research use only
KU-55933 ATM Inhibitor
Cat.No.S1092
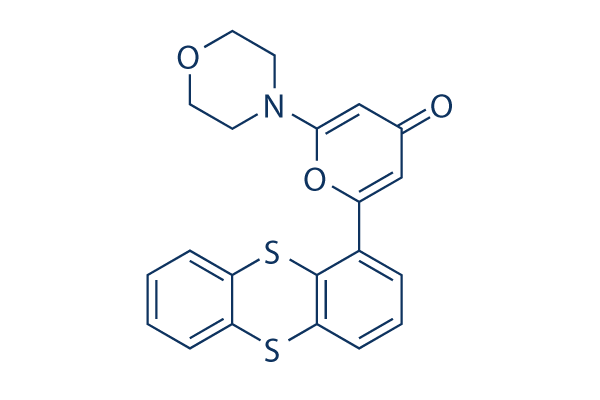
Chemical Structure
Molecular Weight: 395.49
Quality Control
| Related Targets | HDAC PARP DNA-PK WRN DNA/RNA Synthesis Topoisomerase PPAR Sirtuin Casein Kinase eIF |
|---|---|
| Other ATM/ATR Inhibitors | Ceralasertib (AZD6738) AZD1390 Berzosertib (VE-822) Lartesertib (M4076) Camonsertib (RP-3500) KU-60019 VE-821 AZ20 AZD0156 Mirin |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| DU-145 | Growth Inhibition Assay | IC50=3.27352 μM | SANGER | |||
| HuO-3N1 | Growth Inhibition Assay | IC50=4.17142 μM | SANGER | |||
| LAMA-84 | Growth Inhibition Assay | IC50=4.58465 μM | SANGER | |||
| CAL-72 | Growth Inhibition Assay | IC50=5.48084 μM | SANGER | |||
| LoVo | Growth Inhibition Assay | IC50=6.93239 μM | SANGER | |||
| HH | Growth Inhibition Assay | IC50=8.27671 μM | SANGER | |||
| SK-MEL-3 | Growth Inhibition Assay | IC50=8.28575 μM | SANGER | |||
| KM12 | Growth Inhibition Assay | IC50=9.21142 μM | SANGER | |||
| NCI-H1437 | Growth Inhibition Assay | IC50=9.8097 μM | SANGER | |||
| NCI-H1838 | Growth Inhibition Assay | IC50=11.1865 μM | SANGER | |||
| J-RT3-T3-5 | Growth Inhibition Assay | IC50=11.2417 μM | SANGER | |||
| GOTO | Growth Inhibition Assay | IC50=11.6996 μM | SANGER | |||
| LB2241-RCC | Growth Inhibition Assay | IC50=11.7186 μM | SANGER | |||
| ES7 | Growth Inhibition Assay | IC50=11.788 μM | SANGER | |||
| KP-N-YS | Growth Inhibition Assay | IC50=12.6354 μM | SANGER | |||
| CAL-12T | Growth Inhibition Assay | IC50=13.617 μM | SANGER | |||
| COLO-684 | Growth Inhibition Assay | IC50=14.1569 μM | SANGER | |||
| DOK | Growth Inhibition Assay | IC50=15.3329 μM | SANGER | |||
| Hs-578-T | Growth Inhibition Assay | IC50=15.4182 μM | SANGER | |||
| D-423MG | Growth Inhibition Assay | IC50=15.5236 μM | SANGER | |||
| DBTRG-05MG | Growth Inhibition Assay | IC50=15.6111 μM | SANGER | |||
| VM-CUB-1 | Growth Inhibition Assay | IC50=15.9849 μM | SANGER | |||
| KG-1 | Growth Inhibition Assay | IC50=16.0996 μM | SANGER | |||
| 8305C | Growth Inhibition Assay | IC50=16.1889 μM | SANGER | |||
| HuH-7 | Growth Inhibition Assay | IC50=16.2674 μM | SANGER | |||
| LXF-289 | Growth Inhibition Assay | IC50=16.2747 μM | SANGER | |||
| NCI-H1793 | Growth Inhibition Assay | IC50=16.4712 μM | SANGER | |||
| ChaGo-K-1 | Growth Inhibition Assay | IC50=16.6568 μM | SANGER | |||
| GCIY | Growth Inhibition Assay | IC50=16.7905 μM | SANGER | |||
| SK-MEL-28 | Growth Inhibition Assay | IC50=17.0475 μM | SANGER | |||
| NCI-SNU-1 | Growth Inhibition Assay | IC50=17.1269 μM | SANGER | |||
| CTB-1 | Growth Inhibition Assay | IC50=17.2259 μM | SANGER | |||
| NCI-H82 | Growth Inhibition Assay | IC50=17.4573 μM | SANGER | |||
| HCC2998 | Growth Inhibition Assay | IC50=17.6733 μM | SANGER | |||
| NCI-H2030 | Growth Inhibition Assay | IC50=18.1997 μM | SANGER | |||
| HuP-T3 | Growth Inhibition Assay | IC50=18.5888 μM | SANGER | |||
| 697 | Growth Inhibition Assay | IC50=19.0201 μM | SANGER | |||
| MLMA | Growth Inhibition Assay | IC50=19.0557 μM | SANGER | |||
| HCC70 | Growth Inhibition Assay | IC50=19.489 μM | SANGER | |||
| A704 | Growth Inhibition Assay | IC50=19.8305 μM | SANGER | |||
| D-283MED | Growth Inhibition Assay | IC50=20.5339 μM | SANGER | |||
| U031 | Growth Inhibition Assay | IC50=21.1489 μM | SANGER | |||
