research use only
AZ 628 Raf inhibitor
Cat.No.S2746
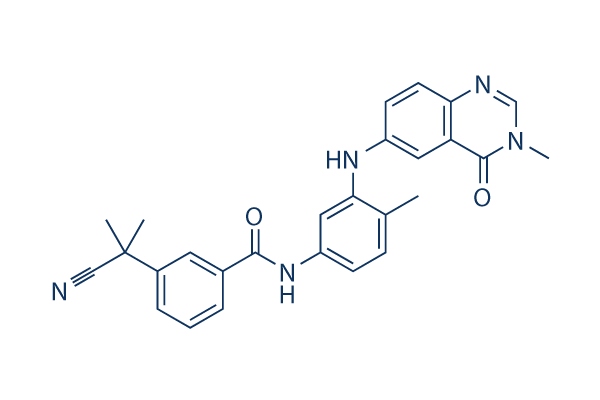
Chemical Structure
Molecular Weight: 451.52
Quality Control
| Related Targets | ERK p38 MAPK JNK MEK Ras KRas S6 Kinase MAP4K TAK1 Mixed Lineage Kinase |
|---|---|
| Other Raf Inhibitors | LY3009120 Exarafenib (KIN-2787) GDC-0879 Avutometinib (Ro5126766, CH5126766) PLX-4720 SB590885 TAK-632 GW5074 RAF265 (CHIR-265) PLX7904 |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| A375 | Function assay | 75 mins | Inhibition of BRAF V600E mutant-mediated ERK phosphorylation in human A375 cells after 75 mins, IC50=0.015μM | 23398453 | ||
| Sf9 | Function assay | 20 mins | Inhibition of human recombinant N-terminus His-tagged BRAF V600E mutant expressed in Sf9 cells coexpressing CDC37 using biotinylated MEK as substrate and MgCl2 incubated for 20 mins prior to MgCl2 addition measured after 1 hr in presence of 10 uM ATP, IC50=0.034μM | 23398453 | ||
| Sf9 | Function assay | 20 mins | Inhibition of human recombinant N-terminus His-tagged BRAF V600E mutant expressed in Sf9 cells coexpressing CDC37 using biotinylated MEK as substrate and MgCl2 incubated for 20 mins prior to MgCl2 addition measured after 1 hr in presence of 5 mM ATP, IC50=0.27μM | 23398453 | ||
| Sf9 | Function assay | 20 mins | Inhibition of human recombinant N-terminus His-tagged BRAF expressed in Sf9 cells coexpressing CDC37 using biotinylated MEK as substrate and MgCl2 incubated for 20 mins prior to MgCl2 addition measured after 1 hr in presence of 5 mM ATP, IC50=2.64μM | 23398453 | ||
| NIH/3T3 | Function assay | 75 mins | Inhibition of wild type BRAF in mouse NIH/3T3 cells assessed as decrease in ERK phosphorylation after 75 mins by Western blotting analysis | 23398453 | ||
| CCD-1070SK | Function assay | 75 mins | Inhibition of wild type BRAF in human CCD-1070SK cells assessed as decrease in ERK phosphorylation after 75 mins by Western blotting analysis | 23398453 | ||
| MCF7 | Function assay | 75 mins | Inhibition of wild type BRAF in human MCF7 cells assessed as decrease in ERK phosphorylation after 75 mins by Western blotting analysis | 23398453 | ||
| A549 | Function assay | 75 mins | Inhibition of wild type BRAF in human A549 cells assessed as decrease in ERK phosphorylation after 75 mins by Western blotting analysis | 23398453 | ||
| PC3 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human PC3 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| PANC1 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human PANC1 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| NCI-H23 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human NCI-H23 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| NCI-H460 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human NCI-H460 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| Calu6 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human Calu6 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| HCT15 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human HCT15 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| MDA-MB-231 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human MDA-MB-231 cells expressing BRAF G464V mutant after 72 hrs by MTS assay | 23398453 | ||
| SKBR3 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human SKBR3 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| MCF7 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human MCF7 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| HCT116 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human HCT116 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| MDA-MB-468 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human MDA-MB-468 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| A549 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human A549 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| HT-29 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human HT-29 cells expressing BRAF V600E mutant after 72 hrs by MTS assay | 23398453 | ||
| SW620 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human SW620 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| A2058 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human A2058 cells expressing BRAF V600E mutant after 72 hrs by MTS assay | 23398453 | ||
| DU145 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human DU145 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| SK-MEL-24 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human SK-MEL-24 cells expressing BRAF V600E mutant after 72 hrs by MTS assay | 23398453 | ||
| SK-MEL-28 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human SK-MEL-28 cells expressing BRAF V600E mutant after 72 hrs by MTS assay | 23398453 | ||
