research use only
AMD3100 (Plerixafor) CXCR4 antagonist
Cat.No.S8030
Chemical Structure
Molecular Weight: 502.78
Quality Control
| Related Targets | Hedgehog/Smoothened PKA Adrenergic Receptor AChR 5-HT Receptor Histamine Receptor Dopamine Receptor Ras KRas GPR |
|---|---|
| Other CXCR Inhibitors | AZD5069 SB 225002 Reparixin (Repertaxin) WZ811 Navarixin (SCH-527123) LIT-927 AMG 487 SX-682 SB 265610 LY2510924 |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| CHOK1 | Function assay | Displacement of [125I]SDF1alpha from CCR2/CXCR4 expressed in CHOK1 cells, IC50 = 0.00004 μM. | 17715128 | |||
| CHOK1 | Function assay | Displacement of [125I]MCP1 from CCR2/CXCR4 expressed in CHOK1 cells, IC50 = 0.00009 μM. | 17715128 | |||
| CHOK1 | Function assay | Displacement of [125I]SDF1alpha from CXCR4 expressed in CHOK1 cells, IC50 = 0.00081 μM. | 17715128 | |||
| U87.CD4 | Antiviral assay | Antiviral activity against HIV1 NL43 infected in U87.CD4 cells expressing human CXCR4 H281A mutant, IC50 = 0.0019 μM. | 17599916 | |||
| U87.CD4 | Antiviral assay | Antiviral activity against HIV1 NDK infected in U87.CD4 cells expressing human CXCR4 H281A mutant, IC50 = 0.0019 μM. | 17599916 | |||
| MT4 | Antiviral assay | 5 days | Antiviral activity against HIV1 3B infected in human MT4 cells assessed as reduction in virus-induced cytopathic effect after 5 days by MTT assay, EC50 = 0.002 μM. | 26974376 | ||
| MT4 | Antiviral assay | 5 days | Antiviral activity against HIV2 ROD infected in human MT4 cells assessed as reduction in virus-induced cytopathic effect after 5 days by MTT assay, EC50 = 0.002 μM. | 26974376 | ||
| HEK293 | Antiviral assay | 2 days | Antiviral activity against T20-resistant HIV1 NL4-3 infected in HEK293 cells assessed as inhibition of viral replication after 2 days, IC50 = 0.0023 μM. | 19451305 | ||
| U87.CD4 | Antiviral assay | Antiviral activity against HIV1 clinical isolate 10 infected in U87.CD4 cells expressing human CXCR4 H281A mutant, IC50 = 0.0024 μM. | 17599916 | |||
| U87.CD4 | Antiviral assay | Antiviral activity against HIV1 clinical isolate 10 infected in U87.CD4 cells expressing human wild type CXCR4, IC50 = 0.003 μM. | 17599916 | |||
| PBMC | Function assay | Effective concentration of compound against HIV-1 89.6 strain in PBMC cells, EC50 = 0.0038 μM. | 14698189 | |||
| MT4 | Antiviral assay | 4 days | Antiviral activity against HIV1 3B infected in human MT4 cells assessed as inhibition of virus replication after 4 days by MTT assay, EC50 = 0.004 μM. | 20043638 | ||
| MT-4 | Function assay | Effective concentration against HIV-1(IIIB) replication in MT-4 cells, EC50 = 0.0042 μM. | 8568797 | |||
| HEK293 | Antiviral assay | 2 days | Antiviral activity against HIV1 NL4-3 infected in HEK293 cells assessed as inhibition of viral replication after 2 days, IC50 = 0.0046 μM. | 19451305 | ||
| HEK293 | Antiviral assay | 2 days | Antiviral activity against multidrug resistant HIV1 HXB2 infected in HEK293 cells assessed as inhibition of viral replication after 2 days, IC50 = 0.0053 μM. | 19451305 | ||
| MT-4 | Function assay | Effective concentration against HIV-2(ROD) replication in MT-4 cells, EC50 = 0.0059 μM. | 8568797 | |||
| CEM-CCRF | Function assay | 30 mins | Inhibition of PE-conjugated-12G5 anti-CXCR4 antibody binding to CXCR4 in human CEM-CCRF cells preincubated for 30 mins followed by antibody addition by FACS Canto II cytofluorometric analysis, IC50 = 0.006 μM. | 27571038 | ||
| HEK293 | Antiviral assay | 2 days | Antiviral activity against HIV1 HXB2 infected in HEK293 cells assessed as inhibition of viral replication after 2 days, IC50 = 0.0062 μM. | 19451305 | ||
| HEK293 | Antiviral assay | 2 days | Antiviral activity against NNRTI-resistant HIV1 HXB2 infected in HEK293 cells assessed as inhibition of viral replication after 2 days, IC50 = 0.007 μM. | 19451305 | ||
| U87.CD4 | Antiviral assay | Antiviral activity against HIV1 NDK infected in U87.CD4 cells expressing human wild type CXCR4, IC50 = 0.0076 μM. | 17599916 | |||
| MT-4 | Antiviral assay | Antiviral activity against HIV 1 RIN HIV 1 RIN harboring integrase gene infected in MT-4 cells assessed as inhibition of virus-induced cytopathic effect by MTT assay after 40 passages selected in presence of compound, EC50 = 0.008 μM. | 18378713 | |||
