research use only
SB203580 (Adezmapimod) p38 MAPK inhibitor
Cat.No.S1076
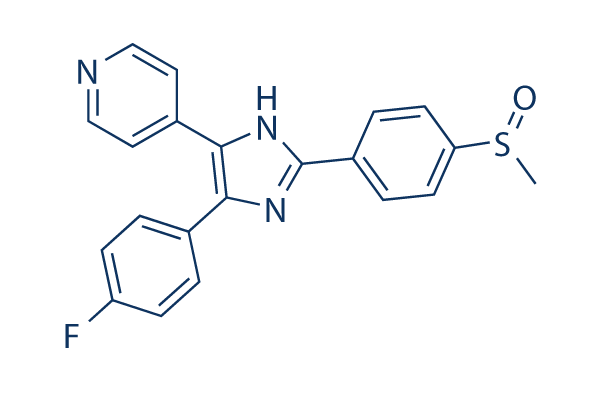
Chemical Structure
Molecular Weight: 377.43
Quality Control
| Related Targets | ERK Raf JNK MEK Ras KRas S6 Kinase MAP4K TAK1 Mixed Lineage Kinase |
|---|---|
| Other p38 MAPK Inhibitors | SB202190 PH-797804 Doramapimod (BIRB 796) Ralimetinib (LY2228820) dimesylate VX-702 Losmapimod SB239063 Asiatic Acid Neflamapimod (VX-745) BMS-582949 |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| PC12 | Kinase Assay | 10 μM | 15 min | Inhibits p38 MAP kinase with IC50 of 0.6 μM | 7750577 | |
| THP-1 | Function Assay | Inhibits LPS-induced TNFalpha production with IC50 of 0.16 μM | 18325768 | |||
| PBMC | Function Assay | 15 min | DMSO | Inhibits the release of interleukin-1-beta with IC50 of 0.037 μM | 12361396 | |
| SW1353 | Function Assay | 1 h | DMSO | Inhibits IL-6 production with IC50 of 0.05 μM | 11140741 | |
| Hela | Function Assay | DMSO | Inhibits Tat-induced HIV1 LTR transactivation in human HeLa cells with IC50 of 0.1 μM | 18926711 | ||
| PC12 | Function Assay | 10 μM | DMSO | Activates Nrf2/ARE in rat PC12 cells assessed as HO-1 protein induction | 21345685 | |
| RAW 264.7 | Cytotoxic Assay | 12.5 μM | 12h | DMSO | Blocks lethal toxin-mediated cytotoxicity | 17485504 |
| HCA2 | Function Assay | 2.5 μM | Inhibits p38 mediated MK2 activation | 17964780 | ||
| U937 | Function Assay | Inhibits JAK-mediated interferon-gamma/anisomycin-induced Stat1 phosphorylation | 18157122 | |||
| U937 | Function Assay | Inhibits JAK-mediated interferon-gamma/anisomycin-induced p38 phosphorylation | 18157122 | |||
| hESCs | Growth Inhibition Assay | 5 μM | DMSO | Induces cardiomyogenic activity assessed as cell growth | 23602399 | |
| RAW264.7 | Function Assay | 10 μM | Induces antiinflammatory activity by inhibiting IL-1β release | 23791078 | ||
| RAW264.7 | Function Assay | 10 μM | Induces antiinflammatory activity by inhibiting iNOS release | 23791078 | ||
| RAW264.7 | Function Assay | 10 μM | Induces antiinflammatory activity by inhibiting NO release | 23791078 | ||
| RAW264.7 | Function Assay | 10 μM | Inhibits LPS-induced p38 MAPK phosphorylation | 24016057 | ||
| IEC-18 | Function Assay | 10 μM | Inhibits LPS-induced p38 MAPK phosphorylation | 23758110 | ||
| Sf9 | Function assay | 30 mins | Inhibition of full-length FLAG-6His-TEV tagged human RIP2 expressed in Sf9 cells assessed as reduction in autophosphorylation pre-incubated for 30 mins followed by incubation with ATP for 2 hrs by ADP-Glo assay, IC50 = 0.01 μM. | 26455654 | ||
| sf21 | Function assay | Binding affinity to wild type human biotin labelled p38 alpha (9 to 352 residues) expressed in sf21 insect cells SPR analysis, Kd = 0.0217 μM. | 28834431 | |||
| PBMC | Function assay | Inhibition of TNF-alpha production by lipopolysaccharide-stimulated human peripheral blood mononuclear cells, IC50 = 0.025 μM. | 9873730 | |||
| PBMC | Function assay | Inhibition of LPS-stimulated TH+NF-alphs production by human peripheral blood mononuclear cells (PBMCs), IC50 = 0.025 μM. | 9784093 | |||
| PBMC | Function assay | Inhibition of p38 MAP kinase related TNF-alpha release from peripheral blood mononuclear cells (PBMC), IC50 = 0.037 μM. | 12061876 | |||
