research use only
UNC1215 DNA/RNA Synthesis antagonist
Cat.No.S7088
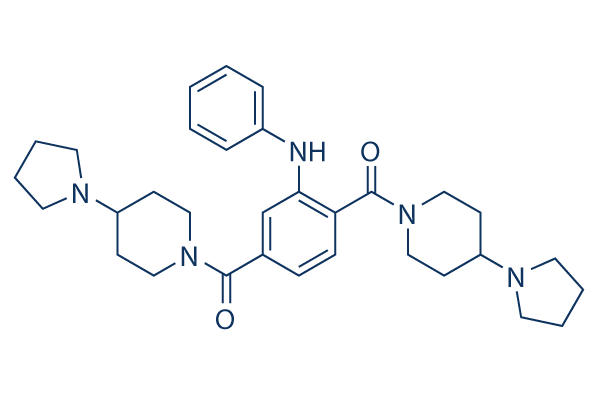
Chemical Structure
Molecular Weight: 529.72
Quality Control
| Related Targets | HDAC JAK BET Histone Methyltransferase PKC PARP HIF PRMT EZH2 AMPK |
|---|---|
| Other DNA/RNA Synthesis Inhibitors | CX-5461 (Pidnarulex) SCR7 Favipiravir (T-705) EED226 RK-33 BMH-21 Carmofur Triapine (3-AP) YK-4-279 Halofuginone |
Solubility
|
In vitro |
DMSO
: 100 mg/mL
(188.77 mM)
Ethanol : 100 mg/mL Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 529.72 | Formula | C32H43N5O2 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 1415800-43-9 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N/A | Smiles | C1CCN(C1)C2CCN(CC2)C(=O)C3=CC(=C(C=C3)C(=O)N4CCC(CC4)N5CCCC5)NC6=CC=CC=C6 | ||
Mechanism of Action
| Features |
The first chemical probe for a Kme-binding protein.
|
|---|---|
| Targets/IC50/Ki |
L3MBTL3
40 nM
L3MBTL3
120 nM(Kd)
L3MBTL3- D274A
3.5 μM
|
| In vitro |
UNC1215 binds L3MBTL3, competitively displacing mono- or dimethyllysine-containing peptides. This probe is greater than 50-fold selective versus other members of the human MBT family. This compound has about a 75-fold selectivity for L3MBTL3 over L3MBTL1. It shows no activity at concentrations up to 30 μM against the tandem Tudor domain of UHRF1, the chromodomain of CBX7 or the PHD domain of JARID1A. X-ray crystallography identifies a unique 2:2 polyvalent mode of interaction between this chemical and L3MBTL3. In cells, it is nontoxic and directly binds L3MBTL3 via the Kme-binding pocket of the MBT domains. This compound increases the cellular mobility of GFP-L3MBTL3 fusion proteins, and point mutants that disrupt the Kme-binding function of GFP-L3MBTL3 phenocopy the effects of this chemical on localization. It is used to reveal a new Kme-dependent interaction of L3MBTL3 with BCLAF1, a protein implicated in DNA damage repair and apoptosis.
|
| Kinase Assay |
AlphaScreen assay
|
|
Compound plates (1 μL at 10 or 30 mM highest concentration) are diluted in 1×assay buffer (20 mM Tris pH 8.0, 25 mM NaCl, 2 mM DTT and 0.05% Tween-20) over 2 steps using a Multimek robotic pipettor and 1 μL is spotted into the wells of 384-well assay Proxiplates. To these plates 9 μL of protein- peptide mix in 1× assay buffer is added by Multidrop and incubated for 30 min at room temperature. At this point 2 μL of streptavidin-conjugate donor and nickel-chelate acceptor beads (45 μg/mL in 1× assay buffer) are added, the plates are allowed to incubate for an additional 30 min in the dark at room temperature. After incubation the plates are read on EnVision mulilabel reader equipped with HTS alpha screen laser. The screens reported are performed up to 10 or 30 μM, and therefore it should be noted that those compounds referred to as inactive are indeed inactive only within the concentration range tested. PHF23 and JARID1A are GST tagged and consequently for these assays GST-acceptor beads are used. It should be noted that any positive binding curves for L3MBTL4 that are generated yielded curves with very shallow slopes, suggesting a nonspecific interaction. The data for the IC50 values is calculated from replicate runs in that the datapoints for each compound concentration are averaged and plotted using 4-parameters curve fitting.
|
References |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