| HSC-3 | Growth Inhibition Assay | IC50=21.1835 μM | SANGER | |||
| JVM-3 | Growth Inhibition Assay | IC50=22.506 μM | SANGER | |||
| Mewo | Growth Inhibition Assay | IC50=22.5073 μM | SANGER | |||
| YH-13 | Growth Inhibition Assay | IC50=22.5123 μM | SANGER | |||
| LB1047-RCC | Growth Inhibition Assay | IC50=22.5879 μM | SANGER | |||
| HCC2157 | Growth Inhibition Assay | IC50=22.8054 μM | SANGER | |||
| SNU-449 | Growth Inhibition Assay | IC50=22.8748 μM | SANGER | |||
| Ramos-2G6-4C10 | Growth Inhibition Assay | IC50=22.96 μM | SANGER | |||
| CHL-1 | Growth Inhibition Assay | IC50=23.7292 μM | SANGER | |||
| SK-MEL-30 | Growth Inhibition Assay | IC50=24.4662 μM | SANGER | |||
| PANC-08-13 | Growth Inhibition Assay | IC50=25.0938 μM | SANGER | |||
| QIMR-WIL | Growth Inhibition Assay | IC50=25.1858 μM | SANGER | |||
| BFTC-905 | Growth Inhibition Assay | IC50=25.5944 μM | SANGER | |||
| GI-1 | Growth Inhibition Assay | IC50=25.7055 μM | SANGER | |||
| MDA-MB-415 | Growth Inhibition Assay | IC50=26.5033 μM | SANGER | |||
| GT3TKB | Growth Inhibition Assay | IC50=26.5342 μM | SANGER | |||
| DEL | Growth Inhibition Assay | IC50=26.8356 μM | SANGER | |||
| KOSC-2 | Growth Inhibition Assay | IC50=26.9075 μM | SANGER | |||
| RVH-421 | Growth Inhibition Assay | IC50=27.2921 μM | SANGER | |||
| EW-13 | Growth Inhibition Assay | IC50=27.4308 μM | SANGER | |||
| 639-V | Growth Inhibition Assay | IC50=27.5119 μM | SANGER | |||
| A2780 | Growth Inhibition Assay | IC50=27.641 μM | SANGER | |||
| SW982 | Growth Inhibition Assay | IC50=27.9052 μM | SANGER | |||
| SW1710 | Growth Inhibition Assay | IC50=28.0981 μM | SANGER | |||
| HCC1569 | Growth Inhibition Assay | IC50=28.4897 μM | SANGER | |||
| MV-4-11 | Growth Inhibition Assay | IC50=28.5735 μM | SANGER | |||
| BHT-101 | Growth Inhibition Assay | IC50=28.6572 μM | SANGER | |||
| Ca9-22 | Growth Inhibition Assay | IC50=28.714 μM | SANGER | |||
| HAL-01 | Growth Inhibition Assay | IC50=28.7615 μM | SANGER | |||
| D-263MG | Growth Inhibition Assay | IC50=29.344 μM | SANGER | |||
| NEC8 | Growth Inhibition Assay | IC50=29.5548 μM | SANGER | |||
| EKVX | Growth Inhibition Assay | IC50=31.5847 μM | SANGER | |||
| EM-2 | Growth Inhibition Assay | IC50=31.6304 μM | SANGER | |||
| MFM-223 | Growth Inhibition Assay | IC50=31.8098 μM | SANGER | |||
| SK-PN-DW | Growth Inhibition Assay | IC50=32.1406 μM | SANGER | |||
| HuO9 | Growth Inhibition Assay | IC50=32.5282 μM | SANGER | |||
| MHH-PREB-1 | Growth Inhibition Assay | IC50=32.6234 μM | SANGER | |||
| OVCAR-4 | Growth Inhibition Assay | IC50=32.8363 μM | SANGER | |||
| NCI-H1648 | Growth Inhibition Assay | IC50=32.8651 μM | SANGER | |||
| MKN1 | Growth Inhibition Assay | IC50=34.1101 μM | SANGER | |||
| KYSE-450 | Growth Inhibition Assay | IC50=34.6444 μM | SANGER | |||
| ES8 | Growth Inhibition Assay | IC50=34.8975 μM | SANGER | |||
| MS-1 | Growth Inhibition Assay | IC50=34.9554 μM | SANGER | |||
| HOP-92 | Growth Inhibition Assay | IC50=35.9277 μM | SANGER | |||
| SKG-IIIa | Growth Inhibition Assay | IC50=36.2561 μM | SANGER | |||
| TE-11 | Growth Inhibition Assay | IC50=36.5243 μM | SANGER | |||
| SK-NEP-1 | Growth Inhibition Assay | IC50=37.6744 μM | SANGER | |||
| DB | Growth Inhibition Assay | IC50=37.9185 μM | SANGER | |||
| IA-LM | Growth Inhibition Assay | IC50=38.0239 μM | SANGER | |||
