation of P-AKT | 24436048 | ||
| TC32 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for TC32 cells | 29435139 | |||
| A673 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for A673 cells | 29435139 | |||
| Saos-2 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for Saos-2 cells | 29435139 | |||
| RD | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for RD cells | 29435139 | |||
| MG 63 (6-TG R) | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for MG 63 (6-TG R) cells | 29435139 | |||
| OHS-50 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for OHS-50 cells | 29435139 | |||
| LAN-5 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for LAN-5 cells | 29435139 | |||
| NB-EBc1 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for NB-EBc1 cells | 29435139 | |||
| SK-N-SH | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for SK-N-SH cells | 29435139 | |||
| Rh41 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for Rh41 cells | 29435139 | |||
| A673 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for A673 cells) | 29435139 | |||
| BT-37 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for BT-37 cells | 29435139 | |||
| MG 63 (6-TG R) | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for MG 63 (6-TG R) cells | 29435139 | |||
| Rh30 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for Rh30 cells | 29435139 | |||
| OHS-50 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for OHS-50 cells | 29435139 | |||
| SJ-GBM2 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SJ-GBM2 cells | 29435139 | |||
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SK-N-MC cells | 29435139 | |||
| NB-EBc1 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for NB-EBc1 cells | 29435139 | |||
| LAN-5 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for LAN-5 cells | 29435139 | |||
| Rh18 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for Rh18 cells | 29435139 | |||
| NB1643 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for NB1643 cells | 29435139 | |||
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for SK-N-MC cells | 29435139 | |||
| SJ-GBM2 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for SJ-GBM2 cells | 29435139 | |||
| TC32 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for TC32 cells | 29435139 | |||
| Rh18 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for Rh18 cells | 29435139 | |||
| Saos-2 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for Saos-2 cells | 29435139 | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
5%TFA : 8 mg/mL
DMSO
: Insoluble
Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 480.52 | Formula | C23H28N8O4 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 1032568-63-0 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | BAY 80-6946 | Smiles | COC1=C(C=CC2=C3NCCN3C(=NC(=O)C4=CN=C(N=C4)N)N=C21)OCCCN5CCOCC5 | ||
Mechanism of Action
| Targets/IC50/Ki |
PI3Kα
(Cell-free assay) 0.5 nM
PI3Kδ
(Cell-free assay) 0.7 nM
PI3Kβ
(Cell-free assay) 3.7 nM
PI3Kγ
(Cell-free assay) 6.4 nM
|
|---|---|
| In vitro |
In both KPL4 cells and LPA-stimulated PC3 cells, BAY 80-6946 reduces pAKT levels. In a subset of human cancer cell lines with PIK3CA mutations and/or overexpression of HER2, BAY 80-6946 shows antiproliferative activity and induces apoptosis. [1] The combination of HER2-targeted therapies and BAY 80-6946 inhibits growth more effectively than either therapy used alone, and can restore sensitivity to trastuzumab and lapatinib in cells. [2] |
| Kinase Assay |
Biochemical lipid kinase assays
|
|
The effect of BAY 80-6946 on PI3Kα, PI3Kβ, and PI3Kγ activity is measured by the inhibition of 33P incorporation into phosphatidylinositol (PI) in 384-well MaxiSorp® plates coated with 2 µg/well of PI and phosphatidylserine (PS) (1:1 molar ratio). In each PI3K isoform assay, 9 µL of reaction buffer (50 mM MOPSO, pH 7.0, 100 mM NaCl, 4 mM MgCl2, 0.1% BSA) containing 7.5 ng of His-tagged N-terminal truncated p110α or p110β protein, or 25 ng of purified human p110γ protein, is used. The reaction is started by adding 5 µL of a 40-µM ATP solution containing 20 µCi/mL [33>/sup>P]-ATP. After 2 hours incubation at room temperature, the reaction is terminated by addition of 5 µL of a 25-mM EDTA solution. The plates are washed and Ultima Gold™ scintillation cocktail (25 µL) is then added. The radioactivity incorporated into the immobilized PI substrate is determined with a BetaPlate Liquid Scintillation Counter.
|
|
| In vivo |
In rat KPL4 or HCT116 tumor xenograft model, BAY 80-6946 (6 mg/kg, i.v.) induces 100% complete tumor regression. In nude mice with Lu7860 erlotinib-resistant, patient-derived NSCLC and MAXF1398 patient-derived luminal breast tumor models, BAY 80-6946 (14 mg/kg, i.v.) also causes tumor growth inhibition. [1] |
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | p-FoxO4(T28) / p-S6(S235/236) / p-4E-BP1(S65) / p-4E-BP1(T37/46) / p-HER3(Y1197) / HER3 / p-IGF1Rβ/ IGF1R / p-HER2 / p-EGFR / p-STAT3 / p-ERK p-AKT / AKT / p-PRAS40(T246) / p-GSK3β(S9) / cleaved caspase-3 / cleaved caspase-7 / PI3K-p85 |
|
24436048 |
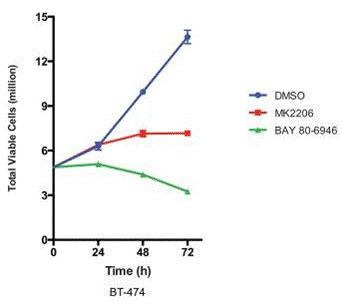
| Growth inhibition assay | Cell viability |
|
24436048 |
Clinical Trial Information
(data from https://clinicaltrials.gov, updated on 2024-05-22)
| NCT Number | Recruitment | Conditions | Sponsor/Collaborators | Start Date | Phases |
|---|---|---|---|---|---|
| NCT05082025 | Active not recruiting | Endometrial Cancer|Ovarian Cancer |
M.D. Anderson Cancer Center|Bayer |
September 27 2022 | Phase 2 |
| NCT05217914 | Active not recruiting | Relapsed or Refractory Indolent Non-Hodgkin Lymphoma |
Bayer |
July 1 2022 | -- |
| NCT04939272 | Suspended | Recurrent Mantle Cell Lymphoma|Refractory Mantle Cell Lymphoma |
City of Hope Medical Center|National Cancer Institute (NCI) |
June 29 2022 | Phase 1|Phase 2 |
| NCT04572763 | Active not recruiting | Diffuse Large B Cell Lymphoma|Relapsed Diffuse Large B-Cell Lymphoma|Refractory Diffuse Large B-Cell Lymphoma |
Dana-Farber Cancer Institute|AbbVie|Bayer |
September 8 2021 | Phase 1|Phase 2 |
| NCT04803123 | Terminated | Leukemia Acute Lymphocytic |
Dorothy Sipkins MD PhD|Bayer|Duke University |
June 21 2021 | Early Phase 1 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