| MT-4 | Antiviral assay | Antiviral activity against HIV 1 RIN HIV 1 RIN harboring integrase gene infected in MT-4 cells assessed as inhibition of virus-induced cytopathic effect by MTT assay after 60 passages selected in presence of compound, EC50 = 0.008 μM. | 18378713 | |||
| MT-4 | Antiviral assay | Antiviral activity against HIV 1 RIN harboring integrase gene infected in MT-4 cells assessed as inhibition of virus-induced cytopathic effect by MTT assay after 20 passages selected in presence of compound, EC50 = 0.008 μM. | 18378713 | |||
| HEK293 | Antiviral assay | 2 days | Antiviral activity against NRTI-resistant HIV1 HXB2 infected in HEK293 cells assessed as inhibition of viral replication after 2 days, IC50 = 0.009 μM. | 19451305 | ||
| HEK293 | Antiviral assay | 2 days | Antiviral activity against PI-resistant HIV1 HXB2 infected in HEK293 cells assessed as inhibition of viral replication after 2 days, IC50 = 0.0092 μM. | 19451305 | ||
| U87.CD4 | Antiviral assay | Antiviral activity against HIV1 NL43 infected in U87.CD4 cells expressing human wild type CXCR4, IC50 = 0.014 μM. | 17599916 | |||
| MT-4 | Antiviral assay | Antiviral activity against HIV 1 RIN harboring integrase gene infected in MT-4 cells assessed as inhibition of virus-induced cytopathic effect by MTT assay, EC50 = 0.014 μM. | 18378713 | |||
| U87.CD4 | Antiviral assay | Antiviral activity against HIV1 NDK infected in U87.CD4 cells expressing human CXCR4 D171N mutant, IC50 = 0.017 μM. | 17599916 | |||
| CD4+ T | Function assay | Antagonist activity at CXCR4 in human CD4+ T cells assessed as inhibition of CXCL12-mediated cytosolic calcium level preincubated with compounds followed by CXCL12 stimulation by calcium 4 dye-based FLIPR assay, IC50 = 0.018 μM. | 29494843 | |||
| U87.CD4 | Antiviral assay | Antiviral activity against HIV1 clinical isolate 10 infected in U87.CD4 cells expressing human CXCR4 D171N mutant, IC50 = 0.019 μM. | 17599916 | |||
| MT4 | Antiviral assay | 5 days | Antiviral activity against T-cell line-tropic HIV1 NL4-3 infected in human MT4 cells assessed as inhibition of virus-induced cytopathogenicity after 5 days by MTT assay in presence of 5 uM of chloroquine, EC50 = 0.025 μM. | 26094944 | ||
| Jurkat | Function assay | Antagonist activity at CXCR4 in human Jurkat cells assessed as inhibition of SDF1-induced cell migration, IC50 = 0.0274 μM. | 19188071 | |||
| MT-4 | Antiviral assay | Antiviral activity against HIV 1 3B harboring integrase L34M mutant infected in MT-4 cells assessed as inhibition of virus-induced cytopathic effect by MTT assay selected after 40 passages in presence of compound, EC50 = 0.028 μM. | 18378713 | |||
| MT4 | Antiviral assay | 5 days | Antiviral activity against T-cell line-tropic HIV1 NL4-3 infected in human MT4 cells assessed as inhibition of virus-induced cytopathogenicity after 5 days by MTT assay, EC50 = 0.032 μM. | 26094944 | ||
| MT-4 | Antiviral assay | Antiviral activity against HIV 1 3B infected in MT-4 cells assessed as inhibition of virus-induced cytopathic effect by MTT assay, EC50 = 0.034 μM. | 18378713 | |||
| MT4 | Antiviral assay | 5 days | Antiviral activity against T-cell line-tropic HIV1 NL4-3 infected in human MT4 cells assessed as inhibition of virus-induced cytopathogenicity after 5 days by MTT assay in presence of 2.5 uM of chloroquine, EC50 = 0.039 μM. | 26094944 | ||
| U87.CD4 | Antiviral assay | Antiviral activity against HIV1 NL43 infected in U87.CD4 cells expressing human CXCR4 D171N mutant, IC50 = 0.046 μM. | 17599916 | |||
| MT-4 | Antiviral assay | Antiviral activity against HIV 1 3B harboring integrase E92Q S230N double mutant infected in MT-4 cells assessed as inhibition of virus-induced cytopathic effect by MTT assay selected after 20 passages in presence of compound, EC50 = 0.049 μM. | 18378713 | |||
| MT-4 | Antiviral assay | Antiviral activity against HIV 1 3B harboring integrase E92Q, S230N and L34M triple mutant infected in MT-4 cells assessed as inhibition of virus-induced cytopathic effect by MTT assay selected after 60 passages in presence of compound, EC50 = 0.056 μM. | 18378713 | |||
