research use only
CX-5461 (Pidnarulex) RNA Polymerase I inhibitor
Cat.No.S2684
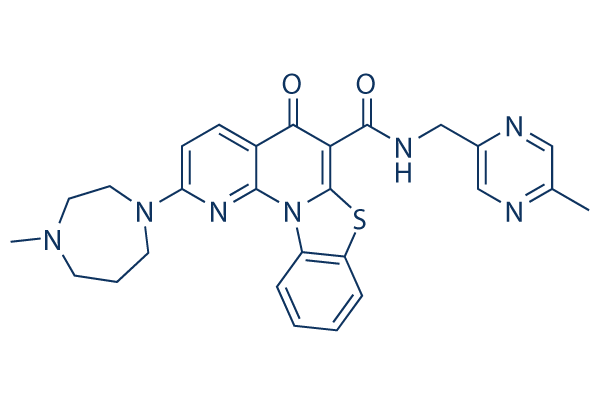
Chemical Structure
Molecular Weight: 513.61
Quality Control
| Related Targets | HDAC PARP ATM/ATR DNA-PK WRN Topoisomerase PPAR Sirtuin Casein Kinase eIF |
|---|---|
| Other DNA/RNA Synthesis Inhibitors | SCR7 Favipiravir (T-705) EED226 RK-33 BMH-21 Carmofur Triapine (3-AP) YK-4-279 Halofuginone Tegafur (FT-207) |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| MNNG | Cell viability assay | 72 h | IC50=0.5-1.5 µM | 27729807 | ||
| U2-OS | Cell viability assay | 72 h | IC50=0.5-1.5 µM | 27729807 | ||
| RS4;11 | Proliferation assay | 250 nM | 24 h | time dependent decrease in proliferation relative to their DMSO treated controls | 26472108 | |
| SEM | Proliferation assay | 250 nM | 24 h | time dependent decrease in proliferation relative to their DMSO treated controls | 26472108 | |
| KOPN-8 | Proliferation assay | 250 nM | 24 h | time dependent decrease in proliferation relative to their DMSO treated controls | 26472108 | |
| NALM-6 | Proliferation assay | 250 nM | 24 h | time dependent decrease in proliferation relative to their DMSO treated controls | 26472108 | |
| NB-EBc1 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for NB-EBc1 cells | 29435139 | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: Insoluble
Water : Insoluble Ethanol : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 513.61 | Formula | C27H27N7O2S |
Storage (From the date of receipt) | 3 years-20°C(in the dark) powder |
|---|---|---|---|---|---|
| CAS No. | 1138549-36-6 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N/A | Smiles | CC1=CN=C(C=N1)CNC(=O)C2=C3N(C4=CC=CC=C4S3)C5=C(C2=O)C=CC(=N5)N6CCCN(CC6)C | ||
Mechanism of Action
| Targets/IC50/Ki |
Pol I-driven transcription of rRNA
(HCT-116, A375, MIA PaCa-2 cells) 142 nM
|
|---|---|
| In vitro |
CX-5461 (Pidnarulex) is found to selectively inhibit rRNA synthesis (Pol I IC50=142 nM; Pol II IC50 > 25 μM; selectivity ~200-fold) in the HCT-116 cells. This selective inhibition is confirmed in two other human solid tumor cell lines; melanoma A375 (Pol I IC50 = 113 nM; Pol II IC50 > 25 μM) and pancreatic carcinoma MIA PaCa-2 (Pol I IC50=54 nM; Pol II IC50 ~25 mM). It possesses 250- to 300-fold selectivity for inhibition of rRNA transcription versus DNA replication and protein translation. The compound exhibits broad antiproliferative potency in a panel of cancer cell lines in human cancer cell lines, but has minimal effect on viability of nontransformed human cells. The median EC50 across all tested cell lines is 147 nM, yet all normal cell lines have EC50 values of approximately 5, 000 nM. Evaluation of the antiproliferative dose response for HCT-116, A375, and MIA PaCa-2 cell lines yield EC50 values of 167, 58, and 74 nM. It induces autophagy and senescence in solid tumor cancer cells, rather than apoptosis, through a p53-independent process. |
| Kinase Assay |
Pol I and Pol II Transcription Assay
|
|
Two short-lived RNA transcripts (half-lives ~20-30 minutes), one produced by Pol I and another by Pol II, are quantitated by qRT-PCR as a measure of CX-5461 (Pidnarulex)-related effects on transcription. The 45S pre-rRNA served as the Pol I transcript and the mRNA for the protooncogene c-myc served as the comparator Pol II transcript. Both Pol I and Pol II transcription are known to be affected by general cellular stress. To minimize the potential effects of such stress, cells are exposed to test agents for only a short period of time (2 hours). This is sufficient time for these transcripts to be reduced by greater than 90% if this compound affects their synthesis.