| PBMC | Function assay | Inhibition of LPS-induced IL1-beta production in human peripheral blood mononuclear cells(PBMCs), IC50 = 0.048 μM. | 9784093 | |||
| SW1353 | Function assay | Compound was evaluated for its ability to inhibit IL-6 production in SW1353 cells treated with cytokines IL-1 and TNF, IC50 = 0.05 μM. | 10999467 | |||
| SW1353 | Function assay | Inhibition of TNF and IL-1 induced IL-6 production in human chondro-sarcoma SW 1353 cells, IC50 = 0.05 μM. | 10999468 | |||
| THP1 | Function assay | Inhibition of LPS-stimulated IL1 production in human THP1 cells, IC50 = 0.05 μM. | 18077363 | |||
| THP1 | Function assay | Inhibition of LPS-stimulated TNFalpha production in human THP1 cells, IC50 = 0.05 μM. | 18077363 | |||
| THP1 | Function assay | Inhibition of LPS-induced tumor necrosis factor-alpha (TNF-alpha) production in THP-1 cells, EC50 = 0.06 μM. | 12086485 | |||
| THP1 | Function assay | Inhibition of tumor necrosis factor alpha release by THP-1 cells, IC50 = 0.07 μM. | 15658855 | |||
| monocytic cell | Function assay | Inhibitory activity against TNF-alpha production using lipopolysaccharide stimulated human monocytic cells, IC50 = 0.072 μM. | 12729637 | |||
| THP1 | Function assay | Inhibitory activity against LPS-stimulated TNF-alpha production in human monocytic cells (THP-1), IC50 = 0.072 μM. | 15139749 | |||
| THP-1 | Function assay | Inhibitory activity against lipopolysaccharide stimulated TNF-alpha production in THP-1 cells (human monocytic cells), IC50 = 0.072 μM. | 15837310 | |||
| THP1 | Function assay | Inhibition of LPS-stimulated TNFalpha production in THP1 cells, IC50 = 0.072 μM. | 16750367 | |||
| PBMC | Function assay | Inhibition of staphylococcal enterotoxin B(SEB) stimulated tumor necrosis factor alpha (TNF-alpha) production by human peripheral blood mononuclear cells(PBMCs), IC50 = 0.16 μM. | 9784093 | |||
| PBMC | Function assay | Concentration required to inhibit lipopolysaccharide induced TNF-alpha release was determined in human peripheral blood mononuclear cells, IC50 = 0.19 μM. | 15454231 | |||
| Sf9 | Function assay | Inhibition of full-length FLAG-6His-TEV tagged human RIP2 expressed in Sf9 cells by fluorescent polarization assay, IC50 = 0.2 μM. | 26455654 | |||
| HEK293F | Function assay | Inhibition of sodium arsenate activated N-terminal GST-tagged Brugia malayi MPK1 expressed in HEK293F cells using FAM-p38tide as substrate by IMAP assay, IC50 = 0.22 μM. | 29541362 | |||
| HEK-293 | Function assay | Inhibition of RIP2K in MDP-stimulated HEK-293 cells over-expressing NOD2 assessed as IL8 secretion, IC50 = 0.25 μM. | 27109867 | |||
| whole blood cell | Function assay | 0.01 to 100 uM | Inhibition of p38-related IL1-beta release by whole blood cells at 10 e-4 to 10 e-8 M, IC50 = 0.35 μM. | 12852754 | ||
| PBMC | Function assay | Inhibition of p38 MAP kinase related TNF-alpha release from peripheral blood mononuclear cells (PBMC), IC50 = 0.59 μM. | 12061876 | |||
| PBMC | Function assay | Inhibition of the release of tumor necrosis factor alpha from peripheral blood mononuclear cells, IC50 = 0.59 μM. | 12361396 | |||
| PBMC | Function assay | Inhibitory activity against TNF-alpha release in PBM cells, IC50 = 0.59 μM. | 12852754 | |||
