research use only
Niraparib tosylate PARP inhibitor
Cat.No.S7625
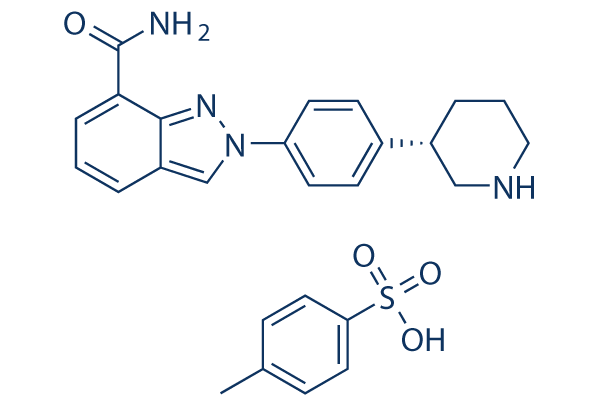
Chemical Structure
Molecular Weight: 492.59
Quality Control
| Related Targets | HDAC ATM/ATR DNA-PK WRN DNA/RNA Synthesis Topoisomerase PPAR Sirtuin Casein Kinase eIF |
|---|---|
| Other PARP Inhibitors | XAV-939 AZD5305 (Saruparib) Veliparib (ABT-888) PJ34 HCl AG-14361 Iniparib (BSI-201) G007-LK UPF 1069 A-966492 Pamiparib |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| HeLa | Function assay | Inhibition of PARP in hydrogen peroxide-induced human HeLa cells assessed as inhibition DNA-damage-induced PARylation, EC50 = 0.004 μM. | 19873981 | |||
| MDA-MB-436 | Antiproliferative assay | 6 days | Antiproliferative activity against human MDA-MB-436 cells expressing BRCA1 5396 + 1G>A mutant after 6 days by cell titer-blue assay, CC50 = 0.018 μM. | 19873981 | ||
| HeLa | Antiproliferative assay | 7 days | Antiproliferative activity against BRCA1 deficient human HeLa cells after 7 days by cell titer-blue assay, CC50 = 0.033 μM. | 19873981 | ||
| HeLa | Function assay | Inhibition of PARP in hydrogen peroxide-induced human HeLa cells assessed as inhibition DNA-damage-induced PARylation, EC90 = 0.045 μM. | 19873981 | |||
| Capan1 | Antiproliferative assay | 13 days | Antiproliferative activity against human Capan1 cells expressing BRCA2 6174delT mutation and loss of wild-type allele after 13 days by cell titer-blue assay, CC50 = 0.09 μM. | 19873981 | ||
| HeLa | Antiproliferative assay | 7 days | Antiproliferative activity against human HeLa cells expressing wild type BRCA1 after 7 days by cell titer-blue assay, CC50 = 0.86 μM. | 19873981 | ||
| MDA-MB-436 | Antitumor assay | 50 mg/kg | 33 days | Antitumor activity against human MDA-MB-436 cells expressing BRCA1 5396 + 1G>A mutant xenografted in CD1 mouse assessed as tumor regression at 50 mg/kg, po bid for 33 days | 19873981 | |
| MDA-MB-436 | Antitumor assay | 100 mg/kg | 33 days | Antitumor activity against human MDA-MB-436 cells expressing BRCA1 5396 + 1G>A mutant xenografted in CD1 mouse assessed as tumor regression at 100 mg/kg, po qd for 33 days | 19873981 | |
| Jurkat | Function assay | 96 hrs | Inhibition of PARP1 in human Jurkat cells assessed as reduction of cell viability after 96 hrs by MTS assay in presence of 100 uM of temozolomide, EC50 = 0.2 μM. | 23850199 | ||
| Jurkat | Function assay | 96 hrs | Inhibition of PARP1 in human Jurkat cells assessed as reduction of cell viability after 96 hrs by MTS assay, EC50 = 31 μM. | 23850199 | ||
| CAPAN-1 | Function assay | Inhibition of PARP in BRCA2-deficient human CAPAN-1 cells assessed as inhibition of hydrogen peroxide-induced PARylation by cell-based assay, IC50 = 0.0035 μM. | 25761096 | |||
| HeLa | Function assay | Inhibition of PARP in human HeLa cells assessed as inhibition of hydrogen peroxide-induced PARylation by cell-based assay, EC50 = 0.004 μM. | 25761096 | |||
| A2780 | Function assay | Inhibition of PARP in human A2780 cells assessed as inhibition of hydrogen peroxide-induced PARylation by cell-based assay, IC50 = 0.004 μM. | 25761096 | |||
| A549 | Cytotoxicity assay | 5 to 7 days | Cytotoxicity against human A549 cells transfected with BRCA2 shRNA assessed as inhibition of cell proliferation after 5 to 7 days by CellTiter-Blue assay, CC50 = 0.011 μM. | 25761096 | ||