| COLO-829 | Growth Inhibition Assay | IC50=38.4159 μM | SANGER | |||
| TGBC11TKB | Growth Inhibition Assay | IC50=39.1408 μM | SANGER | |||
| CAL-51 | Growth Inhibition Assay | IC50=40.0612 μM | SANGER | |||
| NCI-H2228 | Growth Inhibition Assay | IC50=40.3662 μM | SANGER | |||
| C32 | Growth Inhibition Assay | IC50=40.4024 μM | SANGER | |||
| KU-19-19 | Growth Inhibition Assay | IC50=40.7683 μM | SANGER | |||
| KNS-62 | Growth Inhibition Assay | IC50=40.8381 μM | SANGER | |||
| FADU | Growth Inhibition Assay | IC50=41.2502 μM | SANGER | |||
| CAL-33 | Growth Inhibition Assay | IC50=42.6749 μM | SANGER | |||
| CHP-134 | Growth Inhibition Assay | IC50=42.8496 μM | SANGER | |||
| HDLM-2 | Growth Inhibition Assay | IC50=42.9084 μM | SANGER | |||
| NBsusSR | Growth Inhibition Assay | IC50=43.0725 μM | SANGER | |||
| SW954 | Growth Inhibition Assay | IC50=43.1053 μM | SANGER | |||
| HCC1806 | Growth Inhibition Assay | IC50=43.411 μM | SANGER | |||
| VMRC-RCZ | Growth Inhibition Assay | IC50=43.4586 μM | SANGER | |||
| A549 | Growth Inhibition Assay | IC50=43.931 μM | SANGER | |||
| NKM-1 | Growth Inhibition Assay | IC50=43.9558 μM | SANGER | |||
| DMS-273 | Growth Inhibition Assay | IC50=44.7567 μM | SANGER | |||
| TYK-nu | Growth Inhibition Assay | IC50=45.1234 μM | SANGER | |||
| KALS-1 | Growth Inhibition Assay | IC50=45.146 μM | SANGER | |||
| A101D | Growth Inhibition Assay | IC50=45.4456 μM | SANGER | |||
| G-361 | Growth Inhibition Assay | IC50=46.2138 μM | SANGER | |||
| KARPAS-299 | Growth Inhibition Assay | IC50=46.3516 μM | SANGER | |||
| RS4-11 | Growth Inhibition Assay | IC50=46.542 μM | SANGER | |||
| HT-1376 | Growth Inhibition Assay | IC50=46.7426 μM | SANGER | |||
| SK-N-AS | Growth Inhibition Assay | IC50=46.7822 μM | SANGER | |||
| MG-63 | Growth Inhibition Assay | IC50=46.9036 μM | SANGER | |||
| EPLC-272H | Growth Inhibition Assay | IC50=46.9503 μM | SANGER | |||
| BALL-1 | Growth Inhibition Assay | IC50=47.832 μM | SANGER | |||
| LCLC-97TM1 | Growth Inhibition Assay | IC50=48.202 μM | SANGER | |||
| HO-1-N-1 | Growth Inhibition Assay | IC50=48.9676 μM | SANGER | |||
| MFE-280 | Growth Inhibition Assay | IC50=49.4617 μM | SANGER | |||
| NCI-H526 | Growth Inhibition Assay | IC50=49.8163 μM | SANGER | |||
| D-566MG | Growth Inhibition Assay | IC50=49.9096 μM | SANGER | |||
| BB30-HNC | Growth Inhibition Assay | IC50=49.9498 μM | SANGER | |||
| SK-N-DZ | Growth Inhibition Assay | IC50=50.0481 μM | SANGER | |||
| HepG2 | Growth Inhibition Assay | 10 μM | 24 h | blocks SC-III3-induced S phase arrest | 25527123 | |
| HepG2 | Function Assay | 10 μM | 24 h | suppresses the phosphorylations of ATM on Ser1981, Chk1 on Ser345, Chk2 on Thr68, and Cdk2 on Tyr15 induced by SC-III3 | 25527123 | |
| KATO III | Growth Inhibition Assay | 2.5/5/7.5 μM | DMSO | enhances the toxicity of olaparib | 24841718 | |
| hTCEpi | Growth Inhibition Assay | 10 μM | DMSO | prevents the cytopathic effect of HSV-1 | 24370835 | |
| MCF10A | Growth Inhibition Assay | 10 μM | 24 h | DMSO | potentiates the cytotoxicity of GA | 24150595 |
| HL-60 | Function Assay | 10 μM | 0.5 h | DMSO | reduces phosphorylation of Chk2 | 23934411 |
| MCF-7 | Growth Inhibition Assay | 1-100μM | 24 h | FBS | inhibits the cell proliferation | 23185347 |
| HeLa | Growth Inhibition Assay | 1-100μM | 24 h | FBS | inhibits the cell proliferation | 23185347 |
