
- Inhibitors
- By product type
- Natural Products
- Inducing Agents
- Peptides
- Antibiotics
- Antibody-drug Conjugates(ADC)
- PROTAC
- Hydrotropic Agents
- Dyes
- By Signaling Pathways
- PI3K/Akt/mTOR
- Epigenetics
- Methylation
- Immunology & Inflammation
- Protein Tyrosine Kinase
- Angiogenesis
- Apoptosis
- Autophagy
By research - Antibodies
- Compound Libraries
- Popular Compound Libraries
- Customize Library
- Clinical and FDA-approved Related
- Bioactive Compound Libraries
- Inhibitor Related
- Natural Product Related
- Metabolism Related
- Cell Death Related
- By Signaling Pathway
- By Disease
- Anti-infection and Antiviral Related
- Neuronal and Immunology Related
- Fragment and Covalent Related
- FDA-approved Drug Library
- FDA-approved & Passed Phase I Drug Library
- Preclinical/Clinical Compound Library
- Bioactive Compound Library-I
- Bioactive Compound Library-Ⅱ
- Kinase Inhibitor Library
- Express-Pick Library
- Natural Product Library
- Human Endogenous Metabolite Compound Library
- Alkaloid Compound LibraryNew
- Angiogenesis Related compound Library
- Anti-Aging Compound Library
- Anti-alzheimer Disease Compound Library
- Antibiotics compound Library
- Anti-cancer Compound Library
- Anti-cancer Compound Library-Ⅱ
- Anti-cancer Metabolism Compound Library
- Anti-Cardiovascular Disease Compound Library
- Anti-diabetic Compound Library
- Anti-infection Compound Library
- Antioxidant Compound Library
- Anti-parasitic Compound Library
- Antiviral Compound Library
- Apoptosis Compound Library
- Autophagy Compound Library
- Calcium Channel Blocker LibraryNew
- Cambridge Cancer Compound Library
- Carbohydrate Metabolism Compound LibraryNew
- Cell Cycle compound library
- CNS-Penetrant Compound Library
- Covalent Inhibitor Library
- Cytokine Inhibitor LibraryNew
- Cytoskeletal Signaling Pathway Compound Library
- DNA Damage/DNA Repair compound Library
- Drug-like Compound Library
- Endoplasmic Reticulum Stress Compound Library
- Epigenetics Compound Library
- Exosome Secretion Related Compound LibraryNew
- FDA-approved Anticancer Drug LibraryNew
- Ferroptosis Compound Library
- Flavonoid Compound Library
- Fragment Library
- Glutamine Metabolism Compound Library
- Glycolysis Compound Library
- GPCR Compound Library
- Gut Microbial Metabolite Library
- HIF-1 Signaling Pathway Compound Library
- Highly Selective Inhibitor Library
- Histone modification compound library
- HTS Library for Drug Discovery
- Human Hormone Related Compound LibraryNew
- Human Transcription Factor Compound LibraryNew
- Immunology/Inflammation Compound Library
- Inhibitor Library
- Ion Channel Ligand Library
- JAK/STAT compound library
- Lipid Metabolism Compound LibraryNew
- Macrocyclic Compound Library
- MAPK Inhibitor Library
- Medicine Food Homology Compound Library
- Metabolism Compound Library
- Methylation Compound Library
- Mouse Metabolite Compound LibraryNew
- Natural Organic Compound Library
- Neuronal Signaling Compound Library
- NF-κB Signaling Compound Library
- Nucleoside Analogue Library
- Obesity Compound Library
- Oxidative Stress Compound LibraryNew
- Plant Extract Library
- Phenotypic Screening Library
- PI3K/Akt Inhibitor Library
- Protease Inhibitor Library
- Protein-protein Interaction Inhibitor Library
- Pyroptosis Compound Library
- Small Molecule Immuno-Oncology Compound Library
- Mitochondria-Targeted Compound LibraryNew
- Stem Cell Differentiation Compound LibraryNew
- Stem Cell Signaling Compound Library
- Natural Phenol Compound LibraryNew
- Natural Terpenoid Compound LibraryNew
- TGF-beta/Smad compound library
- Traditional Chinese Medicine Library
- Tyrosine Kinase Inhibitor Library
- Ubiquitination Compound Library
-
Cherry Picking
You can personalize your library with chemicals from within Selleck's inventory. Build the right library for your research endeavors by choosing from compounds in all of our available libraries.
Please contact us at info@selleckchem.com to customize your library.