| MDA-MB-436 | Cytotoxicity assay | Cytotoxicity against human MDA-MB-436 cells carrying BRCA1 mutant assessed as inhibition of cell proliferation, CC50 = 0.018 μM. | 25761096 | |||
| SUM1315MO2 | Cytotoxicity assay | 12 days | Cytotoxicity against human SUM1315MO2 cells carrying BRCA1 mutant assessed as inhibition of cell proliferation after 12 days by CellTiter-Blue assay, CC50 = 0.02 μM. | 25761096 | ||
| DoTc2-4510 | Cytotoxicity assay | 5 to 7 days | Cytotoxicity against human DoTc2-4510 cells carrying BRCA2 mutant assessed as inhibition of cell proliferation after 5 to 7 days by CellTiter-Blue assay, CC50 = 0.023 μM. | 25761096 | ||
| SUM149PT | Cytotoxicity assay | 5 to 7 days | Cytotoxicity against human SUM149PT cells carrying BRCA1 mutant assessed as inhibition of cell proliferation after 5 to 7 days by CellTiter-Blue assay, CC50 = 0.024 μM. | 25761096 | ||
| HeLa | Cytotoxicity assay | 5 to 7 days | Cytotoxicity against human HeLa cells transfected with BRCA1 shRNA assessed as reduction of cell viability after 5 to 7 days by CellTiter-Blue assay, CC50 = 0.034 μM. | 25761096 | ||
| HeLa | Function assay | Inhibition of PARP in human HeLa cells assessed as inhibition of hydrogen peroxide-induced PARylation by cell-based assay, IC90 = 0.046 μM. | 25761096 | |||
| CAPAN-1 | Function assay | Inhibition of PARP in BRCA2-deficient human CAPAN-1 cells assessed as inhibition of hydrogen peroxide-induced PARylation by cell-based assay, IC90 = 0.05 μM. | 25761096 | |||
| A2780 | Function assay | Inhibition of PARP in human A2780 cells assessed as inhibition of hydrogen peroxide-induced PARylation by cell-based assay, IC90 = 0.052 μM. | 25761096 | |||
| UWB1.289 | Cytotoxicity assay | 5 to 7 days | Cytotoxicity against human UWB1.289 cells carrying BRCA1 mutant assessed as inhibition of cell proliferation after 5 to 7 days by CellTiter-Blue assay, CC50 = 0.056 μM. | 25761096 | ||
| Capan1 | Cytotoxicity assay | Cytotoxicity against BRCA2-deficient human Capan1 cells, CC50 = 0.09 μM. | 25761096 | |||
| HeLa | Cytotoxicity assay | 5 to 7 days | Cytotoxicity against wild type human HeLa cells assessed as inhibition of cell proliferation after 5 to 7 days by CellTiter-Blue assay, CC50 = 0.852 μM. | 25761096 | ||
| UWB1.289 | Cytotoxicity assay | 5 to 7 days | Cytotoxicity against human UWB1.289 cells expressing BRCA1 assessed as inhibition of cell proliferation after 5 to 7 days by CellTiter-Blue assay, CC50 = 0.975 μM. | 25761096 | ||
| A549 | Cytotoxicity assay | 5 to 7 days | Cytotoxicity against wild type human A549 cells assessed as inhibition of cell proliferation after 5 to 7 days by CellTiter-Blue assay, CC50 = 1.76 μM. | 25761096 | ||
| BT20 | Cytotoxicity assay | 5 to 7 days | Cytotoxicity against human BT20 cells assessed as inhibition of cell proliferation after 5 to 7 days by CellTiter-Blue assay, CC50 = 2.2 μM. | 25761096 | ||
| MDA-MB-436 | Antitumor assay | 80 mg/kg | 1 to 2 weeks | Antitumor activity against human MDA-MB-436 cells harboring BRCA1 mutant xenografted in immunocompromised mouse assessed as tumor growth inhibition at 80 mg/kg, po qd for 1 to 2 weeks | 25761096 | |
| MDA-MB-436 | Antitumor assay | 80 mg/kg | 4 weeks | Antitumor activity against human MDA-MB-436 cells harboring BRCA1 mutant xenografted in immunocompromised mouse assessed as complete and sustained tumor regression at 80 mg/kg, po qd for 4 weeks | 25761096 | |