| MT-4 | Function assay | Effective concentration of compound against HIV-1 IIIB strain in MT-4 cells, EC50 = 0.065 μM. | 14698189 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 H281A mutant expressed in HEK293 cells, IC50 = 0.0727 μM. | 19451305 | |||
| IR983F | Function assay | Displacement of [125I]CXCL12 from CXCR4 in rat IR983F cells, IC50 = 0.108 μM. | 19053768 | |||
| CEM-SS | Function assay | Effective concentration of compound against HIV-1 LAI strain in CEM-SS cells, EC50 = 0.127 μM. | 14698189 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 V196A mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.14 μM. | 17599916 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 D181A mutant expressed in HEK293 cells, IC50 = 0.1437 μM. | 19451305 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 7 of human CXCR4 H281A mutant expressed in COS7 cells, Ki = 0.16 μM. | 17599916 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 F172A mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.17 μM. | 17599916 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 V280A mutant expressed in HEK293 cells, IC50 = 0.1753 μM. | 19451305 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 H281A mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.19 μM. | 17599916 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 V112A mutant expressed in HEK293 cells, IC50 = 0.1966 μM. | 19451305 | |||
| COS7 | Function assay | Antagonist activity at human wild type CXCR4 expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.22 μM. | 17599916 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 E275A mutant expressed in HEK293 cells, IC50 = 0.2356 μM. | 19451305 | |||
| CEM | Function assay | Displacement of [125I]CXCL12 from CXCR4 in human CEM cells, IC50 = 0.245 μM. | 19053768 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 V99A mutant expressed in HEK293 cells, IC50 = 0.2585 μM. | 19451305 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 H203A mutant expressed in HEK293 cells, IC50 = 0.259 μM. | 19451305 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 I284A mutant expressed in HEK293 cells, IC50 = 0.2658 μM. | 19451305 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to wild type CXCR4 expressed in HEK293 cells, IC50 = 0.2891 μM. | 19451305 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 H203A mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.29 μM. | 17599916 | |||
| COS7 | Function assay | Antagonist activity at transmembrane domain 6 of human CXCR4 I259A mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.29 μM. | 17599916 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 T287A mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.29 μM. | 17599916 | |||
| HPBALL | Function assay | 3 hrs | Displacement of 12G5-CXCL12 from CXCR4 in human HPBALL cells after 3 hrs by FACS analysis, IC50 = 0.29 μM. | 29494843 | ||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 H113A mutant expressed in HEK293 cells, IC50 = 0.2964 μM. | 19451305 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 W283A mutant expressed in HEK293 cells, IC50 = 0.3002 μM. | 19451305 | |||
| CEM-SS | Antiviral assay | Antiviral activity against HIV1 LAI in human CEM-SS cells assessed as inhibition of viral replication, IC50 = 0.32 μM. | 19356827 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 L120F mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.33 μM. | 17599916 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 Q200W mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.4 μM. | 17599916 | |||