You could select:
- Bioreagents
- qPCR
- 2x SYBR Green qPCR Master Mix
- 2x SYBR Green qPCR Master Mix(Low ROX)
- 2x SYBR Green qPCR Master Mix(High ROX)
- Protein Assay
- Protein A/G Magnetic Beads for IP
- Anti-Flag magnetic beads
- Anti-Flag Affinity Gel
- Anti-Myc magnetic beads
- Anti-HA magnetic beads
- Poly DYKDDDDK Tag Peptide lyophilized powder
- Protease Inhibitor Cocktail
- Protease Inhibitor Cocktail (EDTA-Free, 100X in DMSO)
- Phosphatase Inhibitor Cocktail (2 Tubes, 100X)
- Cell Biology
- Cell Counting Kit-8 (CCK-8)
- Animal Experiment
- Mouse Direct PCR Kit (For Genotyping)
- Featured Products
- MRTX1133
- Nab-Paclitaxel
- KP-457
- IAG933
- RMC-6236 (Daraxonrasib)
- RMC-7977
- Zoldonrasib (RMC-9805)
- GsMTx4
- Navitoclax (ABT-263)
- TSA (Trichostatin A)
- Y-27632 Dihydrochloride
- SB431542
- SB202190
- MK-2206 Dihydrochloride
- LY294002
- Alisertib (MLN8237)
- XAV-939
- CHIR-99021 (Laduviglusib)
- Bafilomycin A1 (Baf-A1)
- Thiazovivin (TZV)
- CP-673451
- Verteporfin
- DAPT
- Galunisertib (LY2157299)
- MG132
- SBE-β-CD
- Tween 80
- Bavdegalutamide (ARV-110)
- Z-VAD-FMK
- Wnt-C59 (C59)
- IWR-1-endo
- (+)-JQ1
- 3-Deazaneplanocin A (DZNep) Hydrochloride
- RepSox (E-616452)
- Erastin
- Q-VD-Oph
- Puromycin Dihydrochloride
- Cycloheximide
- Telaglenastat (CB-839)
- A-83-01
- Ceralasertib (AZD6738)
- Liproxstatin-1
- Emricasan (IDN-6556)
- PMA (Phorbol 12-myristate 13-acetate)
- Dibutyryl cAMP (Bucladesine) sodium
- Nedisertib (M3814)
- PLX5622
- IKE (Imidazole Ketone Erastin)
- STM2457
- Saruparib (AZD5305)
- New Products
- Contact Us
research use only
Vismodegib (GDC-0449) Hedgehog inhibitor
Cat.No.S1082
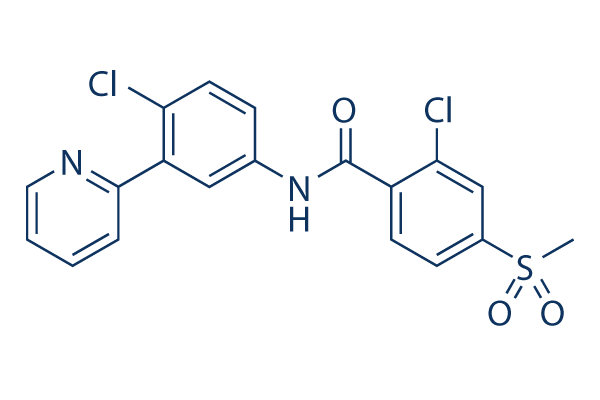
Chemical Structure
Molecular Weight: 421.3
Quality Control
Batch:
Purity:
99.99%
99.99
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| IGROV-1 | Growth Inhibition Assay | IC50=0.07248 μM | SANGER | |||
| HCE-T | Growth Inhibition Assay | IC50=1.32247 μM | SANGER | |||
| D-542MG | Growth Inhibition Assay | IC50=1.86737 μM | SANGER | |||
| 23132-87 | Growth Inhibition Assay | IC50=4.40147 μM | SANGER | |||
| HDLM-2 | Growth Inhibition Assay | IC50=8.04766 μM | SANGER | |||
| ACN | Growth Inhibition Assay | IC50=8.50109 μM | SANGER | |||
| HuO-3N1 | Growth Inhibition Assay | IC50=9.60108 μM | SANGER | |||
| BHT-101 | Growth Inhibition Assay | IC50=11.38 μM | SANGER | |||
| KYSE-150 | Growth Inhibition Assay | IC50=11.5841 μM | SANGER | |||
| MC-IXC | Growth Inhibition Assay | IC50=12.2292 μM | SANGER | |||
| D-423MG | Growth Inhibition Assay | IC50=12.7657 μM | SANGER | |||
| NY | Growth Inhibition Assay | IC50=14.8903 μM | SANGER | |||
| HOS | Growth Inhibition Assay | IC50=15.6719 μM | SANGER | |||
| NB7 | Growth Inhibition Assay | IC50=15.891 μM | SANGER | |||
| DMS-273 | Growth Inhibition Assay | IC50=16.6713 μM | SANGER | |||
| MDA-MB-361 | Growth Inhibition Assay | IC50=17.2711 μM | SANGER | |||
| DU-145 | Growth Inhibition Assay | IC50=18.32 μM | SANGER | |||
| NCI-H82 | Growth Inhibition Assay | IC50=19.8386 μM | SANGER | |||
| NCI-SNU-1 | Growth Inhibition Assay | IC50=20.0196 μM | SANGER | |||
| GCT | Growth Inhibition Assay | IC50=20.8824 μM | SANGER | |||
| C2BBe1 | Growth Inhibition Assay | IC50=21.1058 μM | SANGER | |||
| LB2241-RCC | Growth Inhibition Assay | IC50=21.8441 μM | SANGER | |||
| COLO-829 | Growth Inhibition Assay | IC50=22.1871 μM | SANGER | |||
| EW-11 | Growth Inhibition Assay | IC50=22.8022 μM | SANGER | |||
| NCI-H526 | Growth Inhibition Assay | IC50=23.4717 μM | SANGER | |||
| SF295 | Growth Inhibition Assay | IC50=24.0252 μM | SANGER | |||
| D-566MG | Growth Inhibition Assay | IC50=25.2943 μM | SANGER | |||
| 8505C | Growth Inhibition Assay | IC50=25.6331 μM | SANGER | |||