| whole blood cell | Function assay | 0.01 to 100 uM | Inhibition of p38-related TNF-alpha release by whole blood cells at 10 e-4 to 10 e-8 M, IC50 = 0.94 μM. | 12852754 | ||
| BMDC | Antiinflammatory assay | 1 hr | Antiinflammatory activity in C57BL/6 mouse BMDC cells assessed as inhibition of LPS-induced IL-6 production preincubated for 1 hr prior to LPS-stimulation measured after 18 hrs by ELISA, IC50 = 3.5 μM. | 24047259 | ||
| BMDC | Antiinflammatory assay | 1 hr | Antiinflammatory activity in C57BL/6 mouse BMDC cells assessed as inhibition of LPS-induced IL-12 p40 production preincubated for 1 hr prior to LPS-stimulation measured after 18 hrs by ELISA, IC50 = 5 μM. | 24047259 | ||
| HEK-293 | Function assay | Inhibition of RIP2K in Pam2CSK4-stimulated HEK-293 cells over-expressing TLR2 assessed as IL8 secretion, IC50 = 6.31 μM. | 27109867 | |||
| BMDC | Antiinflammatory assay | 1 hr | Antiinflammatory activity in C57BL/6 mouse BMDC cells assessed as inhibition of LPS-induced TNFalpha production preincubated for 1 hr prior to LPS-stimulation measured after 18 hrs by ELISA, IC50 = 7.5 μM. | 24047259 | ||
| macrophage-like cell | Antiinflammatory assay | 30 mins | Antiinflammatory activity in human HL-60 derived macrophage-like cells assessed as inhibition of LPS-induced TNFalpha production treated 30 mins before LPS stimulation measured after 24 hrs by ELISA, IC50 = 8.6 μM. | 23811089 | ||
| WS | Function assay | 2.5 uM | Inhibition of anisomycin-induced P38-alpha activation in human human TERT-immortalised WS cells assessed as HSP27 phosphorylation at 2.5 uM | 17659871 | ||
| WS | Function assay | Inhibition of MK2 mediated HSP27 phosphorylation in immortalised WS cells assessed as reduction in F-actin stress fibres | 17964780 | |||
| WS | Function assay | Inhibition of p38 mediated MK2 activation in WS cells | 17964780 | |||
| A673 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for A673 cells | 29435139 | |||
| Saos-2 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for Saos-2 cells | 29435139 | |||
| BT-37 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for BT-37 cells | 29435139 | |||
| RD | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for RD cells | 29435139 | |||
| BT-12 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for BT-12 cells | 29435139 | |||
| OHS-50 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for OHS-50 cells | 29435139 | |||
| SJ-GBM2 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SJ-GBM2 cells | 29435139 | |||
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SK-N-MC cells | 29435139 | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 38 mg/mL
(100.68 mM)
Water : Insoluble Ethanol : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 377.43 | Formula | C21H16FN3OS |
Storage (From the date of receipt) | 3 years -20°C(in the dark) powder |
|---|---|---|---|---|---|
| CAS No. | 152121-47-6 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | RWJ 64809, PB 203580 | Smiles | CS(=O)C1=CC=C(C=C1)C2=NC(=C(N2)C3=CC=NC=C3)C4=CC=C(C=C4)F | ||
Mechanism of Action
| Features |
First reported p38 inhibitor.
|
|---|---|
| Targets/IC50/Ki |
p38 MAPK
(THP-1 cells) 0.3 μM-0.5 μM