| MT4 | Antiviral assay | Antiviral activity against HIV1 3B in human MT4 cells assessed as inhibition of viral replication, IC50 = 0.41 μM. | 19356827 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 E277A mutant expressed in HEK293 cells, IC50 = 0.4695 μM. | 19451305 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 5 of human CXCR4 Q200A mutant expressed in COS7 cells, Ki = 0.56 μM. | 17599916 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 H281A mutant expressed in HEK293 cells, IC90 = 0.5722 μM. | 19451305 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 I259W mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.63 μM. | 17599916 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 Y255A mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.66 μM. | 17599916 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 H113A mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.74 μM. | 17599916 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 D181A mutant expressed in HEK293 cells, IC90 = 0.7956 μM. | 19451305 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 V280A mutant expressed in HEK293 cells, IC90 = 0.8212 μM. | 19451305 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 V112A mutant expressed in HEK293 cells, IC90 = 0.8213 μM. | 19451305 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 H203A mutant expressed in HEK293 cells, IC90 = 0.8606 μM. | 19451305 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from human wild type CXCR4 expressed in COS7 cells, Ki = 0.89 μM. | 17599916 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 Q200A mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 0.93 μM. | 17599916 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to CXCR4 E275A mutant expressed in HEK293 cells, IC90 = 0.9302 μM. | 19451305 | |||
| HEK293 | Function assay | Inhibition of Mab 12G5 binding to wild type CXCR4 expressed in HEK293 cells, IC90 = 0.9711 μM. | 19451305 | |||
| MDA-MB-231 | Function assay | 10 mins | Displacement of biotinylated TN14003 from CXCR4 CXCL12 binding domain in human MDA-MB-231 cells preincubated for 10 mins followed by biotinylated TN14003 addition measured after 30 mins using streptavidin-conjugated rhodamine by fluorescence microscopic a, EC = 1 μM. | 27179215 | ||
| MDA-MB-231 | Function assay | 10 mins | Inhibition of biotinylated TN14003 binding to CXCR4 in human MDA-MB-231 cells preincubated for 10 mins followed by TN14003 addition measured after 30 mins by rhodamine dye-based microscopic analysis, EC = 1 μM. | 29529500 | ||
| MDA-MB-231 | Function assay | 10 mins | Displacement of biotinylated-TN14003 from CXCR4 in human MDA-MB-231 cells assessed as reduction in fluorescence preincubated for 10 mins followed by biotinylated-TN14003 addition measured after 30 mins by streptavidin-rhodamine staining based microscopic , EC = 1 μM. | 27914361 | ||
| MDA-MB-231 | Function assay | 10 mins | Inhibition of biotinylated TN14003 binding to CXCR4 in human MDA-MB-231 cells assessed as reduction in fluorescence preincubated for 10 mins followed by biotinylated-TN14003 addition measured after 30 mins by streptavidin-rhodamine staining based immunofl, EC = 1 μM. | 28521261 | ||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 7 of human CXCR4 I284A mutant expressed in COS7 cells, Ki = 1.1 μM. | 17599916 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 I284A mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 1.2 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 6 of human CXCR4 I259A mutant expressed in COS7 cells, Ki = 1.5 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 5 of human CXCR4 H203A mutant expressed in COS7 cells, Ki = 1.8 μM. | 17599916 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 D171N mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 1.8 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 3 of human CXCR4 H113A mutant expressed in COS7 cells, Ki = 2 μM. | 17599916 | |||
| U87.CD4 | Antiviral assay | Antiviral activity against HIV1 clinical isolate 10 infected in U87.CD4 cells expressing human CXCR4 D262N mutant, IC50 = 2.541 μM. | 17599916 | |||
| U87.CD4 | Antiviral assay | Antiviral activity against HIV1 clinical isolate 10 infected in U87.CD4 cells expressing human CXCR4 D171ND262N mutant, IC50 = 2.694 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 5 of human CXCR4 Q200W mutant expressed in COS7 cells, Ki = 2.7 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 5 of human CXCR4 G207F mutant expressed in COS7 cells, Ki = 2.7 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 5 of human CXCR4 G207W mutant expressed in COS7 cells, Ki = 3 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 7 of human CXCR4 T287A mutant expressed in COS7 cells, Ki = 3.1 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 