| SK-MEL-3 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human SK-MEL-3 cells expressing BRAF V600E mutant after 72 hrs by MTS assay | 23398453 | ||
| MALME-3M | Antiproliferative assay | 72 hrs | Antiproliferative activity against human MALME-3M cells expressing BRAF V600E mutant after 72 hrs by MTS assay | 23398453 | ||
| COLO205 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human COLO205 cells expressing BRAF V600E mutant after 72 hrs by MTS assay | 23398453 | ||
| A375 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human A375 cells expressing BRAF V600E mutant after 72 hrs by MTS assay | 23398453 | ||
| A375 | Function assay | 75 mins | Inhibition of BRAF V600E mutant in human A375 cells assessed as decrease in ERK phosphorylation after 75 mins by Western blotting analysis | 23398453 | ||
| MiaPaCa | Antiproliferative assay | 72 hrs | Antiproliferative activity against human MiaPaCa cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| NIH:OVCAR3 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human NIH:OVCAR3 cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| LNCaP-FGC | Antiproliferative assay | 72 hrs | Antiproliferative activity against human LNCaP-FGC cells expressing wild type BRAF after 72 hrs by MTS assay | 23398453 | ||
| Malme-3M | Function assay | Inhibition of B-Raf V600E mutant in human Malme-3M cells assessed as basal ERK phosphorylation, IC50=0.027μM | 24675381 | |||
| A673 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for A673 cells | 29435139 | |||
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SK-N-MC cells | 29435139 | |||
| NB-EBc1 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for NB-EBc1 cells | 29435139 | |||
| SK-N-SH | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SK-N-SH cells | 29435139 | |||
| NB1643 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for NB1643 cells | 29435139 | |||
| RD | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for RD cells | 29435139 | |||
| ACN | Growth inhibition assay | Inhibition of human ACN cell growth in a cell viability assay, IC50=0.0001009μM | SANGER | |||
| SH-4 | Growth inhibition assay | Inhibition of human SH-4 cell growth in a cell viability assay, IC50=0.003009μM | SANGER | |||
| LB2518-MEL | Growth inhibition assay | Inhibition of human LB2518-MEL cell growth in a cell viability assay, IC50=0.004758μM | SANGER | |||
| A101D | Growth inhibition assay | Inhibition of human A101D cell growth in a cell viability assay, IC50=0.005819μM | SANGER | |||
| HT-144 | Growth inhibition assay | Inhibition of human HT-144 cell growth in a cell viability assay, IC50=0.006606μM | SANGER | |||
| SIG-M5 | Growth inhibition assay | Inhibition of human SIG-M5 cell growth in a cell viability assay, IC50=0.009455μM | SANGER | |||
| DU-4475 | Growth inhibition assay | Inhibition of human DU-4475 cell growth in a cell viability assay, IC50=0.01023μM | SANGER | |||
| OCI-AML2 | Growth inhibition assay | Inhibition of human OCI-AML2 cell growth in a cell viability assay, IC50=0.017μM | SANGER | |||
| COLO-829 | Growth inhibition assay | Inhibition of human COLO-829 cell growth in a cell viability assay, IC50=0.02565μM | SANGER | |||
| PSN1 | Growth inhibition assay | Inhibition of human PSN1 cell growth in a cell viability assay, IC50=0.02737μM | SANGER | |||
| UACC-257 | Growth inhibition assay | Inhibition of human UACC-257 cell growth in a cell viability assay, IC50=0.02749μM | SANGER | |||
| RKO | Growth inhibition assay | Inhibition of human RKO cell growth in a cell viability assay, IC50=0.03767μM | SANGER | |||
| MMAC-SF | Growth inhibition assay | Inhibition of human MMAC-SF cell growth in a cell viability assay, IC50=0.04007μM | SANGER | |||
| Calu-6 | Growth inhibition assay | Inhibition of human Calu-6 cell growth in a cell viability assay, IC50=0.05147μM | SANGER | |||
| LS-513 | Growth inhibition assay | Inhibition of human LS-513 cell growth in a cell viability assay, IC50=0.07415μM | SANGER | |||
| LS-411N | Growth inhibition assay | Inhibition of human LS-411N cell growth in a cell viability assay, IC50=0.09161μM | SANGER | |||
| NOS-1 | Growth inhibition assay | Inhibition of human NOS-1 cell growth in a cell viability assay, IC50=0.09265μM | SANGER | |||
| SW872 | Growth inhibition assay | Inhibition of human SW872 cell growth in a cell viability assay, IC50=0.09648μM | SANGER | |||
| KMOE-2 | Growth inhibition assay | Inhibition of human KMOE-2 cell growth in a cell viability assay, IC50=0.11588μM | SANGER | |||
| MZ7-mel | Growth inhibition assay | Inhibition of human MZ7-mel cell growth in a cell viability assay, IC50=0.11727μM | SANGER | |||
| CGTH-W-1 | Growth inhibition assay | Inhibition of human CGTH-W-1 cell growth in a cell viability assay, IC50=0.1213μM | SANGER | |||
