research use only
Palomid 529 (P529) mTOR inhibitor
Cat.No.S2238
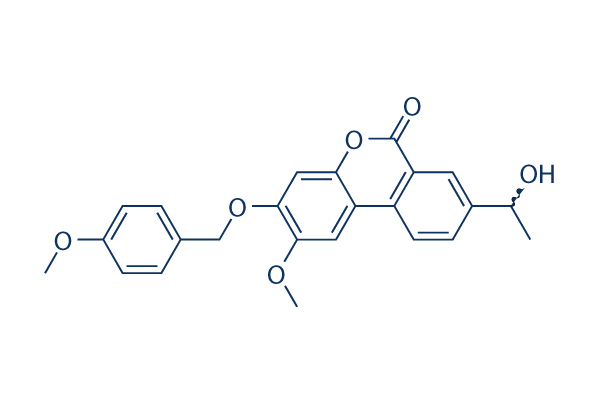
Chemical Structure
Molecular Weight: 406.43
Quality Control
| Related Targets | PI3K Akt GSK-3 ATM/ATR DNA-PK AMPK PDPK1 PTEN PP2A PDK |
|---|---|
| Other mTOR Inhibitors | Torin 1 Torin 2 AZD8055 Ridaforolimus (Deforolimus, MK-8669) Sapanisertib (MLN0128, INK-128) Torkinib (PP242) MHY1485 Vistusertib (AZD2014) KU-0063794 OSI-027 |
Solubility
|
In vitro |
DMSO
: 81 mg/mL
(199.29 mM)
Water : Insoluble Ethanol : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 406.43 | Formula | C24H22O6 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 914913-88-5 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | SG 00529 | Smiles | CC(C1=CC2=C(C=C1)C3=CC(=C(C=C3OC2=O)OCC4=CC=C(C=C4)OC)OC)O | ||
Mechanism of Action
| Targets/IC50/Ki |
mTORC1
mTORC2
|
|---|---|
| In vitro |
Palomid 529 (P529) inhibits proliferation and increases apoptosis of endothelial cells, inhibiting both VEGF-driven and bFGF-driven endothelial cell proliferation with IC50 of 20 nM and 30 nM, respectively. It retains the ability to induce endothelial cell apoptosis and decreases VEGF-A–driven phosphorylation of pAktS473, pGSK3βS9, and pS6. However, this compound prevents neither phosphorylated mitogen-activated protein kinase (pMAPK) nor pAktT308 as potently as pAktS473. It not only reduces the proliferative response in the ischemic retina but also improves the organization and structure of the vessels that form. P529 shows a potent antiproliferative activity in the NCI-60 cell lines panel, with growth inhibitory 50 (GI50) <35 μM. In addition, it significantly enhances the antiproliferative effect of radiation in prostate cancer cells (PC-3), giving rise to a concentration dependent growth inhibition on PC-3 cells. Doses of 2 and 7μM resulted in 30 and 60% growth inhibition, respectively. It inhibits the radiation-induced p-Akt activation and decreases Bcl-2/Bax ratio in PC-3. This compound not only inhibits radiation-induced overexpression of Id-1 and VEGF but also down-regulates radiation-induced MMP-2 and MMP-9. |
| Kinase Assay |
Estrogen receptor binding assays
|
|
The proteins are produced with rabbit reticulocyte lysates. The amount of template used in each reaction is determined empirically and expression is monitored in parallel reactions where [35S]methionine is incorporated into the receptor followed by gel electrophoresis and exposure to film. Binding reactions of the estrogen receptors (ER) and Palomid 529 (P529) are carried out in 100 mL final volumes in TEG buffer [10 mM Tris (pH 7.5), 1.5 mM EDTA, 10% glycerol]. In vitro transcribed-translated receptor (5 μL) is used in each binding reaction in the presence of 0.5 nM [3H]estradiol (E2). This compound is routinely tested from 10−11 to 10−6 M and diluted in ethanol. The reactions are incubated at 4 °C overnight and bound E2 is quantified by adding 200 mL dextran-coated charcoal. After a 15-minutes rotation at 4 °C, the tubes are centrifuged for 10 minutes and 150 mL of the supernatant are added to 5 mL scintillation mixture for determination of cpm by liquid scintillation counting. The maximum binding is determined by competing bound E2 with only the ethanol vehicle. Controls for background are included in each experiment using 5 mL unprogrammed rabbit reticulocyte lysate. This value, typically 10% to 15% of the maximal counts, is subtracted from all values. The data are plotted and Ki values are calculated. Experiments are conducted at least thrice in duplicate.
|
|
| In vivo |
Palomid 529 (P529) shows a dose-dependent inhibition of the Ad-VEGF-A–driven angiogenesis following its treatment. It inhibits C6V10 glioma tumor growth in nude mice following i.p. dosing and decreases AktS473 but not AktT308 signaling. This compound also inhibits tumor growth, angiogenesis, and vascular permeability. Treatment of PC-3 tumour-bearing mice with it reduced tumour growth to 57.1% compared with controls. It is an effective suppressor of Müller cell proliferation, glial scar formation, and photoreceptor cell death in a rabbit model of retinal detachment (RD). Palomid 529 significantly suppresses Brca1-deficient tumor growth in mice through inhibition of both Akt and mTOR signaling. |
References |
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