4 of human CXCR4 F174A mutant expressed in COS7 cells, Ki = 3.2 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from extracellular loop 2 of human CXCR4 D182A mutant expressed in COS7 cells, Ki = 3.2 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 6 of human CXCR4 Y256A mutant expressed in COS7 cells, Ki = 4 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from extracellular loop 2 of human CXCR4 N176A mutant expressed in COS7 cells, Ki = 4.1 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 5 of human CXCR4 V196A mutant expressed in COS7 cells, Ki = 4.6 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 6 of human CXCR4 I259W mutant expressed in COS7 cells, Ki = 4.6 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 4 of human CXCR4 F172A mutant expressed in COS7 cells, Ki = 4.7 μM. | 17599916 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 D262N mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 4.7 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 6 of human CXCR4 S263A mutant expressed in COS7 cells, Ki = 5.4 μM. | 17599916 | |||
| CCRF-CEM | Function assay | 30 mins | Inhibition of anti-CXCR4 PE antibody clone 12G5 binding to CXCR4 in human CCRF-CEM cells preincubated for 30 mins followed by anti-CXCR4 PE antibody clone 12G5 addition measured after 30 mins by flow cytometric method, IC50 = 6.2 μM. | 29125295 | ||
| COS7 | Function assay | Antagonist activity at human CXCR4 E288A mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 6.4 μM. | 17599916 | |||
| MT4 | Cytotoxicity assay | Cytotoxicity against human MT4 cells by MTT assay, CC50 = 6.5 μM. | 19356827 | |||
| COS7 | Function assay | Antagonist activity at human CXCR4 A175F mutant expressed in COS7 cells coexpressing G protein Gqi4myr assessed as inhibition of CXCL12-induced phosphatidylinositol turnover, Ki = 8.5 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 6 of human CXCR4 Y255A mutant expressed in COS7 cells, Ki = 9.7 μM. | 17599916 | |||
| MT-4 | Function assay | Concentration required to inhibit syncytia formation by 50% on HIV-1 infected MT-4 cells, EC50 = 10 μM. | 9925728 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 4 of human CXCR4 D171N mutant expressed in COS7 cells, Ki = 13 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from extracellular loop 2 of human CXCR4 D187A mutant expressed in COS7 cells, Ki = 14 μM. | 17599916 | |||
| HL60 | Function assay | Displacement of [125I]SDF1alpha from CXCR4 in human HL60 cells, IC50 = 15.2 μM. | 19188071 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from extracellular loop 2 of human CXCR4 A175F mutant expressed in COS7 cells, Ki = 36 μM. | 17599916 | |||
| COS7 | Function assay | Displacement of [125I]12G5 antibody from transmembrane domain 6 of human CXCR4 D262N mutant expressed in COS7 cells, Ki = 46 μM. | 17599916 | |||
| COS, TZM-bl | Function assay | Inhibition of HIV1 NL4-3 envelope glycoprotein 120-mediated membrane fusion between virus-transfected african green monkey COS cells and human TZM-bl cells by luciferase-based cell-cell fusion assay in presence of IC9564 | 17954689 | |||
| CXCR4+/CD4+/U87 | Function assay | Inhibition of HIV1 92TH594 infected CXCR4+/CD4+/U87 cells to assess co-receptor tropism as luciferase activity | 17116663 | |||
| CXCR4+/CD4+/U87 | Function assay | Inhibition of HIV1 HXB2 infected CXCR4+/CD4+/U87 cells to assess co-receptor tropism as luciferase activity | 17116663 | |||
| U87 | Function assay | 1000 nM | Antagonist activity at CXCR4 in human U87 cells assessed as inhibition of SDF1-induced modulation of cAMP production at 1000 nM by TR-FRET assay | 17958344 | ||
| CHOK1 | Function assay | Induction of [125I]MCP1 dissociation from CCR2/CXCR4 expressed in CHOK1 cells by non-equilibrium binding assay | 17715128 | |||
| CHOK1 | Function assay | Antagonist activity at CXCR4 expressed in CHOK1 cells assessed as inhibition of SDF1-alpha-induced signaling by aequorin-based assay | 17715128 | |||
| CHOK1 | Function assay | Antagonist activity at CCR2/CXCR4 expressed in CHOK1 cells assessed as inhibition of SDF1-alpha-induced signaling by aequorin-based assay | 17715128 | |||
