research use only
Ralimetinib (LY2228820) dimesylate p38 MAPK inhibitor
Cat.No.S1494
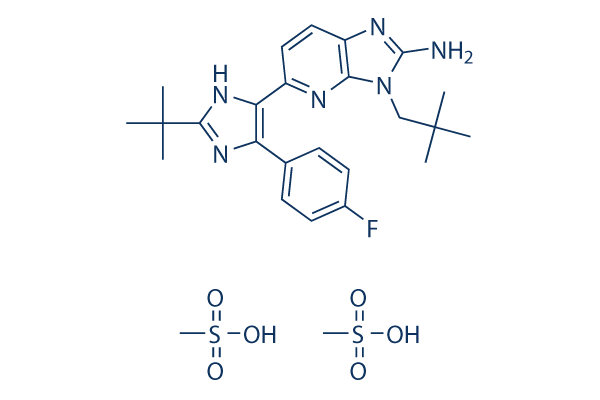
Chemical Structure
Molecular Weight: 612.74
Quality Control
| Related Targets | ERK Raf JNK MEK Ras KRas S6 Kinase MAP4K TAK1 Mixed Lineage Kinase |
|---|---|
| Other p38 MAPK Inhibitors | SB202190 SB203580 (Adezmapimod) PH-797804 Doramapimod (BIRB 796) VX-702 Losmapimod SB239063 Neflamapimod (VX-745) BMS-582949 Asiatic Acid |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| RPMI-8226 | Kinase assay | ~800 nM | DMSO | inhibits phosphorylation of HSP27 | 18397345 | |
| U266 | Kinase assay | ~800 nM | DMSO | inhibits phosphorylation of HSP27 | 18397345 | |
| MM.1S | Kinase assay | ~800 nM | DMSO | inhibits phosphorylation of HSP27 | 18397345 | |
| RPMI-Dox40 | Kinase assay | ~800 nM | DMSO | inhibits phosphorylation of HSP27 | 18397345 | |
| RPMI-LR5 | Kinase assay | ~800 nM | DMSO | inhibits phosphorylation of HSP27 | 18397345 | |
| INA-6 | Kinase assay | ~800 nM | DMSO | inhibits phosphorylation of HSP27 | 18397345 | |
| RPMI-8226 | Cytoxicity assay | ~1000 nM | DMSO | no significant cytotoxicity | 18397345 | |
| U266 | Cytoxicity assay | ~1000 nM | DMSO | no significant cytotoxicity | 18397345 | |
| MM.1S | Cytoxicity assay | ~1000 nM | DMSO | no significant cytotoxicity | 18397345 | |
| RPMI-Dox40 | Cytoxicity assay | ~1000 nM | DMSO | no significant cytotoxicity | 18397345 | |
| RPMI-LR5 | Cytoxicity assay | ~1000 nM | DMSO | no significant cytotoxicity | 18397345 | |
| INA-6 | Cytoxicity assay | ~1000 nM | DMSO | no significant cytotoxicity | 18397345 | |
| CD14+ | Function assay | ~800 nM | DMSO | inhibits osteoclastogenesis from CD14 positive cells | 18397345 | |
| U-87-MG | Function assay | 1 μM | DMSO | reduces tumor-driven cord formation | 23335506 | |
| MDA-MB-231 | Function assay | 1 μM | DMSO | reduces tumor-driven cord formation | 23335506 | |
| A-2780 | Function assay | 1 μM | DMSO | reduces tumor-driven cord formation | 23335506 | |
| SK-OV-3 | Function assay | 1 μM | DMSO | reduces tumor-driven cord formation | 23335506 | |
| LXFA-629 | Function assay | 1 μM | DMSO | reduces tumor-driven cord formation | 23335506 | |
| NCI-H1650 | Function assay | 1 μM | DMSO | reduces tumor-driven cord formation | 23335506 | |
| PC-3 | Function assay | 1 μM | DMSO | reduces tumor-driven cord formation | 23335506 | |
| RAW264.7 | Function assay | ~20 μM | DMSO | inhibits Anisomycin-stimulated MK2 phosphorylation with IC50 of 35.3 nM | 24356814 | |
| mouse peritoneal macrophages | Function assay | ~20 μM | DMSO | LPS/IFN-γ–stimulated TNF-α production with IC50 of 6.3 nM | 24356814 | |
| A549 | Function assay | ~20 μM | DMSO | inhibits LPS-induced CXCL8 production with IC50 of 144.9 nM | 24356814 | |
| MDA-231 | Function assay | ~10 μM | suppresses DKK-1 expression | 26407843 | ||
| MCF-7 | Function assay | ~10 μM | suppresses DKK-1 expression | 26407843 | ||
| MDA-435 | Function assay | ~10 μM | suppresses DKK-1 expression | 26407843 | ||
| PC3 | Function assay | ~10 μM | DMSO | suppresses DKK-1 expression | 26913608 | |
| TC32 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for TC32 cells | 29435139 | |||
| A673 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for A673 cells | 29435139 | |||
| DAOY | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for DAOY cells | 29435139 | |||
| BT-37 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for BT-37 cells | 29435139 | |||
| RD | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for RD cells | 29435139 | |||
| BT-12 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for BT-12 cells | 29435139 | |||
| NB1643 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for NB1643 cells | 29435139 | |||
