research use only
PIK-III Autophagy inhibitor
Cat.No.S7683
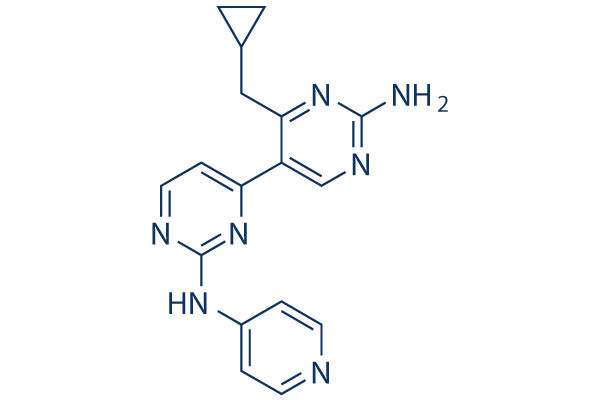
Chemical Structure
Molecular Weight: 319.36
Quality Control
| Related Targets | CXCR Nrf2 Mitophagy LRRK2 ULK FKBP Heme Oxygenase cGAS LC3 Cell wall |
|---|---|
| Other Autophagy Inhibitors | Resveratrol (trans-Resveratrol) Spautin-1 Lys05 Trihydrochloride DC661 Spermidine Autophinib SMER28 NSC 185058 EN6 CA-5f |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| U2OS | Function assay | 2 hrs | Inhibition of VPS34 in human U2OS cells incubated for 2 hrs by GFP-FYVE reporter gene assay, IC50=0.012μM | 26819669 | ||
| DLD1 | Function assay | 1 to 10 uM | 24 hrs | Inhibition of VPS34 in human DLD1 cells assessed as prevention of NCOA4 protein degradation at 1 to 10 uM incubated for 24 hrs by Western blot analysis | 26819669 | |
| DLD1 | Function assay | 1 to 10 uM | 24 hrs | Inhibition of VPS34 in human DLD1 cells assessed as prevention of p62 degradation at 1 to 10 uM incubated for 24 hrs by Western blot analysis | 26819669 | |
| DLD1 | Function assay | 1 to 10 uM | 24 hrs | Inhibition of VPS34 in human DLD1 cells assessed as prevention of NBR1 protein degradation at 1 to 10 uM incubated for 24 hrs by Western blot analysis | 26819669 | |
| DLD1 | Function assay | 1 to 10 uM | 24 hrs | Inhibition of VPS34 in human DLD1 cells assessed as prevention of NDP52 protein degradation at 1 to 10 uM incubated for 24 hrs by Western blot analysis | 26819669 | |
| DLD1 | Function assay | 1 to 10 uM | 24 hrs | Inhibition of VPS34 in human DLD1 cells assessed as prevention of FTH1 protein degradation at 1 to 10 uM incubated for 24 hrs by Western blot analysis | 26819669 | |
| VERO-E6 | Toxicity assay | 48 hrs | Toxicity CC50 against VERO-E6 cells determined at 48 hours by high content imaging (same conditions as 2_LEY without exposure to 0.01 MOI SARS CoV-2 virus), CC50=2.11μM | ChEMBL | ||
| VERO-E6 | Function assay | 48 hrs | Determination of IC50 values for inhibition of SARS-CoV-2 induced cytotoxicity of VERO-E6 cells after 48 hours exposure to 0.01 MOI SARS CoV-2 virus by high content imaging, IC50=7.92μM | ChEMBL | ||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 63 mg/mL
(197.26 mM)
Ethanol : 63 mg/mL Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 319.36 | Formula | C17H17N7 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 1383716-40-2 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | VPS34-IN2 | Smiles | C1CC1CC2=NC(=NC=C2C3=NC(=NC=C3)NC4=CC=NC=C4)N | ||
Mechanism of Action
| Targets/IC50/Ki |
Vps34
(Cell-free assay) 0.018μM
PI3Kδ
(Cell-free assay) 1.2μM
|
|---|---|
| In vitro |
VPS34 enzymatic function is essential for LC3 lipidation in mammalian cells and PIK-III is a robust inhibitor of autophagy and LC3 lipidation in mammalian cells.
In H4 cells, this compound inhibits the formation of autolysosomes and increases the cytosolic signal of LC3 under basal conditions and when autophagy is induced with the mTOR inhibitor AZD8055.
In a CCCP-induced mitophagy model, it inhibits the clearance of mitochondria.This chemical treatment leads to an increase in the levels of LC3-I in H4 and PSN1 cells.
In Panc10.05 cells, this inhibitor increases the levels of LC3-II in parallel with LC3-I suggesting a cell type-specific response.
|
References |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