| CP66-MEL | Growth inhibition assay | Inhibition of human CP66-MEL cell growth in a cell viability assay, IC50=0.13176μM | SANGER | |||
| K5 | Growth inhibition assay | Inhibition of human K5 cell growth in a cell viability assay, IC50=0.1342μM | SANGER | |||
| HL-60 | Growth inhibition assay | Inhibition of human HL-60 cell growth in a cell viability assay, IC50=0.15441μM | SANGER | |||
| ONS-76 | Growth inhibition assay | Inhibition of human ONS-76 cell growth in a cell viability assay, IC50=0.16662μM | SANGER | |||
| NB69 | Growth inhibition assay | Inhibition of human NB69 cell growth in a cell viability assay, IC50=0.19131μM | SANGER | |||
| LAMA-84 | Growth inhibition assay | Inhibition of human LAMA-84 cell growth in a cell viability assay, IC50=0.19242μM | SANGER | |||
| EM-2 | Growth inhibition assay | Inhibition of human EM-2 cell growth in a cell viability assay, IC50=0.19591μM | SANGER | |||
| IST-MEL1 | Growth inhibition assay | Inhibition of human IST-MEL1 cell growth in a cell viability assay, IC50=0.20413μM | SANGER | |||
| HCC2998 | Growth inhibition assay | Inhibition of human HCC2998 cell growth in a cell viability assay, IC50=0.22187μM | SANGER | |||
| GDM-1 | Growth inhibition assay | Inhibition of human GDM-1 cell growth in a cell viability assay, IC50=0.22947μM | SANGER | |||
| MZ1-PC | Growth inhibition assay | Inhibition of human MZ1-PC cell growth in a cell viability assay, IC50=0.27999μM | SANGER | |||
| BB49-HNC | Growth inhibition assay | Inhibition of human BB49-HNC cell growth in a cell viability assay, IC50=0.3518μM | SANGER | |||
| D-392MG | Growth inhibition assay | Inhibition of human D-392MG cell growth in a cell viability assay, IC50=0.35607μM | SANGER | |||
| 697 | Growth inhibition assay | Inhibition of human 697 cell growth in a cell viability assay, IC50=0.36388μM | SANGER | |||
| ML-2 | Growth inhibition assay | Inhibition of human ML-2 cell growth in a cell viability assay, IC50=0.36886μM | SANGER | |||
| MEG-01 | Growth inhibition assay | Inhibition of human MEG-01 cell growth in a cell viability assay, IC50=0.376μM | SANGER | |||
| SK-MEL-1 | Growth inhibition assay | Inhibition of human SK-MEL-1 cell growth in a cell viability assay, IC50=0.38362μM | SANGER | |||
| KS-1 | Growth inhibition assay | Inhibition of human KS-1 cell growth in a cell viability assay, IC50=0.39337μM | SANGER | |||
| NCI-SNU-1 | Growth inhibition assay | Inhibition of human NCI-SNU-1 cell growth in a cell viability assay, IC50=0.44786μM | SANGER | |||
| SK-NEP-1 | Growth inhibition assay | Inhibition of human SK-NEP-1 cell growth in a cell viability assay, IC50=0.47226μM | SANGER | |||
| A4-Fuk | Growth inhibition assay | Inhibition of human A4-Fuk cell growth in a cell viability assay, IC50=0.48375μM | SANGER | |||
| NB14 | Growth inhibition assay | Inhibition of human NB14 cell growth in a cell viability assay, IC50=0.49331μM | SANGER | |||
| NCI-H747 | Growth inhibition assay | Inhibition of human NCI-H747 cell growth in a cell viability assay, IC50=0.50637μM | SANGER | |||
| K-562 | Growth inhibition assay | Inhibition of human K-562 cell growth in a cell viability assay, IC50=0.66695μM | SANGER | |||
| NCI-H1648 | Growth inhibition assay | Inhibition of human NCI-H1648 cell growth in a cell viability assay, IC50=0.67266μM | SANGER | |||
| LAN-6 | Growth inhibition assay | Inhibition of human LAN-6 cell growth in a cell viability assay, IC50=0.68236μM | SANGER | |||
| BV-173 | Growth inhibition assay | Inhibition of human BV-173 cell growth in a cell viability assay, IC50=0.69154μM | SANGER | |||
| NKM-1 | Growth inhibition assay | Inhibition of human NKM-1 cell growth in a cell viability assay, IC50=0.73537μM | SANGER | |||
| MZ2-MEL | Growth inhibition assay | Inhibition of human MZ2-MEL cell growth in a cell viability assay, IC50=0.79367μM | SANGER | |||
| BE-13 | Growth inhibition assay | Inhibition of human BE-13 cell growth in a cell viability assay, IC50=0.80575μM | SANGER | |||
| HUTU-80 | Growth inhibition assay | Inhibition of human HUTU-80 cell growth in a cell viability assay, IC50=0.98331μM | SANGER | |||
| CTV-1 | Growth inhibition assay | Inhibition of human CTV-1 cell growth in a cell viability assay, IC50=1.20065μM | SANGER | |||
| NOMO-1 | Growth inhibition assay | Inhibition of human NOMO-1 cell growth in a cell viability assay, IC50=1.20638μM | SANGER | |||
| GI-ME-N | Growth inhibition assay | Inhibition of human GI-ME-N cell growth in a cell viability assay, IC50=1.29212μM | SANGER | |||
| TE-12 | Growth inhibition assay | Inhibition of human TE-12 cell growth in a cell viability assay, IC50=1.31417μM | SANGER | |||
| ALL-PO | Growth inhibition assay | Inhibition of human ALL-PO cell growth in a cell viability assay, IC50=1.34869μM | SANGER | |||
| ST486 | Growth inhibition assay | Inhibition of human ST486 cell growth in a cell viability assay, IC50=1.38662μM | SANGER | |||
