research use only
Xevinapant (AT406) IAP antagonist
Cat.No.S2754
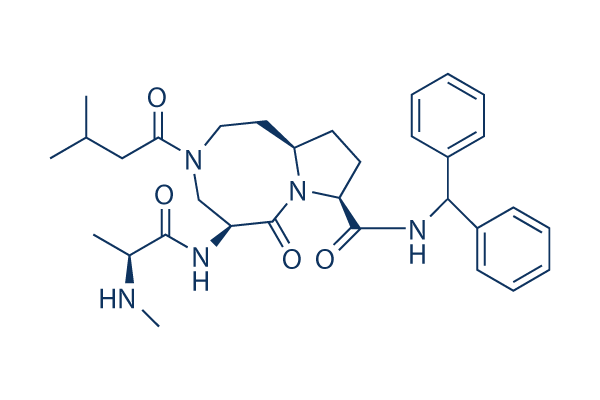
Chemical Structure
Molecular Weight: 561.71
Quality Control
| Related Targets | Bcl-2 Caspase PD-1/PD-L1 Ferroptosis p53 Apoptosis related Synthetic Lethality STAT TNF-alpha Ras |
|---|---|
| Other IAP Inhibitors | BV6 SM-164 Birinapant (TL32711) LCL161 GDC-0152 AZD5582 Tolinapant (ASTX660) |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| EVSA-T | Cytotoxicity assay | 72 hrs | Cytotoxicity against human sensitive EVSA-T cells assessed as inhibition of cell growth after 72 hrs by Alamar Blue assay, EC50 = 0.0021 μM. | 26218264 | ||
| MDA-MB-231 | Cytotoxicity assay | 72 hrs | Cytotoxicity against human sensitive MDA-MB-231 cells assessed as inhibition of cell growth after 72 hrs by Alamar Blue assay, EC50 = 0.019 μM. | 26218264 | ||
| HEK293 | Function assay | 2 hrs | Antagonist activity at full-length FLAG-tagged XIAP (unknown origin) transfected in HEK293 cells assessed as inhibition of interaction with caspase 9 after 2 hrs by immunoprecipitation assay, EC50 = 0.034 μM. | 26218264 | ||
| SKOV3 | Growth inhibition assay | 4 days | Growth inhibition of human SKOV3 cells after 4 days by WST8 assay, IC50 = 0.142 μM. | 21443232 | ||
| MDA-MB-231 | Growth inhibition assay | 4 days | Growth inhibition of human MDA-MB-231 cells after 4 days by WST8 assay, IC50 = 0.144 μM. | 21443232 | ||
| MDA-MB-231 | Function assay | 10 nM | Induction of degradation of cIAP1 in human MDA-MB-231 cells at 10 nM by Western blot analysis | 21443232 | ||
| MDA-MB-231 | Function assay | 100 nM | 15 mins | Induction of degradation of cIAP1 in human MDA-MB-231 cells at 100 nM after 15 mins by Western blot analysis | 21443232 | |
| MDA-MB-231 | Apoptosis assay | 1.5 uM | 12 hrs | Induction of apoptosis in human MDA-MB-231 cells assessed as activation of caspase-3 activity at 1.5 uM after 12 hrs by Western blot analysis | 21443232 | |
| MDA-MB-231 | Apoptosis assay | 1.5 uM | 12 hrs | Induction of apoptosis in human MDA-MB-231 cells assessed as activation of PARP cleavage at 1.5 uM after 12 hrs by Western blot analysis | 21443232 | |
| MDA-MB-231 | Antitumor assay | 30 mg/kg | 2 weeks | Antitumor activity against human MDA-MB-231 cells xenografted in SCID mouse assessed inhibition of tumor growth at 30 mg/kg, po administered daily for 5 days/week for 2 weeks | 21443232 | |
| MDA-MB-231 | Antitumor assay | 100 mg/kg | 2 weeks | Antitumor activity against human MDA-MB-231 cells xenografted in SCID mouse assessed inhibition of tumor growth at 100 mg/kg, po administered daily for 5 days/week for 2 weeks | 21443232 | |