|
|
| In vivo |
CX-5461 (Pidnarulex) is orally bioavailable and demonstrates in vivo antitumor activity against human solid tumors in murine xenograft models. This compound demonstrates significant MIA PaCa-2 TGI with TGI equal to 69% on day 31. Likewise, it demonstrates significant A375 TGI with TGI equal to 79% on day 32. |
References |
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | Cyclin D1 / p53 / p16 53BP1 / γ-H2AX / p-ATM Bcl-2 / Bax / Caspase3 cleaved PARP / cleaved Caspase-9 / cleaved caspase-3 p-AMPK / AMPK / p-mTOR / mTOR EG5 / Histone H3 / p-Histone H3 (S10) |
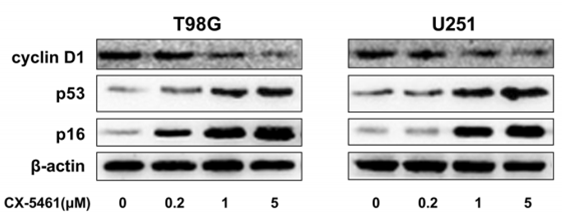
|
29631594 |
| Immunofluorescence | Vimentin / Phalloidin / Snail1 Rictor / Calnexin Fibrillarin LC3 γH2AX / 53BP1 / RPA / RAD51 chromosome / telomere F-actin / Aurora B |
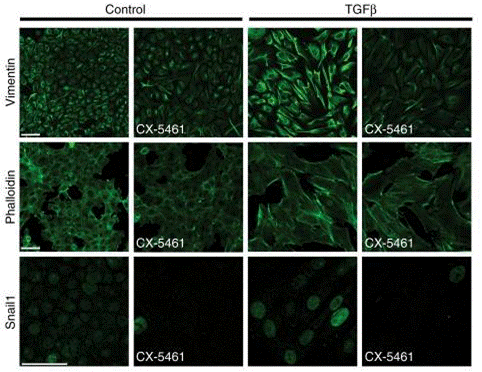
|
31068593 |
| Growth inhibition assay | Cell viability |
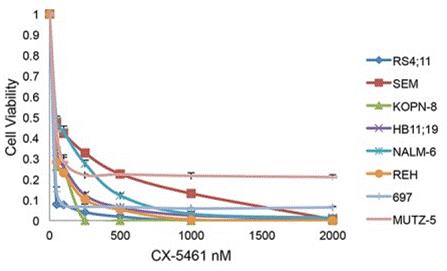
|
26061708 |
Clinical Trial Information
(data from https://clinicaltrials.gov, updated on 2024-05-22)
| NCT Number | Recruitment | Conditions | Sponsor/Collaborators | Start Date | Phases |
|---|---|---|---|---|---|
| NCT02719977 | Completed | Cancer |
Canadian Cancer Trials Group|Senhwa Biosciences Inc.|Stand Up To Cancer |
June 13 2016 | Phase 1 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.
Frequently Asked Questions
Question 1:
I want to make it for further in vivo treatment.
Answer:
This compound has poor solubility in common vehicles. It can be dissolved in DMF at 3 mg/ml with warming.






