| MDA-MB-436 | Antitumor assay | 50 mg/kg | Antitumor activity against human MDA-MB-436 cells harboring BRCA1 mutant xenografted in immunocompromised mouse assessed as tumor growth inhibition at 50 mg/kg, po administered daily | 25761096 | ||
| MDA-MB-436 | Antitumor assay | 80 mg/kg | 3 weeks | Antitumor activity against human MDA-MB-436 cells harboring BRCA1 mutant xenografted in immunocompromised mouse assessed as tumor shrinkage at 80 mg/kg, po qd for 3 weeks | 25761096 | |
| Capan1 | Cytotoxicity assay | Cytotoxicity against BRCA2-deficient human Capan1 cells, EC50 = 0.65 μM. | 26652717 | |||
| TC32 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for TC32 cells | 29435139 | |||
| A673 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for A673 cells | 29435139 | |||
| NB1643 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for NB1643 cells | 29435139 | |||
| A673 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for A673 cells) | 29435139 | |||
| BT-37 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for BT-37 cells | 29435139 | |||
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SK-N-MC cells | 29435139 | |||
| NB-EBc1 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for NB-EBc1 cells | 29435139 | |||
| LAN-5 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for LAN-5 cells | 29435139 | |||
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for SK-N-MC cells | 29435139 | |||
| TC32 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for TC32 cells | 29435139 | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 99 mg/mL
(200.97 mM)
Water : Insoluble Ethanol : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 492.59 | Formula | C19H20N4O.C7H8O3S |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 1038915-73-9 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | MK 4827 tosylate | Smiles | CC1=CC=C(C=C1)S(=O)(=O)O.C1CC(CNC1)C2=CC=C(C=C2)N3C=C4C=CC=C(C4=N3)C(=O)N | ||
Mechanism of Action
| Targets/IC50/Ki |
PARP2
(Cell-free assay) 2.1 nM
PARP1
(Cell-free assay) 3.8 nM
|
|---|---|
| In vitro |
Micromolar concentrations of niraparib radiosensitizes tumor cell lines derived from lung, breast, and prostate cancers independently of their p53 status but not cell lines derived from normal tissues. Niraparib also sensitizes tumor cells to H2O2 and converts H2O2-induced single strand breaks (SSBs) into DSBs during DNA replication. |
| In vivo |
MK-4827 strongly enhances the effect of radiation on a variety of human tumor xenografts, both p53 wild type and p53 mutant. MK-4827 reduces PAR levels in tumors by 1 h after administration which persisted for up to 24 h. In vivo treatment with MK-4827 and radiation prolonged survival (p<0.01) compared to single modalities. In vivo superiority of MK-4827 plus radiation is further documented by significant elevations of cleaved caspase-3 and γ-H2AX in tumors from the combination group compared to single modality cohorts. |
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | c-PARP /c-caspase 3 / γ-H2AX |
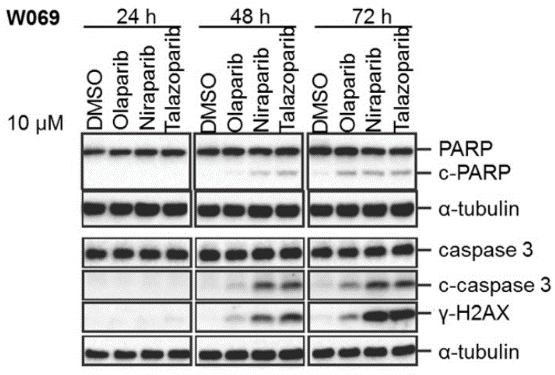
|
29158830 |
| Immunofluorescence | Rad51 / Geminin |
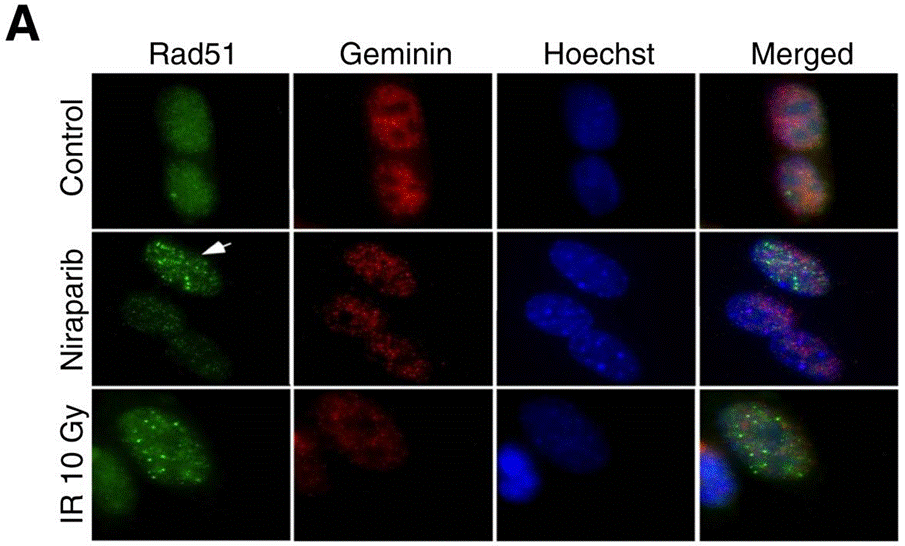
|
27614696 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