| MDA-MB-231 | Antitumor assay | 100 mg/kg | every 3 days | Antitumor activity against human MDA-MB-231 cells xenografted in Balb/c SCID mouse assessed as tumor growth inhibition at 100 mg/kg, po qd measured every 3 days | 26218264 | |
| DAOY | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for DAOY cells | 29435139 | |||
| SJ-GBM2 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SJ-GBM2 cells | 29435139 | |||
| A673 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for A673 cells | 29435139 | |||
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SK-N-MC cells | 29435139 | |||
| BT-37 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for BT-37 cells | 29435139 | |||
| NB-EBc1 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for NB-EBc1 cells | 29435139 | |||
| Saos-2 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for Saos-2 cells | 29435139 | |||
| LAN-5 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for LAN-5 cells | 29435139 | |||
| BT-12 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for BT-12 cells | 29435139 | |||
| OHS-50 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for OHS-50 cells | 29435139 | |||
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for SK-N-MC cells | 29435139 | |||
| TC32 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Confirmatory screen for TC32 cells | 29435139 | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 112 mg/mL
(199.39 mM)
Ethanol : 112 mg/mL Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 561.71 | Formula | C32H43N5O4 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 1071992-99-8 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | ARRY-334543, Debio1143, SM-406 | Smiles | CC(C)CC(=O)N1CCC2CCC(N2C(=O)C(C1)NC(=O)C(C)NC)C(=O)NC(C3=CC=CC=C3)C4=CC=CC=C4 | ||
Mechanism of Action
| Targets/IC50/Ki |
cIAP1-BIR3
(cell-free assay) 1.9 nM(Ki)
cIAP2-BIR3
(cell-free assay) 5.1 nM(Ki)
XIAP-BIR3
(cell-free assay) 66.4 nM(Ki)
|
|---|---|
| In vitro |
AT-406 is a Smac mimetic and appears to mimic closely the AVPI peptide in both hydrogen bonding and hydrophobic interactions with XIAP, with additional hydrophobic contacts with W323 of XIAP. AT-406 is more sensitive to these IAPs than Smac AVPI peptide with 50-100 fold binding affinities. AT-406 (at 1 μM) completely restores the activity of caspase-9, which is suppressed by 500 nM XIAP BIR3 in a cell-free system. In MDA-MB-231 cell, AT-406 induces rapid cellular cIAP1 degradation and also pulls down the cellular XIAP protein. AT-406 effectively inhibits lots of human cancer cell lines and shows IC50 of 144 and 142 nM in MDA-MB-231 cell and SK-OV-3 ovarian cell, with low toxicity against normal-like human breast epithelial MCF-12F cells and primary human normal prostate epithelial cells. AT-406 induces apoptosis in MDA-MB-231 cell by inducing activation of caspase-3 and cleavage of PARP.