| SH-SY5Y | Function Assay | 10 μM | 24 h | inhibits clioquinol-induced phosphorylation of p53 | 22627294 | |
| IMR-32 | Function Assay | 10 μM | 24 h | inhibits clioquinol-induced phosphorylation of p53 | 22627294 | |
| A549 | Function Assay | 10 μM | 1 h | suppresses Nano-Co-induced p53 accumulation | 22559321 | |
| T47D | Function Assay | 20 mM | 24 h | DMSO | prevents IR-induced degradation of IκBα | 21144805 |
| A29 MEF | Function Assay | 10 μM | 1h | blocks the phosphorylation of Akt at Ser473 | 20053781 | |
| MDA-MB-453 | Growth Inhibition Assay | 5-40 μM | 72 h | IC50 of 10 μM | 20053781 | |
| PC-3 | Growth Inhibition Assay | 5-40 μM | 72 h | IC50 of 10 μM | 20053781 | |
| U2OS | Function assay | Inhibition of ATM in human U2OS cells assessed as inhibition of p53 phosphorylation at Ser15 residue, IC50 = 0.25 μM. | 26632965 | |||
| MCF7 | Function assay | 1 hr | Inhibition of ATM kinase in human MCF7 cells after 1 hr by immunofluorescence assay, IC50 = 0.3 μM. | 26632965 | ||
| BJ | Function assay | 10 uM | 10 days | Suppression of senescence in human BJ cells assessed as increase in cell number at 10 uM after 10 days by senescence reversal assay | 16767085 | |
| BJ | Function assay | 10 uM | 10 days | Inhibition of ataxia telangiectasia-mutated in human BJ cells assessed as increase in cell number at 10 uM after 10 days by senescence reversal assay | 16767085 | |
| MCF7 | Function assay | 10 uM | 10 mins | Sensitization of infrared-induced DNA damage in human MCF7 cells assessed as reduction in colony formation at 10 uM pretreated for 10 mins followed by irradiation for 4 hrs measured after 10 days by crystal violet staining analysis | 26632965 | |
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SK-N-MC cells | 29435139 | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 39 mg/mL
(98.61 mM)
Water : Insoluble Ethanol : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 395.49 | Formula | C21H17NO3S2 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 587871-26-9 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | ATM Kinase Inhibitor | Smiles | C1COCCN1C2=CC(=O)C=C(O2)C3=C4C(=CC=C3)SC5=CC=CC=C5S4 | ||
Mechanism of Action
| Targets/IC50/Ki |
ATM
(Cell-free assay) 12.9 nM
|
|---|---|
| In vitro |
KU-55933 inhibits DNA-PK and PI3K with IC50 of 2.5 μM and 16.6 μM, respectively. Besides, this compound also prevents the activity of mTOR with IC50 of 9.3 μM. It is active at the cellular level in ablating a well-characterized ATM-dependent phosphorylation event. This chemical has a dose-dependent effect in inhibiting this ATM-dependent phosphorylation event with IC50 of 300 nM. KU-58050 does not prevent the ATM-dependent phosphorylation of p53 serine 15 until a dose of 30 μM. Addition of this compound has no appreciable effects on UV-induced phosphorylation of H2AX on serine 139, NBS1 on serine 343, CHK1 on serine 345, and SMC1 on serine 966. In stark contrast to the UV responses, it ablates the ionizing radiation-induced phosphorylation of these ATM substrates. This chemical sensitizes HeLa cells to a range of ionizing radiation doses. It inhibits the phosphorylation of Akt induced by growth factors in cancer cells. This compound suppresses the proliferation of cancer cells. Furthermore, suppression of ATM by this chemical improves survival, probably via prevention of downstream activation of TAp63α.
