research use only
AZ20 ATM/ATR inhibitor
Cat.No.S7050
Chemical Structure
Molecular Weight: 412.51
Quality Control
| Related Targets | PI3K Akt mTOR GSK-3 DNA-PK AMPK PDPK1 PTEN PP2A PDK |
|---|---|
| Other ATM/ATR Inhibitors | Ceralasertib (AZD6738) AZD1390 Berzosertib (VE-822) Lartesertib (M4076) Camonsertib (RP-3500) KU-60019 KU-55933 VE-821 AZD0156 Mirin |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| HT29 | Function assay | 1 hr | Inhibition of ATR-mediated CHK1 phosphorylation at serine 345 in human HT29 cells after 1 hr in presence of 4-nitroquinoline 1-oxide, IC50=0.05μM | 23394205 | ||
| LoVo | Growth inhibition assay | 72 hrs | Growth inhibition of human LoVo cells after 72 hrs by MTS assay, GI50=0.2μM | 23394205 | ||
| MDA-MB-468 | Function assay | Inhibition of mTOR-mediated AKT phosphorylation at serine 473 in human MDA-MB-468 cells, IC50=2.4μM | 23394205 | |||
| LoVo | Antitumor assay | 50 mg/kg | 13 days | Antitumor activity against human LoVo cells xenografted in Swiss nu/nu mouse assessed as tumor growth inhibition at 50 mg/kg, po qd for 13 days relative to control | 23394205 | |
| LoVo | Antitumor assay | 25 mg/kg | 13 days | Antitumor activity against human LoVo cells xenografted in Swiss nu/nu mouse assessed as tumor growth inhibition at 25 mg/kg, po bid for 13 days relative to control | 23394205 | |
| HT29 | Function assay | 60 mins | Inhibition of ATR in human HT29 cells after 60 mins by Hoechst 33258 staining-based assay, IC50=0.061μM | 30346772 | ||
| LoVo | Cytotoxicity assay | 72 hrs | Cytotoxicity against human LoVo cells after 72 hrs by MTS assay, GI50=0.2μM | 30346772 | ||
| MDA-MB-468 | Function assay | Inhibition of mTOR in human MDA-MB-468 cells assessed as decrease in 70S6K S235/236 phosphorylation, IC50=0.72μM | 30346772 | |||
| HT-29 | Cytotoxicity assay | 72 hrs | Cytotoxicity against human HT-29 cells after 72 hrs by MTS assay, GI50=0.97μM | 30346772 | ||
| LoVo | Function assay | 25 mg/kg | 8 hrs | Plasma concentration in Swiss nu/nu mouse xenografted with human LoVo cells at 25 mg/kg, po bid after 8 hrs, Cp=1.8μM | 30346772 | |
| MDA-MB-468 | Function assay | Inhibition of mTOR in human MDA-MB-468 cells assessed as decrease in AKT phosphorylation at S473 residue, IC50=2.4μM | 30346772 | |||
| LoVo | Function assay | 50 mg/kg | 8 hrs | Plasma concentration in Swiss nu/nu mouse xenografted with human LoVo cells at 50 mg/kg, po qd after 8 hrs, Cp=3.5μM | 30346772 | |
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 83 mg/mL
(201.2 mM)
Ethanol : 3 mg/mL Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 412.51 | Formula | C21H24N4O3S |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 1233339-22-4 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N/A | Smiles | CC1COCCN1C2=NC(=NC(=C2)C3(CC3)S(=O)(=O)C)C4=C5C=CNC5=CC=C4 | ||
Mechanism of Action
| Features |
ATR-selective inhibitor with high permeability and good stability.
|
|---|---|
| Targets/IC50/Ki |
ATR
(Cell-free assay) 5 nM
mTOR
(Cell-free assay) 38 nM
|
| In vitro |
AZ20 shows good selectivity against all of the PI3K isoforms together with ATM and DNA-PK. In vitro, this compound decreases pChk1 Ser345, pChk1 Ser317 and pChk1 Ser296 levels in a concentration-dependent manner. Prolonged exposure with this chemical increases γH2AX pan-nuclear staining, indicative of replication stress. This is associated with S-phase arrest and increase in phospho-histone H3. It induces growth inhibition and cell death in vitro and its profile of activity is distinct from other cytotoxic agents. The cytotoxic effect of this agent can be increased in combination with the selective ATM inhibitor KU-60019.
|
| In vivo |
Female nude mice bearing LoVo tumors are treated with AZ20 orally at a dose of 25 mg/kg twice daily or 50 mg/kg once daily for 13 days, led to significant tumor growth inhibition. This is associated with a persistent elevation of γH2AX pan-nuclear staining in xenograft tissue, but a transient increase in mouse bone marrow at therapeutic doses, suggesting a favourable therapeutic index.
This compound is assessed for drug−drug interaction (DDI) potential specifically from inhibition of cytochrome P450 enzymes. It is found to inhibit the cytochrome 3A4-mediated metabolism of midazolam by 50% at 10 μM. This chemical has respectable bioavailability in a low dose rat PK study.
|
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Growth inhibition assay | IC50 |
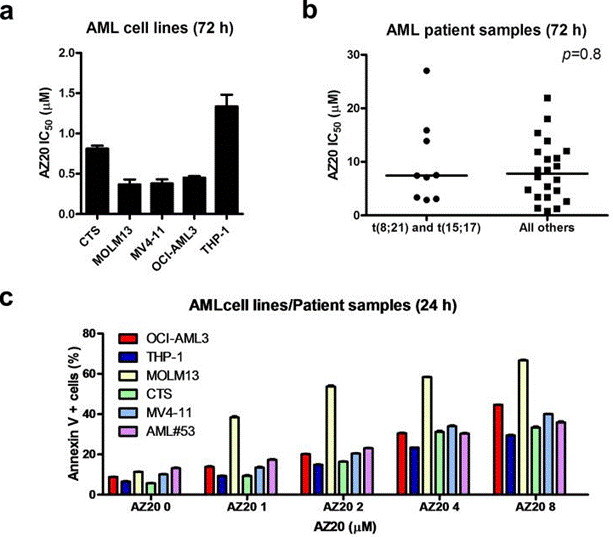
|
28176818 |
| Western blot | p-CDK1 / CDK1 / p-CDK2 / CDK2 γH2AX / RRM1 / RRM2 |
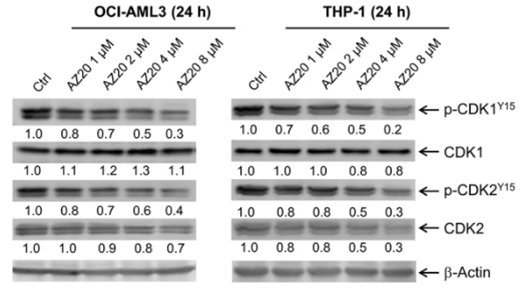
|
28176818 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.
Frequently Asked Questions
Question 1:
If I want to completely block the kinase activity from the in vitro cell lines, how much concentration of it should I use?
Answer:
IC50 5nM was quoted from a previous publication in which the author tested its IC50 in a cell-free assay. In cell culture, many factors, such as membrane permeability and target protein concentration, may affect the efficiency. Each cell line responds to the same compound differently, and it is very difficult to predict the optimized concentration simply based on cell-free data. In cell culture experiments, the required concentration is usually higher. We recommend that you perform a pilot experiment and test different concentrations (50nM to 500μM) to get the optimized condition.






































