research use only
RSL3 Ferroptosis inducer
Cat.No.S8155
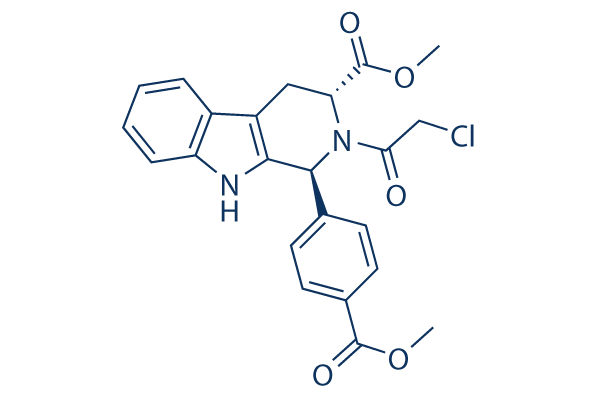
Chemical Structure
Molecular Weight: 440.88
Quality Control
Products Often Used Together with RSL3
| Related Targets | Dehydrogenase HSP Transferase P450 (e.g. CYP17) PDE phosphatase PPAR Vitamin Carbohydrate Metabolism Mitochondrial Metabolism |
|---|---|
| Other Ferroptosis Inhibitors | Imidazole Ketone Erastin (IKE) UAMC-3203 iFSP1 SRS11-92 icFSP1 N6F11 SRS16-86 Bufotalin Erastin2 NPD4928 |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| MEFs and HT1080 cells | Function assay | 0.5 μM | 24 h | 30686534 | ||
| Jurkat | Cell death assay | 0.1 μM | 24 h or 48 h | BV6 cooperates with RSL3 to induce cell death, accompanied by ROS production and lipid peroxidation | 27588473 | |
| Molt-4 | Cell death assay | 0.075 μM | 24 h or 48 h | BV6 cooperates with RSL3 to induce cell death, accompanied by ROS production and lipid peroxidation | 27588473 | |
| RMS13 cells | Cell death assay | 0, 60, 100, 140 and 180 μM | 48 h | Addition of Ferrostatin-1 significantly reduced RSL3- or Erastin-induced loss of cell viability. | 26157704 | |
| BJeLR | Cytotoxicity assay | 0.57 uM | 12 h | Cytotoxicity against human BJeLR cells expressing RAS G12V mutant at 0.57 uM at 12 hrs by trypan blue staining | 22832321 | |
| BJeLR | Cytotoxicity assay | 0.57 uM | 24 h | Cytotoxicity against human BJeLR cells expressing RAS G12V mutant at 0.57 uM at 24 hrs by trypan blue staining | 22832321 | |
| BJeH | Function assay | 6 h | Induction of reactive oxygen species production in human BJeH cells expressing wild type RAS after 6 hrs by DCF-based flow cytometric analysis | 22832321 | ||
| BJeLR | Function assay | 6 h | Induction of reactive oxygen species production in human BJeLR cells expressing RAS G12V mutant after 6 hrs by DCF-based flow cytometric analysis | 22832321 | ||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 88 mg/mL
(199.6 mM)
Ethanol : 28 mg/mL Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 440.88 | Formula | C23H21ClN2O5 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 1219810-16-8 | Download SDF | Storage of Stock Solutions |
|
|
Mechanism of Action
| Targets/IC50/Ki |
Gpx4
(In Calu-1 cells) |
|---|---|
| In vitro |
Ferroptosis-inducing compounds inactivate GPX4 by direct binding or by depleting glutathione.After binding to GPX4, this compound inactivates GPX4 to induce ROS production from lipid peroxidation. Its ability to induce synthetic lethality with oncogenic RAS is rapid and quite potent. This compound inhibits the growth of BJ-TERT/LT/ST/RASV12 and DRD cells as low as 10 ng/mland started to kill sensitive cells as early as 8 hr after treatment.
|
| In vivo |
Prostaglandin-endoperoxide synthase (PTGS) is the key enzyme in prostaglandin biosynthesis. There are two isozymes of PTGS: a constitutive PTGS1 and an inducible PTGS2. PTGS2 encoding cyclooxygenase-2 (COX-2) is significantly upregulated after treatment with RSL3 and this compound in mice.
|
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | GPX4 / Tranferrin / Ferritin HO-1 |
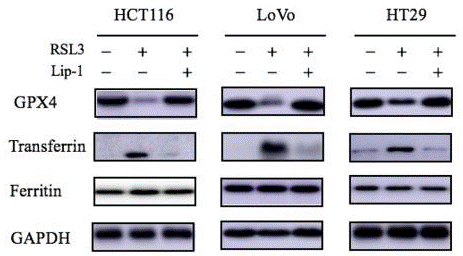
|
30524291 |
| Growth inhibition assay | Cell viability |
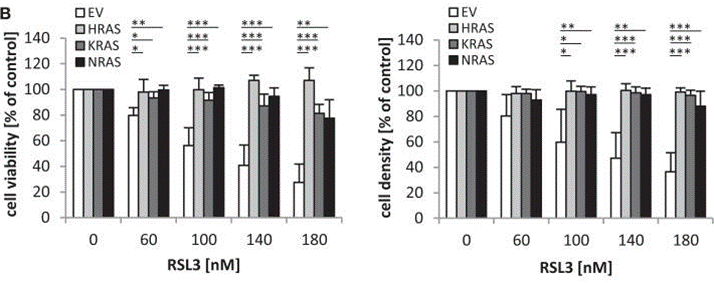
|
26157704 |
| Immunofluorescence | ALOX12 / ALOX15 4-HNE |
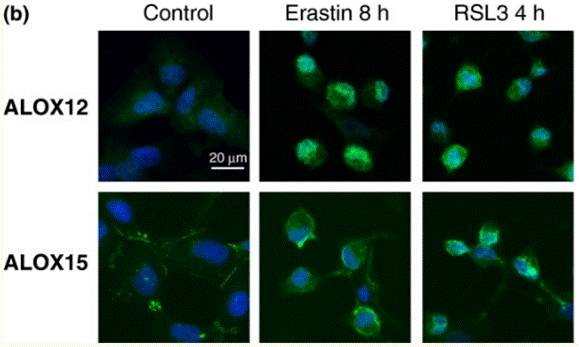
|
28837253 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