| MV-4-11 | Growth inhibition assay | Inhibition of human MV-4-11 cell growth in a cell viability assay, IC50=1.42334μM | SANGER | |||
| SJSA-1 | Growth inhibition assay | Inhibition of human SJSA-1 cell growth in a cell viability assay, IC50=1.43007μM | SANGER | |||
| LB373-MEL-D | Growth inhibition assay | Inhibition of human LB373-MEL-D cell growth in a cell viability assay, IC50=1.66482μM | SANGER | |||
| ECC12 | Growth inhibition assay | Inhibition of human ECC12 cell growth in a cell viability assay, IC50=1.7149μM | SANGER | |||
| SUP-T1 | Growth inhibition assay | Inhibition of human SUP-T1 cell growth in a cell viability assay, IC50=1.73943μM | SANGER | |||
| NB5 | Growth inhibition assay | Inhibition of human NB5 cell growth in a cell viability assay, IC50=1.95406μM | SANGER | |||
| BB30-HNC | Growth inhibition assay | Inhibition of human BB30-HNC cell growth in a cell viability assay, IC50=1.96472μM | SANGER | |||
| DOHH-2 | Growth inhibition assay | Inhibition of human DOHH-2 cell growth in a cell viability assay, IC50=1.98728μM | SANGER | |||
| KM12 | Growth inhibition assay | Inhibition of human KM12 cell growth in a cell viability assay, IC50=2.08103μM | SANGER | |||
| D-502MG | Growth inhibition assay | Inhibition of human D-502MG cell growth in a cell viability assay, IC50=2.26142μM | SANGER | |||
| RPMI-8226 | Growth inhibition assay | Inhibition of human RPMI-8226 cell growth in a cell viability assay, IC50=2.46647μM | SANGER | |||
| EW-24 | Growth inhibition assay | Inhibition of human EW-24 cell growth in a cell viability assay, IC50=2.71195μM | SANGER | |||
| TE-441-T | Growth inhibition assay | Inhibition of human TE-441-T cell growth in a cell viability assay, IC50=2.80668μM | SANGER | |||
| NCI-SNU-5 | Growth inhibition assay | Inhibition of human NCI-SNU-5 cell growth in a cell viability assay, IC50=2.81269μM | SANGER | |||
| BL-70 | Growth inhibition assay | Inhibition of human BL-70 cell growth in a cell viability assay, IC50=2.86352μM | SANGER | |||
| HD-MY-Z | Growth inhibition assay | Inhibition of human HD-MY-Z cell growth in a cell viability assay, IC50=2.9442μM | SANGER | |||
| OPM-2 | Growth inhibition assay | Inhibition of human OPM-2 cell growth in a cell viability assay, IC50=3.0342μM | SANGER | |||
| PF-382 | Growth inhibition assay | Inhibition of human PF-382 cell growth in a cell viability assay, IC50=3.05237μM | SANGER | |||
| SK-MEL-2 | Growth inhibition assay | Inhibition of human SK-MEL-2 cell growth in a cell viability assay, IC50=3.21751μM | SANGER | |||
| QIMR-WIL | Growth inhibition assay | Inhibition of human QIMR-WIL cell growth in a cell viability assay, IC50=3.23473μM | SANGER | |||
| LC-1F | Growth inhibition assay | Inhibition of human LC-1F cell growth in a cell viability assay, IC50=3.30326μM | SANGER | |||
| SU-DHL-1 | Growth inhibition assay | Inhibition of human SU-DHL-1 cell growth in a cell viability assay, IC50=3.52915μM | SANGER | |||
| KG-1 | Growth inhibition assay | Inhibition of human KG-1 cell growth in a cell viability assay, IC50=3.80547μM | SANGER | |||
| MSTO-211H | Growth inhibition assay | Inhibition of human MSTO-211H cell growth in a cell viability assay, IC50=3.95028μM | SANGER | |||
| L-540 | Growth inhibition assay | Inhibition of human L-540 cell growth in a cell viability assay, IC50=3.95889μM | SANGER | |||
| HCC1187 | Growth inhibition assay | Inhibition of human HCC1187 cell growth in a cell viability assay, IC50=4.03307μM | SANGER | |||
| RL95-2 | Growth inhibition assay | Inhibition of human RL95-2 cell growth in a cell viability assay, IC50=4.10119μM | SANGER | |||
| KINGS-1 | Growth inhibition assay | Inhibition of human KINGS-1 cell growth in a cell viability assay, IC50=4.17422μM | SANGER | |||
| SCC-15 | Growth inhibition assay | Inhibition of human SCC-15 cell growth in a cell viability assay, IC50=4.21476μM | SANGER | |||
| EW-12 | Growth inhibition assay | Inhibition of human EW-12 cell growth in a cell viability assay, IC50=4.4199μM | SANGER | |||
| SIMA | Growth inhibition assay | Inhibition of human SIMA cell growth in a cell viability assay, IC50=4.46019μM | SANGER | |||
| NCI-H720 | Growth inhibition assay | Inhibition of human NCI-H720 cell growth in a cell viability assay, IC50=4.7198μM | SANGER | |||
| HCC1599 | Growth inhibition assay | Inhibition of human HCC1599 cell growth in a cell viability assay, IC50=4.8017μM | SANGER | |||
| 8-MG-BA | Growth inhibition assay | Inhibition of human 8-MG-BA cell growth in a cell viability assay, IC50=4.85156μM | SANGER | |||
| RPMI-6666 | Growth inhibition assay | Inhibition of human RPMI-6666 cell growth in a cell viability assay, IC50=5.08842μM | SANGER | |||
| BC-3 | Growth inhibition assay | Inhibition of human BC-3 cell growth in a cell viability assay, IC50=5.25452μM | SANGER | |||