| HT-29 | Growth Inhibition Assay | IC50=26.0431 μM | SANGER | |||
| NBsusSR | Growth Inhibition Assay | IC50=26.8006 μM | SANGER | |||
| BV-173 | Growth Inhibition Assay | IC50=28.3182 μM | SANGER | |||
| CTB-1 | Growth Inhibition Assay | IC50=30.1031 μM | SANGER | |||
| JAR | Growth Inhibition Assay | IC50=32.5371 μM | SANGER | |||
| CAMA-1 | Growth Inhibition Assay | IC50=33.4615 μM | SANGER | |||
| CAL-51 | Growth Inhibition Assay | IC50=34.7176 μM | SANGER | |||
| A172 | Growth Inhibition Assay | IC50=37.4921 μM | SANGER | |||
| QIMR-WIL | Growth Inhibition Assay | IC50=38.0708 μM | SANGER | |||
| AsPC-1 | Growth Inhibition Assay | IC50=38.4651 μM | SANGER | |||
| MKN7 | Growth Inhibition Assay | IC50=39.0079 μM | SANGER | |||
| ONS-76 | Growth Inhibition Assay | IC50=43.3057 μM | SANGER | |||
| RS4-11 | Growth Inhibition Assay | IC50=44.0752 μM | SANGER | |||
| NOS-1 | Growth Inhibition Assay | IC50=44.6031 μM | SANGER | |||
| A101D | Growth Inhibition Assay | IC50=44.8023 μM | SANGER | |||
| HCC1806 | Growth Inhibition Assay | IC50=46.1148 μM | SANGER | |||
| CAL-27 | Growth Inhibition Assay | IC50=47.7246 μM | SANGER | |||
| BT-549 | Growth Inhibition Assay | IC50=48.5315 μM | SANGER | |||
| LCLC-97TM1 | Growth Inhibition Assay | IC50=49.2413 μM | SANGER | |||
| A4-Fuk | Growth Inhibition Assay | IC50=49.849 μM | SANGER | |||
| OVCAR-4 | Growth Inhibition Assay | IC50=50.0601 μM | SANGER | |||
| HD-MY-Z | Growth Inhibition Assay | IC50=50.7764 μM | SANGER | |||
| NCI-H292 | Growth Inhibition Assay | IC50=50.8758 μM | SANGER | |||
| Sk-ChA-1 | Growth Inhibition Assay | 0.25–50 μM | 72 h | IC50=74.54±2.58μM | 25742482 | |
| Mz-ChA-1 | Growth Inhibition Assay | 0.25–50 μM | 72 h | IC50=54.97±3.45μM | 25742482 | |
| Smo-WT | Growth Inhibition Assay | IC50 of 14 nM | 24291104 | |||
| Smo-D473H | Growth Inhibition Assay | IC50 of 7.1 μM | 24291104 | |||
| K562 | Function Assay | 10 μM | 72 h | reduces the expression of Gli1 | 23319824 | |
| T315I BCR-ABL BaF3 | Function Assay | 10 μM | 72 h | reduces the expression of Gli1 | 23319824 | |
| TF-1 BCR-ABL | Function Assay | 10 μM | 72 h | reduces the expression of Gli1 | 23319824 | |
| Shh-Light 2 | Function assay | 2 days | Antagonist activity at Smo in mouse Shh-Light 2 cells assessed as inhibition of Shh-induced Gli1-reporter activity after 2 days by dual-luciferase reporter gene method, IC50 = 0.0015 μM. | 23063522 | ||
| NIH/3T3 | Function assay | Inhibition of hedgehog signaling (unknown origin) expressed in mouse NIH/3T3 cells by Gli-dual-luciferase reporter assay, IC50 = 0.0023 μM. | 27180012 | |||
| NIH/3T3 Light2 | Function assay | Inhibition of hedgehog signaling pathway in mouse NIH/3T3 Light2 cells by Gli-luciferase reporter gene assay, IC50 = 0.0023 μM. | 28688278 | |||
| Light2 | Function assay | 48 hrs | Inhibition of hedgehog signaling pathway in mouse Light2 cells after 48 hrs by Gli-luciferase reporter gene assay, IC50 = 0.00246 μM. | 29739714 | ||
| HEPM | Function assay | Inhibition of SHH in human HEPM cells by Gli-luciferase reporter gene assay, INH = 0.0028 μM. | 19716296 | |||
| Shh Light2 | Function assay | Inhibition of SHH in mouse Shh Light2 cells by GLI-responsive firefly luciferase reporter gene assay, IC50 = 0.003 μM. | 19309080 | |||
| HEPM | Function assay | Inhibition of Hedgehog signaling in human HEPM cells assessed as reduction in receptor-mediated Gli1 mRNA expression by reporter gene assay, INH = 0.003 μM. | 20875741 | |||
| C3H10T1/2 | Function assay | 20 hrs | Inhibition of Smo in mouse C3H10T1/2 cells using human recombinant SHH assessed as effect on SMO/SHH transient transcriptional activation after 20 hrs by Gli-luciferase reporter assay, IC50 = 0.005 μM. | 24900436 | ||