| CHOK1 | Function assay | Induction of [125I]SDF1alpha dissociation from CXCR4 expressed in CHOK1 cells by non-equilibrium binding assay | 17715128 | |||
| U87.CD4 | Function assay | Antagonist activity at human CXCR4 H281A mutant expressed in U87.CD4 cells assessed as inhibition of CXCL12-induced calcium mobilization | 17599916 | |||
| MT2 | Antiviral assay | 1 ug/mL | 4 days | Antiviral activity against HIV1 3B infected in human MT2 cells assessed as inhibition of viral p24 antigen production at 1 ug/mL after 4 days by ELISA | 21168336 | |
| MOLT4 | Function assay | 1000 nM | Inhibition of Mab 12G5 binding to CXCR4 expressed in human MOLT4 cells at 1000 nM by FACS analysis | 19451305 | ||
| TZM-bl | Antiviral assay | 100 uM | 24 hrs | Antiviral activity against HIV NL-Lai infected in human TZM-bl cells assessed as inhibition of viral infection at 100 uM treated before viral infection measured after 24 hrs by luciferase assay | 21783371 | |
| Jurkat | Function assay | 0.01 to 100 uM | Inhibition of CXCR4 in human Jurkat cells assessed as reduction in HIV-Nef-M1-induced mitochondrial membrane depolarization at 0.01 to 100 uM by JC1 dye based fluorescence depolarization assay | 26191361 | ||
| MDA-MB-231 | Function assay | 100 nM | 24 hrs | Antagonist activity at CXCR4-mediated chemotaxis in human MDA-MB-231 cells assessed as inhibition of CXCL12-induced cell invasion at 100 nM after 24 hrs by crystal violet staining-based microscopic matrigel assay | 29494843 | |
| TZM-bl | Antiviral assay | 1 hr | Antiviral activity against HIV1 HXB2 pseudovirus infected in human TZM-bl cells assessed as inhibition of viral entry treated 1 hr post infection measured after 48 hrs by luciferase reporter gene assay | 28266845 | ||
| CXCR4+ | Function assay | 6 mg/kg | 2 hrs | Induction hematopoietic stem cell mobilization in C57BL/6 mouse assessed as increase in CXCR4+ cells in blood at 6 mg/kg, sc measured after 2 hrs by APC-conjugated anti-CXCR4-staining based flow cytometry relative to vehicle control | 29314840 | |
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
Ethanol : 100 mg/mL Water : 15 mg/mL
DMSO
: Insoluble
|
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 502.78 | Formula | C28H54N8 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 110078-46-1 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | JM 3100, SID791 | Smiles | C1CNCCNCCCN(CCNC1)CC2=CC=C(C=C2)CN3CCCNCCNCCCNCC3 | ||
Mechanism of Action
| Targets/IC50/Ki |
CXCL12
(Cell-free assay) 5.7 nM
CXCR4
(Cell-free assay) 44 nM
|
|---|---|
| In vitro |
Plerixafor (AMD3100) inhibits CXCL12-mediated chemotaxis with a potency lightly better than its affinity for CXCR4. It also antagonizes SDF-1/CXCL12 ligand binding with an IC50 of 651 nM. This compound inhibits SDF-1 mediated GTP-binding, SDF-1 mediated calcium flux and SDF-1 stimulated chemotaxis with IC50 of 27 nM, 572 nM and 51 nM, respectively. It does not inhibit calcium flux against cells expressing CXCR3, CCR1, CCR2b, CCR4, CCR5 or CCR7 when stimulated with their cognate ligands, nor does it inhibit receptor binding of LTB4. On its own, it does not induce a calcium flux in the CCRF–CEM cells, which express multiple GPCRs including CXCR4, CCR4 and CCR7. |
| Kinase Assay |
Receptor binding assays
|
|
For the competition binding studies against CXCR4, Plerixafor (AMD3100) is incubated at a concentration range for 3 hours at 4 °C in binding buffer (PBS containing 5 mM MgCl2, 1 mM CaCl2, 0.25% BSA, pH 7.4) with 5 × 105 CCRF–CEM cells and 100 pM 125I-SDF-1α (2200 Ci/mmol) in Milipore DuraporeTM filter plates. Unbound 125I-SDF-1α is removed by washing with cold 50 mM HEPES, 0.5 M NaCl pH 7.4. The competition binding assay against BLT1 is performed on membranes from CHO-S cells expressing recombinant BLT1. The membranes are prepared by mechanical cell lysis followed by high speed centrifugation, re-suspended in 50 mM HEPES, 5 mM MgCl2 buffer and flash frozen. This compound is incubated with the membrane preparation for 1 hour at room temperature in an assay mixture containing 50 mM Tris, pH 7.4, 10 mM MgCl2, 10 mM CaCl2, 4 nM LTB4 mixed with 1 nM 3H-LTB4 (195.0 Ci/mmol) and 8 μg membrane. The unbound 3H-LTB4 is separated by filtration on Millipore Type GF-C filter plates.
