research use only
SCR7 DNA/RNA Synthesis inhibitor
Cat.No.S7742
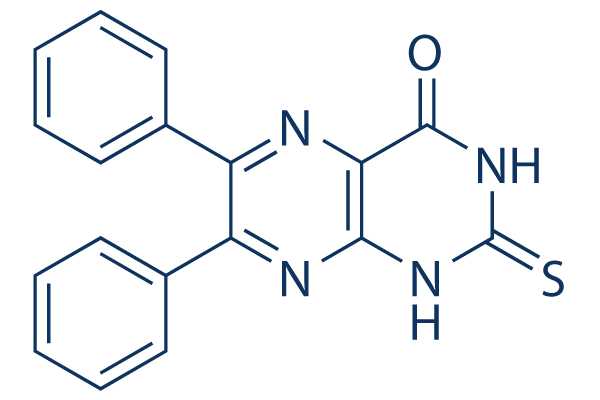
Chemical Structure
Molecular Weight: 332.38
Quality Control
| Related Targets | HDAC PARP ATM/ATR DNA-PK WRN Topoisomerase PPAR Sirtuin Casein Kinase eIF |
|---|---|
| Other DNA/RNA Synthesis Inhibitors | CX-5461 (Pidnarulex) Favipiravir (T-705) EED226 RK-33 BMH-21 Triapine (3-AP) Carmofur Thymidine YK-4-279 Tegafur (FT-207) |
Solubility
|
In vitro |
DMSO
: 66 mg/mL
(198.56 mM)
Ethanol : 23 mg/mL Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 332.38 | Formula | C18H12N4OS |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 14892-97-8 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N/A | Smiles | C1=CC=C(C=C1)C2=NC3=C(NC(=S)NC3=O)N=C2C4=CC=CC=C4 | ||
Mechanism of Action
| Targets/IC50/Ki |
DNA Ligase IV
|
|---|---|
| In vitro |
SCR7 effectively inhibits the formation of multimers at 200 μM and above. This compound successfully inhibits cell proliferation of MCF7, A549, HeLa, T47D, A2780, HT1080, and Nalm6 with IC50 of 40, 34, 44, 8.5, 120, 10, and 50 μM, respectively. It suppresses the NHEJ repair of CRISPR-Cas9-induced DSBs.This chemical increases the efficiency of HDR-mediated genome editing up to 19-fold using CRISPR/Cas9 in mammalian cells and mouse embryos. |
| Kinase Assay |
Complementation of SCR7 Inhibition with Purified Ligase IV
|
|
Complementation experiment is carried out by adding increasing concentrations of purified Ligase IV/XRCC4 complex (30, 60, and 120 fmol) along with the oligomeric DNA substrates (5’ compatible and 5’-5’ noncompatible ends) to the SCR7-treated extracts. Reactions are incubated for 2 h at 25℃. The reaction products are then resolved on 8% denaturing PAGE. The gel is dried and exposed and the signal is detected with a PhosphorImager and analyzed with Multi Gauge (V3.0) software.
|
|
| In vivo |
SCR7 treatment (10 mg/kg, i.m.) significantly reduces breast adenocarcinoma-induced tumor, and exhibits 4-fold increase in lifespan compared with control group. However, in Swiss albino mice with Dalton’s lymphoma tumor model, this compound (20 mg/kg, i.p.) exhibits neither tumor regression nor increase in lifespan. In BALB/c mice, this chemical (20 mg/kg, i.p.) significantly enhances the cytotoxic effects of radiation, VP-16213 and 3-Aminobenzamide on tumor derived from Dalton’s lymphoma (DLA) cells. |
References |
|
Clinical Trial Information
(data from https://clinicaltrials.gov, updated on 2024-05-22)
| NCT Number | Recruitment | Conditions | Sponsor/Collaborators | Start Date | Phases |
|---|---|---|---|---|---|
| NCT03941483 | Completed | Acute Kidney Injury (AKI) |
Astellas Pharma Inc |
November 1 2019 | Phase 2 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.
Frequently Asked Questions
Question 1:
What should I do if I got precipitate out of solution after a freeze thaw and cannot re-suspend it?
Answer:
Some compounds will precipitate out of solution after freeze/thaw, especially at very high concentration. You can warm it up to 45 degree and sonicate it to help it dissolve, of course, adding more DMSO will help too.






