| NIH/3T3 | Function assay | 24 hrs | Inhibition of hedgehog signaling pathway in mouse NIH/3T3 cells measured after 24 hrs by Gli-dual luciferase reporter gene assay, IC50 = 0.005 μM. | 27810591 | ||
| NIH3T3 | Function assay | 48 hrs | Inhibition of smo-mediated hedgehog signaling pathway in mouse NIH3T3 cells expressing GRE-Luc reporter gene assessed as inhibition of SAG-induced GRE-Luc reporter activity after 48 hrs by luminescence assay, IC50 = 0.006 μM. | 29499483 | ||
| Shh-light2 | Function assay | Inhibition of Smo-mediated Hh signaling in human Shh-light2 cells by luciferase reporter gene assay, IC50 = 0.007 μM. | 22268551 | |||
| HEK293 | Function assay | 2 hrs | Displacement of BODIPY-labelled cyclopamine from human Smo receptor expressed in HEK293 cells after 2 hrs by fluorescence microscopy, IC50 = 0.007 μM. | 22268551 | ||
| NIH3T3 | Function assay | Inhibition of hedgehog receptor signaling pathway in mouse NIH3T3 cells transfected with Gli-reporter gene by luciferase reporter gene assay, IC50 = 0.00717 μM. | 24923765 | |||
| NIH/3T3 | Function assay | 48 hrs | Inhibition of Hedgehog signaling pathway in mouse NIH/3T3 cells assessed as reduction in Sonic hedgehog-induced Gli luciferase activity after 48 hrs by Dual-luciferase reporter gene assay, IC50 = 0.00717 μM. | 30249494 | ||
| NIH3T3 | Function assay | 48 hrs | Inhibition of hedgehog signalling in mouse NIH3T3 cells stably transfected with Gli-luciferase construct incubated for 48 hrs by dual luciferase reporter gene assay, IC50 = 0.0072 μM. | 26827136 | ||
| NIH3T3 | Function assay | 48 hrs | Inhibition of Sonic-induced hedgehog signalling in mouse NIH3T3 cells after 48 hrs by Gli-luciferase reporter assay, IC50 = 0.0072 μM. | 26820554 | ||
| NIH3T3 | Function assay | 48 hrs | Inhibition of SHH signaling pathway in mouse NIH3T3 cells measured after 48 hrs by Gli-luciferase reporter assay, IC50 = 0.0072 μM. | 24176396 | ||
| NIH/3T3 | Function assay | 48 hrs | Inhibition of hedgehog signaling pathway in mouse NIH/3T3 cells after 48 hrs by Gli-luciferase reporter gene assay, IC50 = 0.0072 μM. | 24486203 | ||
| NIH/3T3 | Function assay | Inhibition of hedgehog signaling pathway in mouse NIH/3T3 cells by Gli1-luciferase reporter gene assay, IC50 = 0.0072 μM. | 24405704 | |||
| NIH/3T3 | Function assay | Inhibition of hedgehog signaling pathway in mouse NIH/3T3 cells carrying stably transfected Gli-reporter construct by luciferase reporter assay, IC50 = 0.0072 μM. | 28642101 | |||
| C3H10T1/2 | Function assay | 6 hrs | Inhibition of SAG-induced differentiation of mouse mesenchymal pluripotent C3H10T1/2 cells to alkaline phosphatase positive oeseoblasts after 6 hrs, IC50 = 0.011 μM. | 22268551 | ||
| HCC827 | Function assay | Displacement of [3H]-cyclopamine from SMO V404M mutant in gefitinib resistant human HCC827 cells by scintillation counting, Ki = 0.0122 μM. | 28787156 | |||
| C3H10T1/2 | Function assay | Inhibition of SHH in mouse C3H10T1/2 cells by Gli-luciferase reporter gene assay, IC50 = 0.013 μM. | 19716296 | |||
| S12 | Function assay | Inhibition of Hedgehog signaling in mouse S12 cells assessed as reduction in receptor-mediated Gli1 mRNA expression by reporter gene assay, INH = 0.013 μM. | 20875741 | |||
| TM3 | Function assay | 48 hrs | Inhibition of Hh signaling pathway in mouse TM3 cells assessed as downregulation of Gli1 gene expression after 48 hrs by luciferase reporter gene assay, EC50 = 0.013 μM. | 26976215 | ||
| NIH/3T3 | Function assay | Inhibition of hedgehog signaling pathway in mouse NIH/3T3 cells expressing wild type Smo assessed as reduction in Gli mRNA expression by RT-PCR method, IC50 = 0.0144 μM. | 27810591 | |||
| S12 | Function assay | 48 hrs | Inhibition of human SHH pathway in mouse S12 cells assessed as GLI-mediated transcriptional activity after 48 hrs by luciferase reporter gene assay, IC50 = 0.015 μM. | 21438527 | ||
