research use only
Pilaralisib (XL147) PI3K inhibitor
Cat.No.S7645
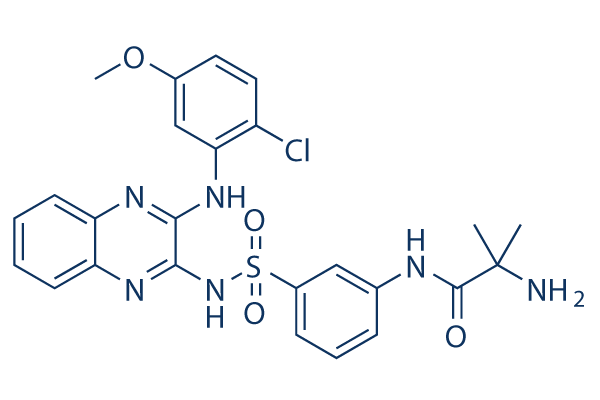
Chemical Structure
Molecular Weight: 541.02
Quality Control
| Related Targets | Akt mTOR GSK-3 ATM/ATR DNA-PK AMPK PDPK1 PTEN PP2A PDK |
|---|---|
| Other PI3K Inhibitors | GDC-0077 (Inavolisib) SAR405 Quercetin (Sophoretin) LY294002 XL147 analogue Tersolisib (STX-478) Buparlisib (BKM120) 740 Y-P (PDGFR 740Y-P) GO-203 TFA Eganelisib (IPI-549) |
Solubility
|
In vitro |
DMSO
: 100 mg/mL
(184.83 mM)
Water : Insoluble Ethanol : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 541.02 | Formula | C25H25ClN6O4S |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 934526-89-3 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N/A | Smiles | CC(C)(C(=O)NC1=CC(=CC=C1)S(=O)(=O)NC2=NC3=CC=CC=C3N=C2NC4=C(C=CC(=C4)OC)Cl)N | ||
Mechanism of Action
| Targets/IC50/Ki |
PI3Kγ
23 nM
PI3Kβ
36 nM
PI3Kδ
36 nM
PI3Kα
39 nM
|
|---|---|
| In vitro |
In Pediatric Preclinical Testing Program (PPTP) cell lines, Pilaralisib (XL147) exhibits cytotoxic activity, with a median relative IC50 value of 10.9 mM (range 2.7 mM to 24.5 mM).
|
| Kinase Assay |
In vitro kinase inhibition assays
|
|
Kinase activity for PI3K isoforms is measured as the percentage of ATP consumed following the kinase reaction using luciferase–luciferin-coupled chemiluminescence, with ATP concentrations approximately equal to the Km for each respective kinase. Kinase reactions are initiated by combining test compounds, ATP and kinase in a 20 μL volume. PI3Kα, PI3Kβ, PI3Kγ, and PI3Kδ final enzyme concentrations are 0.5, 8, 20, and 2 nM, respectively. Of note, 0.5 μL dimethyl sulfoxide (DMSO) containing varying concentrations of this compound is mixed with 10 μL enzyme solution (2×concentration). Kinase reactions are initiated by the addition of 10 μL of liver phosphatidylinositol and ATP solution (2×concentration). Assay concentrations for VPS34, ATP, and phosphatidylinositol are 40 nM, 1 μM, and 5 μM, respectively.
|
|
| In vivo |
In BALB/c nu/nu mice, Pilaralisib (XL147) (100 mg/kg, p.o.) induces tumor growth inhibition for solid glioma xenografts and is well tolerated, with only 0.7% toxicity rate in the treated groups, similar to that observed for control animals. In athymic female mouse, it (100 mg/kg, p.o.) significantly delays tumor growth without significant drug-related toxicity.
|
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | IGF-1R / AKT / MAPK |
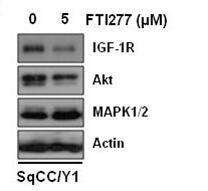
|
22113431 |
| Immunofluorescence | PAK phospho-PAK |
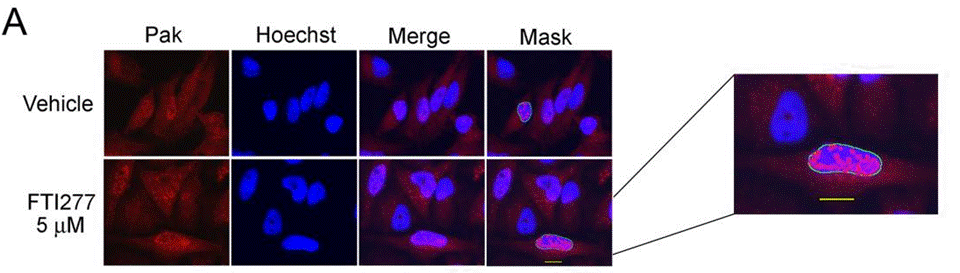
|
23915247 |
Clinical Trial Information
(data from https://clinicaltrials.gov, updated on 2024-05-22)
| NCT Number | Recruitment | Conditions | Sponsor/Collaborators | Start Date | Phases |
|---|---|---|---|---|---|
| NCT01240460 | Completed | Glioblastoma|Astrocytoma Grade IV |
Sanofi |
January 2011 | Phase 1 |
| NCT01013324 | Completed | Endometrial Cancer|Endometrial Neoplasms |
Sanofi |
January 2010 | Phase 2 |
| NCT00756847 | Completed | Cancer|Non-Small Cell Lung Cancer|Endometrial Carcinoma|Ovarian Carcinoma |
Sanofi |
September 2008 | Phase 1 |
| NCT00704392 | Withdrawn | Cancer|Non-small-cell Lung Cancer|Breast Cancer |
Exelixis |
June 2008 | Phase 1 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