|
|
| In vivo |
In diabetic mice, a single topical application of Plerixafor (AMD3100) promotes wound healing by increasing cytokine production, mobilizing bone marrow EPCs, and enhancing the activity of fibroblasts and monocytes/macrophages, thereby increasing both angiogenesis and vasculogenesis. Cohorts of mice are administered with PBS, IGF1, PDGF, SCF, or VEGF for five consecutive days and this compound on the 5th day. The number and size of the colonies are highest in IGF1 plus it injected mice compared to PDGF, SCF and VEGF treated groups, in combination with Plerixafor. |
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Immunofluorescence | CXCR4 β-arrestin2 |
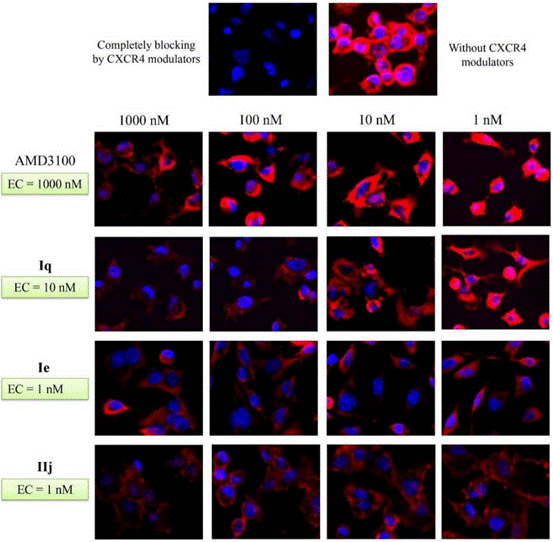
|
28521261 |
Clinical Trial Information
(data from https://clinicaltrials.gov, updated on 2024-05-22)
| NCT Number | Recruitment | Conditions | Sponsor/Collaborators | Start Date | Phases |
|---|---|---|---|---|---|
| NCT05421416 | Not yet recruiting | Stem Cell Transplant Complications |
AHS Cancer Control Alberta |
April 1 2024 | Phase 2 |
| NCT05343572 | Recruiting | Asherman Syndrome|Atrophic Endometrium|Recurrent Implantation Failure |
Hugh Taylor|Yale University |
November 1 2023 | Early Phase 1 |
| NCT05844527 | Recruiting | Wound of Skin|Abdominal Wound |
MedRegen LLC |
November 20 2023 | Phase 2 |
| NCT05411575 | Withdrawn | COVID-19 Acute Respiratory Distress Syndrome|COVID-19 |
4Living Biotech|4P-Pharma |
July 19 2022 | Phase 2 |
| NCT05445128 | Terminated | Sickle Cell Disease |
Ensoma|bluebird bio |
June 24 2022 | Phase 2 |
| NCT05835726 | Recruiting | Multiple Myeloma|Autologous Stem Cell Transplantation|Leukapheresis |
Fondazione Policlinico Universitario Agostino Gemelli IRCCS |
January 1 2022 | -- |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.
Frequently Asked Questions
Question 1:
How about the half-life of it (Cat S8030)?
Answer:
Its biological half-life is 3-5 hours: https://en.wikipedia.org/wiki/Plerixafor.






