| NIH/3T3 | Function assay | Inhibition of SMO in mouse NIH/3T3 cells assessed as inhibition of SAG-induced hedgehog-mediated luminescence signaling by GRE-luciferase reporter gene assay, IC50 = 0.016 μM. | 26119500 | |||
| U2OS | Function assay | 2 hrs | Displacement of [3H]cyclopamine from wild type Smo expressed in U2OS cells after 2 hrs by scintillation counting, Ki = 0.0162 μM. | 23063522 | ||
| NIH3T3 | Function assay | 30 hrs | Inhibition of Smo receptor (unknown origin) expressed in NIH3T3 cells assessed as inhibition of Smo agonist SAG-induced GRE activation after 30 hrs by luciferase reporter gene assay, IC50 = 0.023 μM. | 24726807 | ||
| medulloblastoma cell | Antiproliferative assay | 36 hrs | Antiproliferative activity against Ptch+/- and p53-/- mouse medulloblastoma cells after 36 hrs by Brdu incorporation assay, IC50 = 0.0304 μM. | 28688278 | ||
| Shh Light2 | Function assay | Antagonist activity at smoothened (unknown origin) expressed in mouse Shh Light2 cells co-expressing Gli-dependent reporter gene assessed as inhibition of Hh signaling by dual luciferase reporter gene assay, IC50 = 0.033 μM. | 24491459 | |||
| Light2 | Function assay | Inhibition of hedgehog signaling pathway in mouse Light2 cells in Shh conditioned medium by Gli-luciferase reporter gene assay, IC50 = 0.039 μM. | 28873303 | |||
| Shh Light2 | Function assay | Inhibition of Smo-mediated Hh signalling pathway in mouse Shh Light2 cells by Gli-luciferase reporter gene assay, IC50 = 0.0392 μM. | 27736063 | |||
| ASZ001 | Function assay | 48 hrs | Inhibition of Hedgehog signaling in mouse ASZ001 cells assessed as downregulation of Gli1 mRNA expression after 48 hrs by RT-PCR analysis, IC50 = 0.04 μM. | 24900716 | ||
| NIH/3T3 | Function assay | 48 hrs | Antagonist activity at Smo receptor in mouse NIH/3T3 cells harboring GRE-Luc assessed as inhibition of SAG-induced Hh signaling pathway preincubated with cells followed by SAG addition measured after 48 hrs by luciferase reporter gene assay, IC50 = 0.046 μM. | 29857275 | ||
| Light2 | Function assay | 30 hrs | Inhibition of hedgehog signaling pathway in mouse Light2 cells in Shh conditioned medium after 30 hrs by Gli-Renilla luciferase reporter gene assay, IC50 = 0.05 μM. | 30099257 | ||
| C3H10T1/2 | Function assay | 6 days | Inhibition of hedgehog signaling pathway-mediated differentiation of mouse C3H10T1/2 cells assessed as decrease in SAG-induced ALP activity after 6 days by chemiluminescence-based assay, IC50 = 0.05 μM. | 30099257 | ||
| DaOY | Function assay | 48 hrs | Inhibition of Hedgehog signaling in human DaOY cells assessed as downregulation of Gli1 mRNA expression after 48 hrs by RT-PCR analysis, IC50 = 0.086 μM. | 24900716 | ||
| C3H10T1/2 | Function assay | 1 hr | Inhibition of Hedgehog signaling pathway in mouse C3H10T1/2 cells assessed as reduction in purmorphamine-induced increase in alkaline phosphatase level using CDP-star as substrate pretreated for 1 hr followed by purmorphamine addition measured after 5 day, IC50 = 0.14 μM. | 28947939 | ||
| M210B4 | Function assay | 24 hrs | Inhibition of Hedgehog signaling in mouse M210B4 cells assessed as downregulation of Ptch mRNA expression after 24 hrs by RT-PCR analysis, IC50 = 0.14 μM. | 24900716 | ||
| M210B4 | Function assay | 24 hrs | Inhibition of Hedgehog signaling in mouse M210B4 cells assessed as downregulation of Gli1 mRNA expression after 24 hrs by RT-PCR analysis, IC50 = 0.2 μM. | 24900716 | ||
| Shh Light2 | Function assay | 1 hr | Inhibition of Hedgehog signaling in mouse Shh Light2 cells assessed as reduction in purmorphamine-induced Gli mediated transcriptional activity pretreated for 1 hr followed by purmorphamine addition measured after 48 hrs by luciferase reporter gene assay, IC50 = 0.98 μM. | 28947939 | ||