|
| Kinase Assay |
Fluorescence Polarization Based Assays for XIAP, cIAP1, and cIAP2 BIR3 Proteins
|
|
FL-AT-406 (the fluorescently tagged AT-406) is employed to develop a set of new FP assays for determination of the binding affinities of Smac mimetics to XIAP, cIAP-1, and cIAP-2 BIR3 proteins. The Kd value of FL-AT-406 to each IAP protein is determined by titration experiments using a fixed concentration of FL-AT-406 and different concentrations of the protein up to full saturation. Fluorescence polarization values are measured using an Infinite M-1000 plate reader in Microfluor 2 96-well, black, round-bottom plates. To each well, FL-AT-406 (2, 1, and 1 nM for experiments with XIAP BIR3, cIAP-1 BIR3, and cIAP-2 BIR3, respectively) and different concentrations of the protein are added to a final volume of 125 μL in the assay buffer (100 mM potassium phosphate, pH 7.5, 100 μg/mL bovine γ-globulin, 0.02% sodium azide, with 4% DMSO). Plates are mixed and incubated at room temperature for 2-3 hours with gentle shaking. The polarization values in millipolarization units (mP) are measured at an excitation wavelength of 485 nm and an emission wavelength of 530 nm. Equilibrium dissociation constants (Kd) are then calculated by fitting the sigmoidal dose-dependent FP increases as a function of protein concentrations using Graphpad Prism 5.0 software. In competitive binding experiments for XIAP3 BIR3, AT-406 is incubated with 20 nM XIAP BIR3 protein and 2 nM FL-AT-406 in the assay buffer (100 mM potassium phosphate, pH 7.5; 100 μg/mL bovine γ-globulin; 0.02% sodium azide). In competitive binding experiments for cIAP1 BIR3 protein, 3 nM protein and 1 nM FL-AT-406 are used. In competitive binding experiments for cIAP2 BIR3, 5 nM protein and 1 nM FL-AT-406 are used. For each competitive binding experiment, polarization values are measured after 2-3 hours of incubation using an Infinite M-1000 plate reader. The IC50 value, the inhibitor concentration at which 50% of the bound tracer is displaced, is determined from the plot using nonlinear least-squares analysis. Curve fitting is performed using the PRISM software. A Ki value for AT-406 is calculated.
|
|
| In vivo |
AT-406 has good pharmacokinetic (PK) properties and oral bioavailability in mice, rats, non-human primates, and dogs. In the MDA-MB-231 xenograft, AT-406 effectively induces cIAP1 degradation and processing of procaspase-8, cleavage of PARP in tumor tissues at 100 mg/kg with well toleration even at 200 mg/kg. AT-406 induces significant tumor growth inhibition with p of 0.0012 at 100 mg/kg.
|
References |
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | cIAP-1 / XIAP / Mcl-1 / pS6K1 / Cleaved PARP |
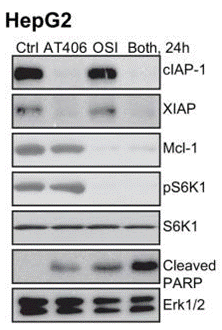
|
28036295 |
| Growth inhibition assay | Cell viability |
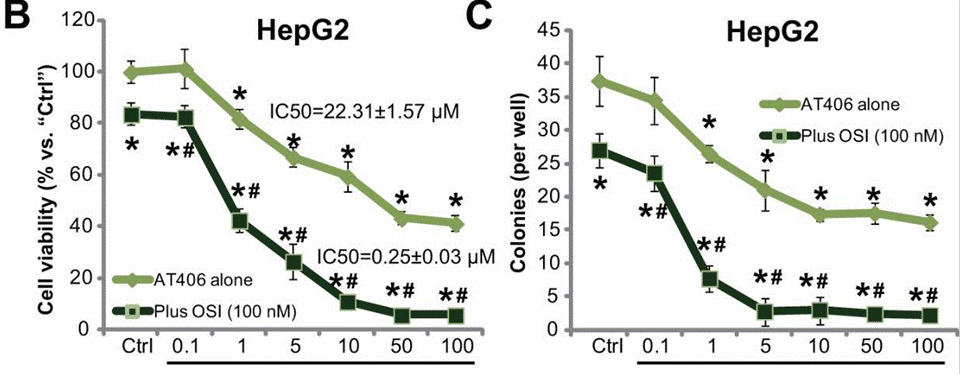
|
28036295 |
Clinical Trial Information
(data from https://clinicaltrials.gov, updated on 2024-05-22)
| NCT Number | Recruitment | Conditions | Sponsor/Collaborators | Start Date | Phases |
|---|---|---|---|---|---|
| NCT01265199 | Terminated | Acute Myelogenous Leukemia (AML) |
Ascenta Therapeutics|The Leukemia and Lymphoma Society |
February 2011 | Phase 1 |
| NCT01078649 | Completed | Cancer|Solid Tumors|Lymphoma|Malignancy |
Debiopharm International SA |
March 29 2010 | Phase 1 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