| LB996-RCC | Growth inhibition assay | Inhibition of human LB996-RCC cell growth in a cell viability assay, IC50=5.28958μM | SANGER | |||
| TE-5 | Growth inhibition assay | Inhibition of human TE-5 cell growth in a cell viability assay, IC50=5.39519μM | SANGER | |||
| MPP-89 | Growth inhibition assay | Inhibition of human MPP-89 cell growth in a cell viability assay, IC50=5.53594μM | SANGER | |||
| DJM-1 | Growth inhibition assay | Inhibition of human DJM-1 cell growth in a cell viability assay, IC50=5.63516μM | SANGER | |||
| LC-2-ad | Growth inhibition assay | Inhibition of human LC-2-ad cell growth in a cell viability assay, IC50=5.66116μM | SANGER | |||
| NEC8 | Growth inhibition assay | Inhibition of human NEC8 cell growth in a cell viability assay, IC50=5.68736μM | SANGER | |||
| KGN | Growth inhibition assay | Inhibition of human KGN cell growth in a cell viability assay, IC50=5.69705μM | SANGER | |||
| MOLT-13 | Growth inhibition assay | Inhibition of human MOLT-13 cell growth in a cell viability assay, IC50=5.82867μM | SANGER | |||
| EW-3 | Growth inhibition assay | Inhibition of human EW-3 cell growth in a cell viability assay, IC50=5.87026μM | SANGER | |||
| HOP-62 | Growth inhibition assay | Inhibition of human HOP-62 cell growth in a cell viability assay, IC50=6.06751μM | SANGER | |||
| EW-22 | Growth inhibition assay | Inhibition of human EW-22 cell growth in a cell viability assay, IC50=6.09281μM | SANGER | |||
| EW-11 | Growth inhibition assay | Inhibition of human EW-11 cell growth in a cell viability assay, IC50=6.27578μM | SANGER | |||
| EB2 | Growth inhibition assay | Inhibition of human EB2 cell growth in a cell viability assay, IC50=6.33123μM | SANGER | |||
| TE-9 | Growth inhibition assay | Inhibition of human TE-9 cell growth in a cell viability assay, IC50=6.42042μM | SANGER | |||
| IST-SL2 | Growth inhibition assay | Inhibition of human IST-SL2 cell growth in a cell viability assay, IC50=6.51935μM | SANGER | |||
| GB-1 | Growth inhibition assay | Inhibition of human GB-1 cell growth in a cell viability assay, IC50=6.57311μM | SANGER | |||
| D-247MG | Growth inhibition assay | Inhibition of human D-247MG cell growth in a cell viability assay, IC50=6.63417μM | SANGER | |||
| TK10 | Growth inhibition assay | Inhibition of human TK10 cell growth in a cell viability assay, IC50=6.64799μM | SANGER | |||
| A388 | Growth inhibition assay | Inhibition of human A388 cell growth in a cell viability assay, IC50=6.70097μM | SANGER | |||
| EHEB | Growth inhibition assay | Inhibition of human EHEB cell growth in a cell viability assay, IC50=7.13498μM | SANGER | |||
| ECC4 | Growth inhibition assay | Inhibition of human ECC4 cell growth in a cell viability assay, IC50=7.2399μM | SANGER | |||
| HAL-01 | Growth inhibition assay | Inhibition of human HAL-01 cell growth in a cell viability assay, IC50=7.45581μM | SANGER | |||
| SR | Growth inhibition assay | Inhibition of human SR cell growth in a cell viability assay, IC50=7.64918μM | SANGER | |||
| KU812 | Growth inhibition assay | Inhibition of human KU812 cell growth in a cell viability assay, IC50=8.02775μM | SANGER | |||
| YT | Growth inhibition assay | Inhibition of human YT cell growth in a cell viability assay, IC50=8.11695μM | SANGER | |||
| MOLT-4 | Growth inhibition assay | Inhibition of human MOLT-4 cell growth in a cell viability assay, IC50=8.51012μM | SANGER | |||
| SK-UT-1 | Growth inhibition assay | Inhibition of human SK-UT-1 cell growth in a cell viability assay, IC50=8.59275μM | SANGER | |||
| MLMA | Growth inhibition assay | Inhibition of human MLMA cell growth in a cell viability assay, IC50=8.65529μM | SANGER | |||
| MFH-ino | Growth inhibition assay | Inhibition of human MFH-ino cell growth in a cell viability assay, IC50=8.9591μM | SANGER | |||
| IM-9 | Growth inhibition assay | Inhibition of human IM-9 cell growth in a cell viability assay, IC50=9.06101μM | SANGER | |||
| NCI-H322M | Growth inhibition assay | Inhibition of human NCI-H322M cell growth in a cell viability assay, IC50=9.28598μM | SANGER | |||
| CESS | Growth inhibition assay | Inhibition of human CESS cell growth in a cell viability assay, IC50=9.3786μM | SANGER | |||
| KE-37 | Growth inhibition assay | Inhibition of human KE-37 cell growth in a cell viability assay, IC50=9.49296μM | SANGER | |||
| U-87-MG | Growth inhibition assay | Inhibition of human U-87-MG cell growth in a cell viability assay, IC50=9.78165μM | SANGER | |||
| EoL-1-cell | Growth inhibition assay | Inhibition of human EoL-1-cell cell growth in a cell viability assay, IC50=9.93708μM | SANGER | |||
| CA46 | Growth inhibition assay | Inhibition of human CA46 cell growth in a cell viability assay, IC50=10.5081μM | SANGER | |||