| NIH/3T3 | Function assay | Inhibition of hedgehog signaling pathway in mouse NIH/3T3 cells harboring Smo D477H mutant assessed as reduction in Gli mRNA expression by RT-PCR method, IC50 = 1.196 μM. | 27810591 | |||
| S12 | Function assay | 48 hrs | Inhibition of human SHH pathway in mouse S12 cells assessed as GLI-mediated transcriptional activity after 48 hrs by luciferase reporter gene assay in presence of 0.5 mg/mL human alpha-1-acid glycoprotein, IC50 = 1.75 μM. | 21438527 | ||
| Calu6 | Function assay | 4 hrs | Plasma concentration in human Calu6 cells xenografted in nude mouse at 75 mg/kg, po bid measured after 4 hrs post fifth dose, Cp = 23 μM. | 19716296 | ||
| Calu-6 | Function assay | Plasma concentration in Calu-6 cells xenografted nude mouse PK/PD model at 75 mg/kg, po bid for 5 days measured 4 hrs post last dose relative to untreated control, Cp = 23 μM. | 20875741 | |||
| T47D | Cytotoxicity assay | 72 hrs | Cytotoxicity against human T47D cells after 72 hrs by MTT assay, IC50 = 41.34 μM. | 29107429 | ||
| LS180 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human LS180 cells after 72 hrs by sulforhodamine-B assay, IC50 = 45 μM. | 30099257 | ||
| LS174T | Antiproliferative assay | 72 hrs | Antiproliferative activity against human LS174T cells after 72 hrs by CellTiter-Glo luminescent cell viability assay, IC50 = 45.81 μM. | 26820554 | ||
| BxPC3 | Antiproliferative assay | 72 hrs | Antiproliferative activity against human BxPC3 cells after 72 hrs by CellTiter-Glo luminescent cell viability assay, IC50 = 47.95 μM. | 26820554 | ||
| medulloblastoma cell | Antitumor assay | Antitumor activity against mouse medulloblastoma cells xenografted in Ptch +/- mouse assessed as decrease in tumor volume at 12.5 mg/kg, po bid. | 19716296 | |||
| S12 | Function assay | 48 hrs | Inhibition of human SHH pathway in mouse S12 cells assessed as GLI-mediated transcriptional activity after 48 hrs by luciferase reporter gene assay in presence of 1 mg/mL human alpha-1-acid glycoprotein. | 21438527 | ||
| Shh Light2 | Function assay | 0.1 uM | 36 hrs | Inhibition of Smo-mediated Hh signalling pathway in mouse Shh Light2 cells assessed as suppression of Gli1-mRNA expression at 0.1 uM measured after 36 hrs by RT-qPCR analysis. | 27736063 | |
| SW1353 | Antiproliferative assay | 10 uM | 48 hrs | Antiproliferative activity against human SW1353 cells assessed as growth inhibition at 10 uM after 48 hrs by MTT assay. | 24491459 | |
| DaOY | Antiproliferative assay | 10 uM | 48 hrs | Antiproliferative activity against human DaOY cells assessed as growth inhibition at 10 uM after 48 hrs by MTT assay. | 24491459 | |
| A549 | Antiproliferative assay | 10 uM | 48 hrs | Antiproliferative activity against human A549 cells assessed as growth inhibition at 10 uM after 48 hrs by MTT assay. | 24491459 | |
| HCT116 | Antiproliferative assay | 10 uM | 48 hrs | Antiproliferative activity against human HCT116 cells assessed as growth inhibition at 10 uM after 48 hrs by MTT assay. | 24491459 | |
| NIH/3T3 | Function assay | 10 uM | Inhibition of Hedgehog signaling in mouse NIH/3T3 cells assessed as down regulation of mPTCH1 mRNA expression at 10 uM. | 24491459 | ||
| MSC | Function assay | 20 to 80 uM | Inhibition of hedgehog signaling in mouse MSC cells assessed as down regulation of alkaline phosphatase at 20 to 80 uM. | 24491459 | ||
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SK-N-MC cells. | 29435139 | |||
| TC32 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for TC32 cells. | 29435139 | |||