| OHS-50 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for OHS-50 cells | 29435139 | |||
| Rh41 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for Rh41 cells | 29435139 | |||
| Rh30 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for Rh30 cells | 29435139 | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
Water : 123 mg/mL
DMSO
: 35 mg/mL
(57.12 mM)
Ethanol : 3 mg/mL |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 612.74 | Formula | C24H29FN6.2CH4O3S |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 862507-23-1 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N/A | Smiles | CC(C)(C)CN1C2=C(C=CC(=N2)C3=C(N=C(N3)C(C)(C)C)C4=CC=C(C=C4)F)N=C1N.CS(=O)(=O)O.CS(=O)(=O)O | ||
Mechanism of Action
| Targets/IC50/Ki |
p38α
(Cell-free assay) 7 nM
|
|---|---|
| In vitro |
LY2228820 inhibits p38α, as well as the level of phosphoMAPKAPK-2 (pMK2) in RAW 264.7 cells, with IC50 values of 7 nM and 34.3 nM, respectively. Furthermore, LY2228820 inhibits lipopolysaccharide (LPS)-induced TNFα formation in murine peritoneal macrophages, with IC50 of 5.2 nM. In multiple myeloma (MM) cells, including INA6, RPMI-8226, U266, and RPMI-Dox40, LY2228820 (200 nM–800 nM) significantly blocks p38MAPK signaling, as revealed by its inhibition on phosphorylation of HSP27, a downstream target of p38MAPK, without affecting the expression level of HSP27. LY2228820 (200 nM–400 nM) enhances cytotoxicity and apoptosis, but LY2228820 alone doesn't inhibit the growth of MM.1S cells. LY2228820 (200 nM–800 nM) also inhibits secretion of IL-6 and MIP-1α in long-term BM stromal cells (LT-BMSCs), BM mononuclear cells (BMMNCs), peripheral blood (PB) CD138+, CD138− or PB CD14+ cells. LY2228820 (400 nM–800 nM) also blocks osteoclastogenesis from CD14+ cells. |
| Kinase Assay |
Inhibition of p38α
|
|
Inhibition of p38α is determined using recombinant human p38α in a standard filter binding protocol using ATP[γ-33P] and EGFR 21-mer peptide as substrate. Functional inhibition of TNFα in murine peritoneal macrophages is determined using LPS stimulation in the presence of LY2228820. To assess p38α activity in cells more directly, RAW 264.7 cells are treated with LY2228820 and then stimulated with anisomycin. The level of p38α activity is detected using a phosphoMAPKAPK-2 (pMK2) (Thr 334) antibody which reacts with a residue specifically phosphorylated by p38α.
|
|
| In vivo |
In LPS-induced mice, LY2228820 effectively inhibits the formation of TNFα with a threshold minimum 50% effective dose (TMED50) less than 1 mg/kg. In a rat model of collagen-inducedarthritis (CIA), LY2228820 displays potent effects on paw swelling, bone erosion, and cartilage destruction, with a threshold minimum 50% effective dose (TMED50)of 1.5 mg/kg. |
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | p-p38 / p38α / p38β / p-MK2 / MK2 / p-HSP27 / HSP27 p-S6K |
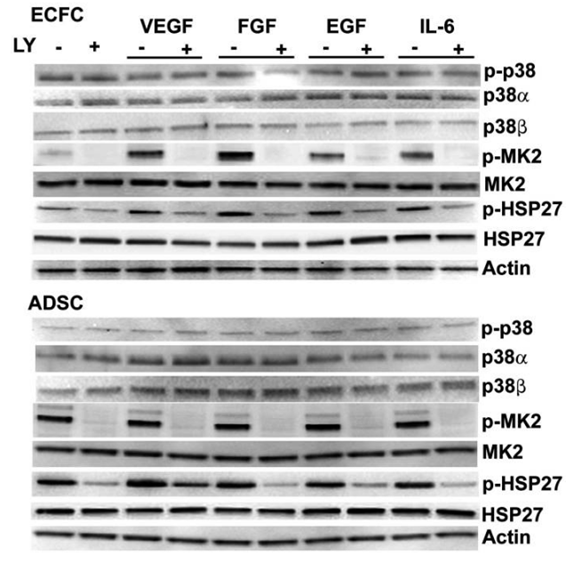
|
23335506 |
Clinical Trial Information
(data from https://clinicaltrials.gov, updated on 2024-05-22)
| NCT Number | Recruitment | Conditions | Sponsor/Collaborators | Start Date | Phases |
|---|---|---|---|---|---|
| NCT02860780 | Completed | Advanced Cancer|Metastatic Cancer|Colorectal Cancer|Non-small Cell Lung Cancer |
Eli Lilly and Company |
August 10 2016 | Phase 1 |
| NCT02364206 | Completed | Adult Glioblastoma |
Centre Jean Perrin|National Cancer Institute France|ARC Foundation for Cancer Research |
June 8 2015 | Phase 1|Phase 2 |
| NCT02322853 | Terminated | Postmenopausal|Metastatic Breast Cancer |
Centre Francois Baclesse|National Cancer Institute France|ARC Foundation for Cancer Research |
January 2015 | Phase 2 |
| NCT01393990 | Completed | Advanced Cancer |
Eli Lilly and Company |
September 4 2008 | Phase 1 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