| LB1047-RCC | Growth inhibition assay | Inhibition of human LB1047-RCC cell growth in a cell viability assay, IC50=10.5566μM | SANGER | |||
| NB1 | Growth inhibition assay | Inhibition of human NB1 cell growth in a cell viability assay, IC50=10.5731μM | SANGER | |||
| LS-123 | Growth inhibition assay | Inhibition of human LS-123 cell growth in a cell viability assay, IC50=10.5844μM | SANGER | |||
| GOTO | Growth inhibition assay | Inhibition of human GOTO cell growth in a cell viability assay, IC50=10.966μM | SANGER | |||
| LB2241-RCC | Growth inhibition assay | Inhibition of human LB2241-RCC cell growth in a cell viability assay, IC50=11.3688μM | SANGER | |||
| RPMI-8402 | Growth inhibition assay | Inhibition of human RPMI-8402 cell growth in a cell viability assay, IC50=11.7395μM | SANGER | |||
| EW-18 | Growth inhibition assay | Inhibition of human EW-18 cell growth in a cell viability assay, IC50=12.2119μM | SANGER | |||
| SF268 | Growth inhibition assay | Inhibition of human SF268 cell growth in a cell viability assay, IC50=12.5572μM | SANGER | |||
| NCI-H2171 | Growth inhibition assay | Inhibition of human NCI-H2171 cell growth in a cell viability assay, IC50=12.7506μM | SANGER | |||
| SCLC-21H | Growth inhibition assay | Inhibition of human SCLC-21H cell growth in a cell viability assay, IC50=13.4345μM | SANGER | |||
| MDA-MB-134-VI | Growth inhibition assay | Inhibition of human MDA-MB-134-VI cell growth in a cell viability assay, IC50=13.8879μM | SANGER | |||
| ES5 | Growth inhibition assay | Inhibition of human ES5 cell growth in a cell viability assay, IC50=13.9428μM | SANGER | |||
| KALS-1 | Growth inhibition assay | Inhibition of human KALS-1 cell growth in a cell viability assay, IC50=14.1325μM | SANGER | |||
| GI-1 | Growth inhibition assay | Inhibition of human GI-1 cell growth in a cell viability assay, IC50=14.1988μM | SANGER | |||
| U-698-M | Growth inhibition assay | Inhibition of human U-698-M cell growth in a cell viability assay, IC50=14.435μM | SANGER | |||
| LP-1 | Growth inhibition assay | Inhibition of human LP-1 cell growth in a cell viability assay, IC50=14.4661μM | SANGER | |||
| NCI-H23 | Growth inhibition assay | Inhibition of human NCI-H23 cell growth in a cell viability assay, IC50=14.724μM | SANGER | |||
| HC-1 | Growth inhibition assay | Inhibition of human HC-1 cell growth in a cell viability assay, IC50=14.7737μM | SANGER | |||
| NCI-H1355 | Growth inhibition assay | Inhibition of human NCI-H1355 cell growth in a cell viability assay, IC50=14.9946μM | SANGER | |||
| LC4-1 | Growth inhibition assay | Inhibition of human LC4-1 cell growth in a cell viability assay, IC50=15.1355μM | SANGER | |||
| CCRF-CEM | Growth inhibition assay | Inhibition of human CCRF-CEM cell growth in a cell viability assay, IC50=15.2306μM | SANGER | |||
| TE-1 | Growth inhibition assay | Inhibition of human TE-1 cell growth in a cell viability assay, IC50=15.5684μM | SANGER | |||
| NTERA-S-cl-D1 | Growth inhibition assay | Inhibition of human NTERA-S-cl-D1 cell growth in a cell viability assay, IC50=16.395μM | SANGER | |||
| LOXIMVI | Growth inhibition assay | Inhibition of human LOXIMVI cell growth in a cell viability assay, IC50=16.4329μM | SANGER | |||
| MONO-MAC-6 | Growth inhibition assay | Inhibition of human MONO-MAC-6 cell growth in a cell viability assay, IC50=16.6532μM | SANGER | |||
| NCI-H716 | Growth inhibition assay | Inhibition of human NCI-H716 cell growth in a cell viability assay, IC50=16.9882μM | SANGER | |||
| JiyoyeP-2003 | Growth inhibition assay | Inhibition of human JiyoyeP-2003 cell growth in a cell viability assay, IC50=17.2361μM | SANGER | |||
| SW954 | Growth inhibition assay | Inhibition of human SW954 cell growth in a cell viability assay, IC50=17.7495μM | SANGER | |||
| CAS-1 | Growth inhibition assay | Inhibition of human CAS-1 cell growth in a cell viability assay, IC50=17.7612μM | SANGER | |||
| SF539 | Growth inhibition assay | Inhibition of human SF539 cell growth in a cell viability assay, IC50=17.8653μM | SANGER | |||
| COLO-800 | Growth inhibition assay | Inhibition of human COLO-800 cell growth in a cell viability assay, IC50=18.0144μM | SANGER | |||
| KY821 | Growth inhibition assay | Inhibition of human KY821 cell growth in a cell viability assay, IC50=18.4608μM | SANGER | |||
| LU-65 | Growth inhibition assay | Inhibition of human LU-65 cell growth in a cell viability assay, IC50=18.5947μM | SANGER | |||
| JVM-2 | Growth inhibition assay | Inhibition of human JVM-2 cell growth in a cell viability assay, IC50=18.6058μM | SANGER | |||
| CRO-AP2 | Growth inhibition assay | Inhibition of human CRO-AP2 cell growth in a cell viability assay, IC50=18.6171μM | SANGER | |||
| NCI-H128 | Growth inhibition assay | Inhibition of human NCI-H128 cell growth in a cell viability assay, IC50=19.2368μM | SANGER | |||