| NIH3T3-Luc | Function assay | 1 uM | 48 hrs | Inhibition of smo-mediated hedgehog signaling pathway in mouse NIH3T3-Luc cells assessed as downregulation of Gli mRNA expression at 1 uM after 48 hrs by real-time PCR analysis. | 29499483 | |
| C3H10T1/2 | Function assay | 24 hrs | Inhibition of hedgehog signaling pathway-mediated GLI1 mRNA expression in mouse C3H10T1/2 cells after 24 hrs by RT-PCR analysis in presence of (1R,3aR,4S,7aR)-7a-methyl-1-((R)-6-methylheptan-2-yl)octahydro-1H-inden-4-yl 3-hydroxybenzoate. | 24730984 | ||
| C3H10T1/2 | Function assay | 24 hrs | Inhibition of hedgehog signaling pathway-mediated GLI1 mRNA expression in mouse C3H10T1/2 cells after 24 hrs by RT-PCR analysis in presence of VD3. | 24730984 | ||
| NIH/3T3 | Function assay | 100 nM | 36 hrs | Inhibition of hedgehog signaling pathway in mouse NIH/3T3 cells assessed as downregulation of Shh-induced Gli1 mRNA expression levels at 100 nM after 36 hrs by RT-qPCR analysis. | 28873303 | |
| C3H10T1/2 | Function assay | 100 nM | 36 hrs | Inhibition of hedgehog signaling pathway in mouse C3H10T1/2 cells assessed as downregulation of Shh-induced Gli1 mRNA expression levels at 100 nM after 36 hrs by RT-qPCR analysis. | 28873303 | |
| NIH/3T3 | Function assay | 100 nM | 36 hrs | Inhibition of hedgehog signaling pathway in mouse NIH/3T3 cells assessed as downregulation of Shh-induced ptch1 mRNA expression levels at 100 nM after 36 hrs by RT-qPCR analysis. | 28873303 | |
| C3H10T1/2 | Function assay | 100 nM | 36 hrs | Inhibition of hedgehog signaling pathway in mouse C3H10T1/2 cells assessed as downregulation of Shh-induced ptch1 mRNA expression levels at 100 nM after 36 hrs by RT-qPCR analysis. | 28873303 | |
| medulloblastoma cell | Antiproliferative assay | up to 30 uM | 72 hrs | Antiproliferative activity against mouse Ptch+/- p53-/- medulloblastoma cells assessed as reduction in cell proliferation up to 30 uM after 72 hrs by MTS assay. | 28873303 | |
| medulloblastoma cell | Apoptosis assay | Induction of apoptosis in mouse Ptch+/- p53-/- medulloblastoma cells implanted in athymic nude mouse at 12.5 mg/kg, ip bid for 15 consecutive days by TUNEL assay. | 28873303 | |||
| NIH/3T3 | Function assay | Inhibition of hedgehog signaling pathway expressed in mouse NIH/3T3 cells assessed as inhibition of hedgehog-induced Gli-2 accumulation at tip of primary cilia by DAPI staining based confocal microscopic analysis. | 27810591 | |||
| NIH/3T3 | Function assay | Inhibition of hedgehog signaling pathway in mouse NIH/3T3 cells assessed as inhibition of hedgehog-induced Smo-EGFP ciliary translocation by DAPI staining based confocal microscopic analysis. | 27810591 | |||
| medulloblastoma cell | Anti-tumor assay | Anti-tumor activity against mouse Ptch+/- p53-/- medulloblastoma cells implanted in athymic nude mouse assessed as tumor growth inhibition at 20 mg/kg, po administered twice daily via gavage measured every other day during compound dosing. | 29739714 | |||
| C3H10T1/2 | Function assay | 4.1 to 1000 nM | Allosteric inhibition of Smo in mouse C3H10T1/2 cells assessed as inhibition of purmorphamine-induced Gli1 transcriptional activity at 4.1 to 1000 nM by qPCr analysis. | 23074541 | ||
| C3H10T1/2 | Function assay | 4.1 to 1000 nM | Competitive inhibition of Smo in mouse C3H10T1/2 cells assessed as inhibition of SAG-induced Gli1 transcriptional activity at 4.1 to 1000 nM by qPCR analysis. | 23074541 | ||
| Click to View More Cell Line Experimental Data | ||||||
Chemical Information, Storage & Stability
| Molecular Weight | 421.3 | Formula | C19H14Cl2N2O3S |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 879085-55-9 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | GDC-0449 | Smiles | CS(=O)(=O)C1=CC(=C(C=C1)C(=O)NC2=CC(=C(C=C2)Cl)C3=CC=CC=N3)Cl | ||
Solubility
|
In vitro |
