research use only
RG108 DNA Methyltransferase inhibitor
Cat.No.S2821
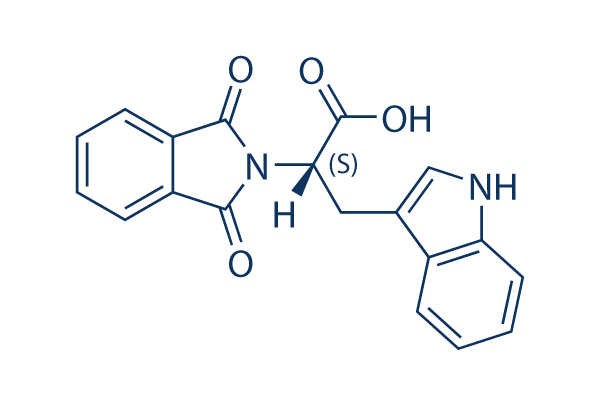
Chemical Structure
Molecular Weight: 334.33
Quality Control
| Related Targets | HDAC JAK BET Histone Methyltransferase PKC PARP HIF PRMT EZH2 AMPK |
|---|---|
| Other DNA Methyltransferase Inhibitors | SGI-1027 Zebularine (NSC 309132) GSK3685032 Gamma-Oryzanol CM272 Bobcat339 β-thujaplicin GSK-3484862 DC-05 2'-Deoxy-5-Fluorocytidine |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| human THP1 cells | Function assay | Inhibition of ACAT-mediated esterified cholesterol accumulation in human THP1 cells exposed to acetyl-LDL during differentiation assessed as effect on foam cell formation, IC50=1.5 μM | 18620381 | |||
| human HepG2 cells | Function assay | Inhibition of ACAT in human HepG2 cells, IC50=0.479 μM | 19167888 | |||
| rat macrophages | Function assay | 24 h | Inhibition of ACAT in rat macrophages assessed as incorporation of extracellular [3H]-oleic acid-BSA complex into the intracellular cholesteryl ester after 24 hrs, IC50=0.479 μM | 19464189 | ||
| MCF7 | Apoptosis assay | 10 uM | 6 hrs | Induction of apoptosis in human MCF7 cells assessed as apoptotic cells at 10 uM after 6 hrs by FITC-Annexin V/propidium iodide staining-based FACS analysis relative to control | 25801160 | |
| MCF7 | Apoptosis assay | 10 uM | 6 hrs | Induction of apoptosis in human MCF7 cells assessed as dead cells at 10 uM after 6 hrs by FITC-Annexin V/propidium iodide staining-based FACS analysis relative to control | 25801160 | |
| MCF7 | Function assay | 100 uM | 1 hr | Competitive inhibition of DNMT1 in (S)-Methyl 2-(2,5-dioxo-2,5-dihydro-1H-pyrrol-1-yl)-3-(5-(prop-2-yn-1-yloxy)-1H-indol-3-yl)propanoate labeled human MCF7 cells at 100 uM preincubated for 1 hr followed by cell labeling for 30 mins and clicked with TER-PE | 25801160 | |
| MCF7 | Function assay | 50 to 500 uM | 1 hr | Competitive inhibition of DNMT1 in (S)-Methyl 2-(2,5-dioxo-2,5-dihydro-1H-pyrrol-1-yl)-3-(5-(prop-2-yn-1-yloxy)-1H-indol-3-yl)propanoate labeled human MCF7 cells at 50 to 500 uM preincubated for 1 hr followed by cell labeling for 1 hr and clicked with TER | 25801160 | |
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SK-N-MC cells | 29435139 | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 66 mg/mL
(197.4 mM)
Ethanol : 66 mg/mL Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 334.33 | Formula | C19H14N2O4 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 48208-26-0 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N-Phthalyl-L-tryptophan | Smiles | C1=CC=C2C(=C1)C(=O)N(C2=O)C(CC3=CNC4=CC=CC=C43)C(=O)O | ||
Mechanism of Action
| Targets/IC50/Ki |
DNA methyltransferase
(Cell-free assay) 115 nM
|
|---|---|
| In vitro |
RG108 effectively blocks DNA methyltransferases in vitro and does not cause covalent enzyme trapping in human cell lines. Incubation of cells with low micromolar concentrations of this compound results in significant demethylation of genomic DNA without any detectable toxicity. Intriguingly, it causes demethylation and reactivation of tumor suppressor genes, but it does not affect the methylation of centromeric satellite sequences. In another study, the synthesis and in vitro analysis of a biotinylated conjugate of this chemical is investigated to evaluate the interactions with DNA methyltransferase enzymes. In a recent study, it is shown this compound can significantly reduce the DNA methyltransferases activity in SM derived iPS cells as compared to the native SMs.
|
| Kinase Assay |
In vitro methylation assay
|
|
The substrate DNA for the in vitro methylation assay is a 798 bp fragment (−423/+375 relative to the initiation codon) from the promoter region of the human p16Ink4a gene. The methylation reaction contains 350 to 400 ng substrate DNA and 4 units of M.SssI methylase (0.5 μM) in a final volume of 50 μL. This compound is added to final concentrations of 10, 100, 200, and 500 μM, respectively. Reactions are done at 37 °C for 2 hours. After completion, the reaction is inactivated at 65 °C for 15 minutes and the DNA is purified using PCR Purification kit. Three hundred nanograms of purified DNA is digested for 3 hours at 60 °C with 30 units of BstUI and analyzed on 2% Tris-borate EDTA agarose gels.
|
References |
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