PKB
(THP-1 cells) 3 μM-5 μM
|
| In vitro |
Adezmapimod (SB203580) inhibits the IL-2-induced proliferation of primary human T cells, murine CT6 T cells, or BAF F7 B cells with an IC50 of 3–5 μm. It also inhibits IL-2-induced p70S6 kinase activation, although the concentration required is slightly higher with an IC50 above 10 μm. This compound inhibits the activity of PDK1 in a dose-dependent manner with an IC50 in the 3–10 μm range. It inhibits p38-MAPK stimulation of MAPKAPK2 with an IC50 of approximately 0.07 μM, whereas total SAPK/JNK activity is inhibited with an IC50 of 3–10 μM. At higher concentrations, it activates the ERK pathway, which subsequently enhances NF-κB transcriptional activity. It also induces autophagy in human hepatocellular carcinoma (HCC) cells. |
| Kinase Assay |
Cellular receptor kinase phosphorylation assay
|
|
4 μg of sheep anti-PKBα is immobilized on 25 μL of protein G-Sepharose overnight (or 1.5 hours) and washed in Buffer A (50 mm Tris, pH 7.5, 1 mm EDTA, 1 mm EGTA, 0.5 mm Na3VO4, 0.1% β-mercaptoethanol, 1% Triton X-100, 50 mm sodium fluoride, 5 mm sodium pyrophosphate, 0.1 mm phenylmethylsulfonyl fluoride, 1 μg/mL pepstatin, leupeptin, and 1 μm microcystin). The immobilized anti-PKB is then incubated with 0.5 ml of lysate (from 5 × 106 cells) for 1.5 hours and washed three times in 0.5 mL of Buffer A supplemented with 0.5 m NaCl, two times in 0.5 mL of Buffer B (50 mm Tris-HCl, pH 7.5, 0.03% (w/v) Brij-35, 0.1 mm EGTA, and 0.1% β-mercaptoethanol), and twice with 100 μl of assay dilution buffer; 5× assay dilution buffer is 100 mm MOPS, pH 7.2, 125 mm β-glycerophosphate, 25 mm EGTA, 5 mm sodium orthovanadate, 5 mm DTT. To the PKB enzyme immune complex is added 10 μL of assay dilution buffer, 40 μm protein kinase A inhibitor peptide, 100 μm PKB-specific substrate peptide, and 10 μCi of [γ-32P]ATP, all made up in assay dilution buffer. The reaction is incubated for 20 minutes at room temperature with shaking, then samples are pulse spun, and 40 μL of reaction volume are removed into another tube to which is added 20 μL of 40% trichloroacetic acid to stop the reaction. This is mixed and incubated for 5 minutes at room temperature, and 40 μL is transferred onto P81 phosphocellulose paper and allowed to bind for 30 seconds. The P81 piece is washed three times in 0.75% phosphoric acid then in acetone at room temperature. γ-32P incorporation is then measured by scintillation counting.
|
|
| In vivo |
Adezmapimod (SB203580) protects pig myocardium against ischemic injury in an in vivo model. It is also effective in preventing and treating the disease in MRL/lpr mice model of Systemic lupus erythematosus (SLE). |
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
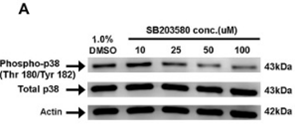
| Western blot | phospho-p38 / p38 p-ASK1 / ASK1 / P-JNK / p-MKK4 / p-ERK phospho-MK2(T334) / phospho-HSP27(S82) / phospho-Akt(T308) / phospho-Akt(S473) |
|
25747578 |
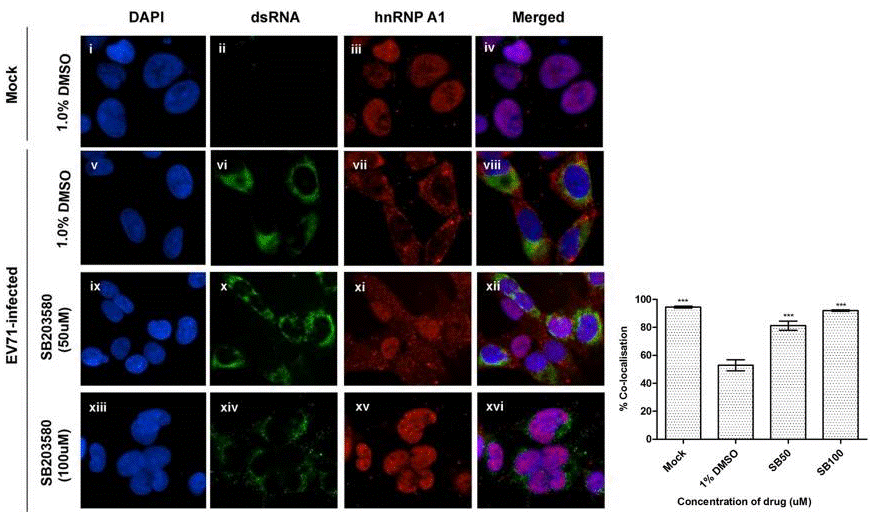
| Immunofluorescence | dsRNA / hnRNP A1 |
|
25747578 |
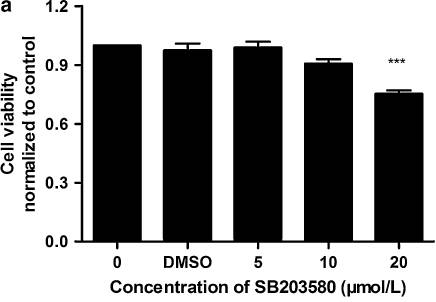
| Growth inhibition assay | Cell viability |
|
23001390 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