DMSO
: 84 mg/mL
(199.38 mM)
Water : Insoluble Ethanol : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
mg/kg
g
μL
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
%
DMSO
%
%
Tween 80
%
ddH2O
%
DMSO
+
%
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Mechanism of Action
| Targets/IC50/Ki |
Hedgehog [1]
(Cell-free assay) 3 nM
|
|---|---|
| In vitro |
Vismodegib (GDC-0449) targets the Hedgehog signaling pathway, blocking the activities of the Hedgehog-ligand cell surface receptors PTCH and/or SMO and suppressing Hedgehog signaling. This compound prevents multiple ATP-binding cassette (ABC) transporters and also blocks ABCG2, Pgp, and MRP1—important ABC transporters associated with MDR. It is a potent inhibitor of ABC transporters, ABCG2/BCRP and ABCB1/Pgp, and is a mild inhibitor of ABCC1/MRP1. In ABCG2-overexpressing HEK293 cells, it increases retention of the fluorescent ABCG2 substrate BODIPY and resensitizes these cells. In Madin-Darby canine kidney II cells engineered to overexpress Pgp or MRP1, it increases the retention of calcein-AM and resensitizes them. It also resensitizes human non-small cell lung carcinoma cells NCI-H460/par and NCI-H460/MX20, which overexpress ABCG2 in response to SN-38. The IC50 values for prevention of ABCG2 and Pgp are about 1.4 μM and 3.0 μM, respectively. [2] Additionally, it alters intracellular Ca2+ homeostasis and inhibits cell growth in resistant lung cancer cells. [3] |
| In vivo |
Vismodegib (GDC-0449) has been used to treat medulloblastoma in animal models. [2] It prevents the growth of primary pancreatic xenografts without non-specifically inhibiting pancreatic cell proliferation. Oral dosing of this compound causes tumor regressions in the Ptch(+/-) allograft model of medulloblastoma at doses ≥25 mg/kg and tumor growth inhibition at doses up to 92 mg/kg dosed twice daily in two ligand-dependent colorectal cancer models, D5123, and 1040830. Analysis of Hh pathway activity and PK/PD modeling reveals that it inhibits Gli1 with a similar IC50 in both the medulloblastoma and D5123 models (0.165 μM and 0.267 μM, respectively). Pathway modulation is linked to efficacy using an integrated PK/PD model revealing a steep relationship where > 50% of the activity of GDC-0449 is associated with >80% repression of the Hh pathway. [4] |
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | Fas / DR4 / DR5 / Cleaved PARP / Bcl-2 / Cleaved caspase-3 / PDGFRα p-GSK3β / GSK3β / p-Akt / Akt / Gli1 Gli1 / SOX2 / OCT4 p53 / Cyclin D1 / p21 Bcl-2 / Bax |
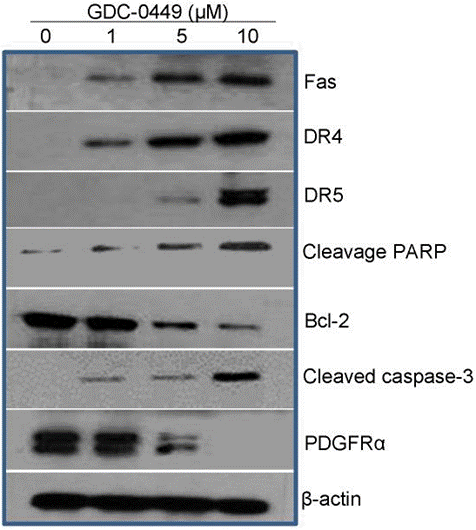
|
22087285 |
| Immunofluorescence | Gli1 / Gli2 |
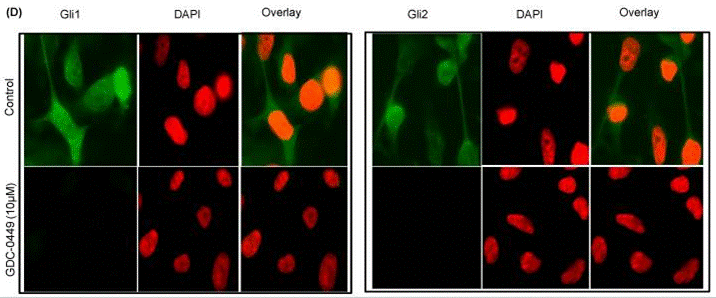
|
22087285 |
| Growth inhibition assay | Cell viability |
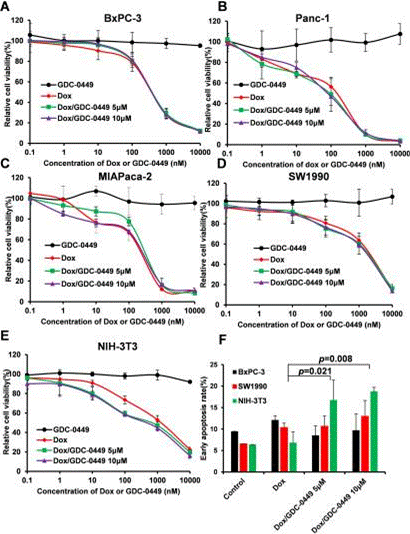
|
29042665 |
Clinical Trial Information
(data from https://clinicaltrials.gov, updated on 2024-05-22)
| NCT Number | Recruitment | Conditions | Sponsor/Collaborators | Start Date | Phases |
|---|---|---|---|---|---|
| NCT06344052 | Recruiting | Basal Cell Carcinoma |
Stamford Pharmaceuticals Inc. |
April 9 2024 | Phase 2 |
| NCT03610022 | Completed | Metastatic Basal Cell Carcinoma|Locally Advanced Basal Cell Carcinoma |
University Hospital Bordeaux |
September 3 2018 | Phase 4 |
| NCT03035188 | Completed | Basal Cell Carcinoma |
SRH Wald-Klinikum Gera GmbH |
January 2017 | Phase 2 |
| NCT02781389 | Completed | Basal Cell Carcinoma |
University Hospital Essen|OnkoDataMed GmbH |
April 29 2016 | -- |
| NCT02593760 | Completed | Myelofibrosis |
Hoffmann-La Roche |
January 25 2016 | Phase 1 |
| NCT02648048 | Completed | Idiopathic Pulmonary Fibrosis |
Hoffmann-La Roche |
January 15 2016 | Phase 1 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