| AM-38 | Growth inhibition assay | Inhibition of human AM-38 cell growth in a cell viability assay, IC50=19.7389μM | SANGER | |||
| NCI-H64 | Growth inhibition assay | Inhibition of human NCI-H64 cell growth in a cell viability assay, IC50=19.9518μM | SANGER | |||
| ES6 | Growth inhibition assay | Inhibition of human ES6 cell growth in a cell viability assay, IC50=20.3657μM | SANGER | |||
| KARPAS-45 | Growth inhibition assay | Inhibition of human KARPAS-45 cell growth in a cell viability assay, IC50=20.4183μM | SANGER | |||
| NCI-H1581 | Growth inhibition assay | Inhibition of human NCI-H1581 cell growth in a cell viability assay, IC50=20.5843μM | SANGER | |||
| KURAMOCHI | Growth inhibition assay | Inhibition of human KURAMOCHI cell growth in a cell viability assay, IC50=20.7537μM | SANGER | |||
| NALM-6 | Growth inhibition assay | Inhibition of human NALM-6 cell growth in a cell viability assay, IC50=21.2955μM | SANGER | |||
| DSH1 | Growth inhibition assay | Inhibition of human DSH1 cell growth in a cell viability assay, IC50=21.3697μM | SANGER | |||
| Becker | Growth inhibition assay | Inhibition of human Becker cell growth in a cell viability assay, IC50=21.5612μM | SANGER | |||
| HH | Growth inhibition assay | Inhibition of human HH cell growth in a cell viability assay, IC50=21.8318μM | SANGER | |||
| EB-3 | Growth inhibition assay | Inhibition of human EB-3 cell growth in a cell viability assay, IC50=21.8562μM | SANGER | |||
| HCC2157 | Growth inhibition assay | Inhibition of human HCC2157 cell growth in a cell viability assay, IC50=22.2184μM | SANGER | |||
| no-10 | Growth inhibition assay | Inhibition of human no-10 cell growth in a cell viability assay, IC50=22.5132μM | SANGER | |||
| D-263MG | Growth inhibition assay | Inhibition of human D-263MG cell growth in a cell viability assay, IC50=22.9998μM | SANGER | |||
| Raji | Growth inhibition assay | Inhibition of human Raji cell growth in a cell viability assay, IC50=23.4081μM | SANGER | |||
| EKVX | Growth inhibition assay | Inhibition of human EKVX cell growth in a cell viability assay, IC50=23.7929μM | SANGER | |||
| LS-1034 | Growth inhibition assay | Inhibition of human LS-1034 cell growth in a cell viability assay, IC50=24.2647μM | SANGER | |||
| SF126 | Growth inhibition assay | Inhibition of human SF126 cell growth in a cell viability assay, IC50=24.9217μM | SANGER | |||
| MC-CAR | Growth inhibition assay | Inhibition of human MC-CAR cell growth in a cell viability assay, IC50=25.0901μM | SANGER | |||
| RL | Growth inhibition assay | Inhibition of human RL cell growth in a cell viability assay, IC50=25.1477μM | SANGER | |||
| KNS-81-FD | Growth inhibition assay | Inhibition of human KNS-81-FD cell growth in a cell viability assay, IC50=25.8563μM | SANGER | |||
| Ramos-2G6-4C10 | Growth inhibition assay | Inhibition of human Ramos-2G6-4C10 cell growth in a cell viability assay, IC50=26.0093μM | SANGER | |||
| TE-8 | Growth inhibition assay | Inhibition of human TE-8 cell growth in a cell viability assay, IC50=26.0599μM | SANGER | |||
| ES8 | Growth inhibition assay | Inhibition of human ES8 cell growth in a cell viability assay, IC50=26.1087μM | SANGER | |||
| SK-PN-DW | Growth inhibition assay | Inhibition of human SK-PN-DW cell growth in a cell viability assay, IC50=27.3933μM | SANGER | |||
| NCI-H2126 | Growth inhibition assay | Inhibition of human NCI-H2126 cell growth in a cell viability assay, IC50=27.6384μM | SANGER | |||
| NCI-SNU-16 | Growth inhibition assay | Inhibition of human NCI-SNU-16 cell growth in a cell viability assay, IC50=27.7795μM | SANGER | |||
| MHH-PREB-1 | Growth inhibition assay | Inhibition of human MHH-PREB-1 cell growth in a cell viability assay, IC50=27.9604μM | SANGER | |||
| HCE-T | Growth inhibition assay | Inhibition of human HCE-T cell growth in a cell viability assay, IC50=29.2587μM | SANGER | |||
| IST-MES1 | Growth inhibition assay | Inhibition of human IST-MES1 cell growth in a cell viability assay, IC50=29.329μM | SANGER | |||
| CTB-1 | Growth inhibition assay | Inhibition of human CTB-1 cell growth in a cell viability assay, IC50=31.889μM | SANGER | |||
| MOLT-16 | Growth inhibition assay | Inhibition of human MOLT-16 cell growth in a cell viability assay, IC50=32.0214μM | SANGER | |||
| BC-1 | Growth inhibition assay | Inhibition of human BC-1 cell growth in a cell viability assay, IC50=32.527μM | SANGER | |||
| A253 | Growth inhibition assay | Inhibition of human A253 cell growth in a cell viability assay, IC50=32.9626μM | SANGER | |||
| D-336MG | Growth inhibition assay | Inhibition of human D-336MG cell growth in a cell viability assay, IC50=33.9563μM | SANGER | |||
| KARPAS-422 | Growth inhibition assay | Inhibition of human KARPAS-422 cell growth in a cell viability assay, IC50=34.5318μM | SANGER | |||
