research use only
Purmorphamine Smoothened Receptor Agonist
Cat.No.S3042
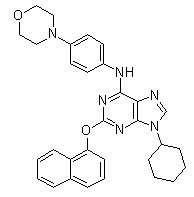
Chemical Structure
Molecular Weight: 520.62
Quality Control
| Related Targets | JAK TGF-beta/Smad Wnt/beta-catenin ERK GSK-3 ROCK PKA Secretase STAT Casein Kinase |
|---|---|
| Other Hedgehog/Smoothened Inhibitors | SAG (Smoothened Agonist) Hydrochloride Cyclopamine (11-Deoxojervine) GANT 61 (NSC 136476) SAG (Smoothened Agonist) SANT-1 HPI-4 (Ciliobrevin A) BMS-833923 Taladegib (LY2940680) Ciliobrevin D Jervine |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| C3H10T1/2 | Function assay | 6 days | Activity at Smo in mouse C3H10T1/2 cells assessed as induction of cell differentiation into osteoblast incubated for 6 days by alkaline phosphatase assay, EC50 = 0.8 μM. | 27429255 | ||
| Shh Light2 | Function assay | 30 hrs | Activation of Shh in mouse Shh Light2 cells after 30 hrs by luciferase reporter gene assay, EC50 = 1 μM. | 16408088 | ||
| C3H10T1/2 | Function assay | Induction of osteogenesis in mouse C3H10T1/2 cells assessed as induction of osteoblast specific marker alkaline phosphatase by immunofluorescence method, EC50 = 1 μM. | 16408003 | |||
| HEK293T | Function assay | 1 hr | Inhibition of BODIPY-cyclopamine binding to Smo expressed in HEK293T cells after 1 hr by fluorescence microscopy, IC50 = 1.5 μM. | 16408088 | ||
| Shh Light2 | Function assay | 30 hrs | Activation of Shh in mouse Shh Light2 cells assessed as beta-galactosidase activity after 30 hrs by luciferase reporter gene assay in presence of 100 nM 3-keto-N-aminoethyl-N'-aminocaproyldihydrocinnamoyl cyclopamine | 16408088 | ||
| HEK293T | Function assay | 5 uM | 4 hrs | Inhibition of BODIPY-cyclopamine binding to Smo N-terminal cysteine domain expressed in HEK293T cells at 5 uM after 4 hrs by fluorescence microscopy | 16408088 | |
| HEK293T | Function assay | 5 uM | 4 hrs | Inhibition of BODIPY-cyclopamine binding to Smo C-terminal cytoplasmic domain expressed in HEK293T cells at 5 uM after 4 hrs by fluorescence microscopy | 16408088 | |
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SK-N-MC cells | 29435139 | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 4 mg/mL
(7.68 mM)
Water : Insoluble Ethanol : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 520.62 | Formula | C31H32N6O2 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 483367-10-8 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | Shh Signaling Antagonist VI | Smiles | C1CCC(CC1)N2C=NC3=C(N=C(N=C32)OC4=CC=CC5=CC=CC=C54)NC6=CC=C(C=C6)N7CCOCC7 | ||
Mechanism of Action
| Targets/IC50/Ki |
Smoothened
(HEK293T cells) ~1.5 μM
|
|---|---|
| In vitro |
Purmorphamine activates the Hedgehog pathway by directly binds and activates Smoothened with IC50 of ~ 1.5 μM in compete with cyclopamine, a Smo antagonist. This compound is a potent inducer of osteogenesis in multipotent C3H10T1/2 cells. The EC50 (based on ALP expression) for this chemical is 1 μM in C3H10T1/2 cells. It (1 μM) and BMP-4 (100 ng/mL) together increase ALP activity more than 90-fold in 3T3-L1 cells. In contrast to BMP-4, this compound induces osteogenesis by activating Hedgehog signaling in multipotent mesenchymal progenitor cells. |
| Kinase Assay |
Binding assay
|
|
Smo binding assays are conducted with BODIPY-cyclopamine and Smo-overexpressing cells as previously described4,5, using CMV promoter-based, SV40 origin-containing expression constructs for Smo-Myc3, the deletion mutant SmoCRD (deletion of amino acids 68 to 182), and SmoCT (deletion of amino acids 556 to 793). HEK 293T cells are grown on poly-D-lysine-treated glass coverslips in 12-well plates until 70% confluency and then transfected with the appropriate expression construct (0.5 g/well) using FuGene 6 according the manufacturer
|
|
| In vivo |
Purmorphamine up-regulates ALP expression in human mesenchymal stem cell-based constructs on rats. |
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | Patch1 / Gli1 / LC3 / p62 |
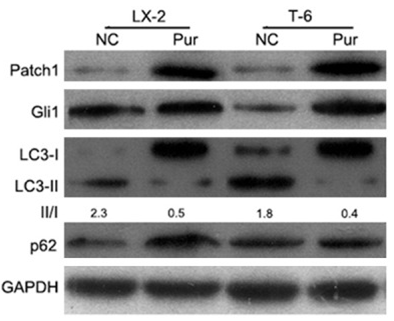
|
26609469 |
| Immunofluorescence | SOX18 |
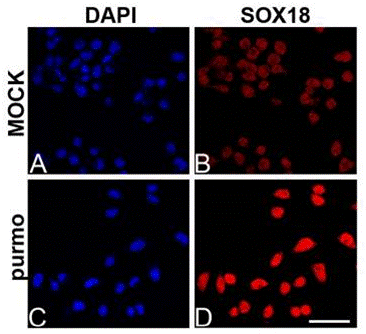
|
26588701 |
| Growth inhibition assay | Cell viability |
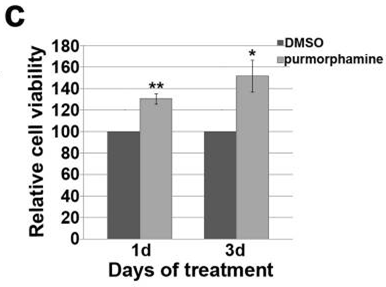
|
26588701 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