|
| Kinase Assay |
Purified enzyme assays
|
|
ATM for use in the in vitro assay is obtained from HeLa nuclear extract by immunoprecipitation with rabbit polyclonal antiserum raised to the COOH-terminal 400 amino acids of ATM in buffer containing 25 mM HEPES (pH 7.4), 2 mM MgCl2, 250 mM KCl, 500 μM EDTA, 100 μM Na3VO4, 10% v/v glycerol, and 0.1% v/v Igepal. ATM-antibody complexes are isolated from nuclear extract by incubating with protein A-Sepharose beads for 1 hour and then through centrifugation to recover the beads. In the well of a 96-well plate, ATM-containing Sepharose beads are incubated with 1 μg of substrate glutathione S-transferase–p53N66 (NH2-terminal 66 amino acids of p53 fused to glutathione S-transferase) in ATM assay buffer [25 mM HEPES (pH 7.4), 75 mM NaCl, 3 mM MgCl2, 2 mM MnCl2, 50 μM Na3VO4, 500 μM DTT, and 5% v/v glycerol] at 37 °C in the presence or absence of this compound. After 10 minutes with gentle shaking, ATP is added to a final concentration of 50 μM and the reaction continued at 37 °C for an additional 1 hour. The plate is centrifuged at 250 × g for 10 minutes (4 °C) to remove the ATM-containing beads, and the supernatant is removed and transferred to a white opaque 96-well plate and incubated at room temperature for 1.5 hours to allow glutathione S-transferase-p53N66 binding. This plate is then washed with PBS, blotted dry, and analyzed by a standard ELISA technique with a phospho-serine 15 p53 antibody. The detection of phosphorylated glutathione S-transferase-p53N66 substrate is performed in combination with a goat antimouse horseradish peroxidase-conjugated secondary antibody. Enhanced chemiluminescence solution is used to produce a signal and chemiluminescent detection is carried out.
|
|
| In vivo |
Suppression of ATM-dependent STAT3 activation by KU-55933 enhances TRAIL-mediated apoptosis through up-regulation of surface DR5 expression, whereas suppression of both STAT3 and NF-κB appeares to be involved in down-regulation of cFLIP accompanied by an additional increase in apoptotic levels. This compound affects TRAIL-mediated apoptosis more strongly than the JAK2 inhibitor, AG490, or overexpression of STAT3β.
|
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | p-AKT(Ser473) / p-AKT(Thr308) PARP / Cleaved PARP / Caspase-3 / Cleaved caspase-3 ATM-S1981 / ATM / p-p53(S15) / p53 p21 / p27 / p53 |
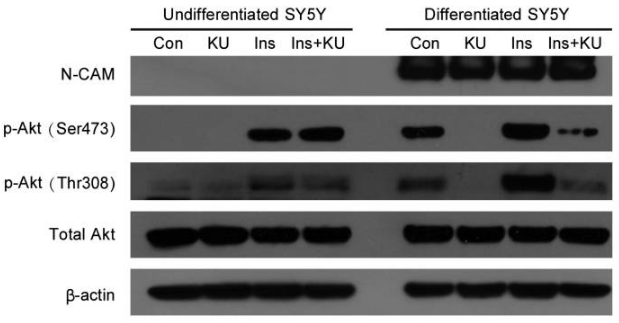
|
22739265 |
| Immunofluorescence | p-S824 KAP1 / ZEBRA |
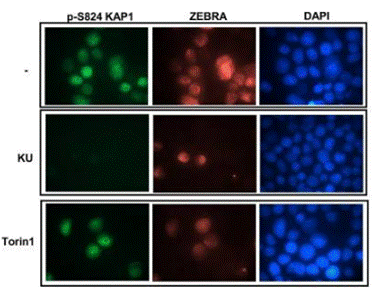
|
28249048 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