| BB65-RCC | Growth inhibition assay | Inhibition of human BB65-RCC cell growth in a cell viability assay, IC50=36.0231μM | SANGER | |||
| WSU-NHL | Growth inhibition assay | Inhibition of human WSU-NHL cell growth in a cell viability assay, IC50=36.1908μM | SANGER | |||
| COLO-824 | Growth inhibition assay | Inhibition of human COLO-824 cell growth in a cell viability assay, IC50=37.3777μM | SANGER | |||
| COR-L279 | Growth inhibition assay | Inhibition of human COR-L279 cell growth in a cell viability assay, IC50=37.3993μM | SANGER | |||
| SK-N-FI | Growth inhibition assay | Inhibition of human SK-N-FI cell growth in a cell viability assay, IC50=38.3192μM | SANGER | |||
| U-266 | Growth inhibition assay | Inhibition of human U-266 cell growth in a cell viability assay, IC50=38.8703μM | SANGER | |||
| A3-KAW | Growth inhibition assay | Inhibition of human A3-KAW cell growth in a cell viability assay, IC50=39.1454μM | SANGER | |||
| Daudi | Growth inhibition assay | Inhibition of human Daudi cell growth in a cell viability assay, IC50=42.5065μM | SANGER | |||
| NCI-H1092 | Growth inhibition assay | Inhibition of human NCI-H1092 cell growth in a cell viability assay, IC50=43.345μM | SANGER | |||
| NB10 | Growth inhibition assay | Inhibition of human NB10 cell growth in a cell viability assay, IC50=43.945μM | SANGER | |||
| LXF-289 | Growth inhibition assay | Inhibition of human LXF-289 cell growth in a cell viability assay, IC50=44.786μM | SANGER | |||
| MHH-NB-11 | Growth inhibition assay | Inhibition of human MHH-NB-11 cell growth in a cell viability assay, IC50=45.1968μM | SANGER | |||
| LOUCY | Growth inhibition assay | Inhibition of human LOUCY cell growth in a cell viability assay, IC50=45.9997μM | SANGER | |||
| DMS-114 | Growth inhibition assay | Inhibition of human DMS-114 cell growth in a cell viability assay, IC50=46.1196μM | SANGER | |||
| NCI-H1882 | Growth inhibition assay | Inhibition of human NCI-H1882 cell growth in a cell viability assay, IC50=46.5042μM | SANGER | |||
| ES3 | Growth inhibition assay | Inhibition of human ES3 cell growth in a cell viability assay, IC50=46.5616μM | SANGER | |||
| OCUB-M | Growth inhibition assay | Inhibition of human OCUB-M cell growth in a cell viability assay, IC50=46.9249μM | SANGER | |||
| TGW | Growth inhibition assay | Inhibition of human TGW cell growth in a cell viability assay, IC50=47.6222μM | SANGER | |||
| OS-RC-2 | Growth inhibition assay | Inhibition of human OS-RC-2 cell growth in a cell viability assay, IC50=48.0389μM | SANGER | |||
| D-542MG | Growth inhibition assay | Inhibition of human D-542MG cell growth in a cell viability assay, IC50=49.6469μM | SANGER | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 90 mg/mL
(199.32 mM)
Water : Insoluble Ethanol : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 451.52 | Formula | C27H25N5O2 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 878739-06-1 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N/A | Smiles | CC1=C(C=C(C=C1)NC(=O)C2=CC(=CC=C2)C(C)(C)C#N)NC3=CC4=C(C=C3)N=CN(C4=O)C | ||
Mechanism of Action
| Targets/IC50/Ki |
C-Raf-1
(Cell-free assay) 29 nM
B-Raf (V600E)
(Cell-free assay) 34 nM
B-Raf
(Cell-free assay) 105 nM
|
|---|---|
| In vitro |
AZ628 prevents activation of number of tyrosine protein kinases including VEGFR2, DDR2, Lyn, Flt1, FMS and others. AZ628 suppresses anchorage-dependent and -independent growth, gives rise to cell cycle arrest, and induces apoptosis in colon and melanoma cell lines harboring B-RafV600E mutation. The profile of AZ628 cross-reactivity suggests that similar to AZ628 may be antiangiogenic based on prevention of VEGFR2. AZ628-resistant clones are approximately 100-fold more resistant to AZ628 than the parental cell line, exhibiting IC50 of approximately 10 μM, compared with 0.1 μM for the parental cell line. Effective suppression of p-ERK1/2 levels is observed in the M14 parental cell line following treatment with increasing concentrations of AZ628. AZ628-resistant clones express elevated CRAF. Elevated CRAF expression is a potential mechanism of acquired resistance to continuous AZ628 exposure, resulting in sustained activation of ERK1/2. p-ERK1/2 activity is not significantly inhibited by exposure to AZ628 in one of these three AZ628-insensitive cell lines (Wm1552C). Unlike in the AZ628-resistant M14 cells in which AZ628 fails to suppress the activation of ERK, AZ628 treatment efficiently attenuates ERK activation in the NRAS mutant melanoma cells. |
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | p-ERK / ERK / p-MEK / MEK / BIM / BRAF / CRAF |

|
21098728 |
| Growth inhibition assay | Cell viability |
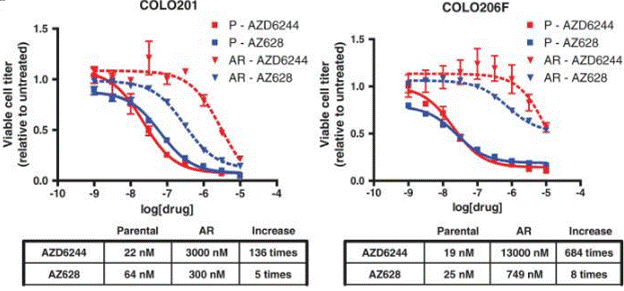
|
21098728 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































