research use only
SB743921 HCl Kinesin inhibitor
Cat.No.S2182
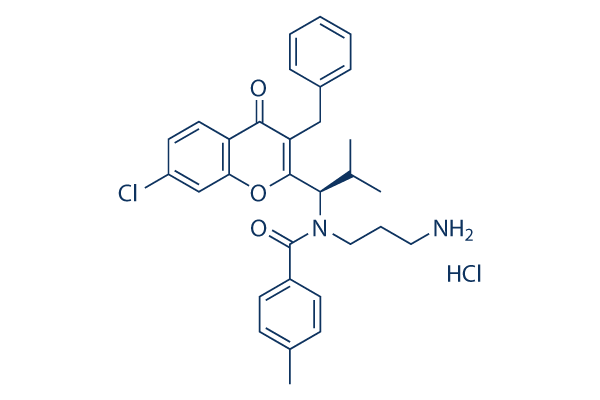
Chemical Structure
Molecular Weight: 553.52
Quality Control
| Related Targets | Akt Wnt/beta-catenin PKC HSP ROCK Microtubule Associated Integrin Bcr-Abl Actin FAK |
|---|---|
| Other Kinesin Inhibitors | GSK-923295 Ispinesib (SB-715992) Monastrol ARQ 621 GW406108X H-Cys(Trt)-OH VLS-1488(KIF18A-IN-6 ) SB-743921 K 858 BTB-1 |
Solubility
|
In vitro |
DMSO
: 100 mg/mL
(180.66 mM)
Ethanol : 100 mg/mL Water : 46 mg/mL |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 553.52 | Formula | C31H33N2O3.HCl |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 940929-33-9 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N/A | Smiles | CC1=CC=C(C=C1)C(=O)N(CCCN)C(C2=C(C(=O)C3=C(O2)C=C(C=C3)Cl)CC4=CC=CC=C4)C(C)C.Cl | ||
Mechanism of Action
| Targets/IC50/Ki |
KSP (MX1 cells)
0.06 nM
KSP (Colo205 cells)
0.07 nM
KSP
0.1 nM(Ki)
KSP (SKOV3 cells)
0.2 nM
KSP (MV522 cells)
1.7 nM
KSP (P388 cells)
14.4 nM
|
|---|---|
| In vitro |
The Ki of SB 743921 for human and mouse KSP is 0.1 nM and 0.12 nM, respectively, while the Ki of SB 743921 for other kinesins including MKLP1, Kin2 is more than 70 μM. SB 743921 blocks assembly of a functional mitotic spindle, thereby causing cell cycle arrest in mitosis and subsequent cell death. SB-743921 has improved potency over ispinesib in both biochemical and cellular assays.
|
| Kinase Assay |
Biochemistry assay
|
|
The motor domains of KSP (amino acids 1–360) is expressed as in Escherichia coli BL21(DE3) as COOH-terminal 6-his fusion proteins. Bacterial pellets are lysed in a microfluidizer with a lysis buffer [50 mM Tris-HCl; 50 mM KCl, 10 mM imidazole, 2 mM MgCl2, 8 mM β-mercaptoethanol, 0.1 mM ATP (pH 7.4)], and proteins are purified using Ni-NTA agarose affinity chromatography, with an elution buffer consisting of 50 mM PIPES, 10% sucrose, 300 mM imidazole, 50 mM KCl, 2 mM MgCl2, mM β-mercaptoethanol, and 0.1 mM ATP (pH 6.8). Steady-state measurements of ATPase activity are performed with a pyruvate kinase–lactate dehydrogenase detection system that coupled the appearance of ADP with oxidation of NADH. Absorbance changes are monitored at 340 nm. All biochemical experiments are performed in PEM25 buffer [25 mM Pipes/KOH (pH 6.8), 2 mM MgCl2, 1 mM EGTA] supplemented with 10 μM SB 743921 for experiments involving microtubules. Rates of ADP release are measured in a stopped-flow apparatus; the decrease in fluorescence of MANT-ATP is monitored. Rates of Pi release are measured in a stopped-flow apparatus, using bacterial phosphate binding protein modified with 7-diethylamino-3-((((2 maleimidyl)ethyl)amino)carbonyl)coumarin (MDCC) dye. Ki estimates of KSP inhibitors are extracted from the dose–response curves, with explicit correction for enzyme concentration. Tubulin polymerization by measuring changes in absorbance at 340 nm is monitored. The assay is performed in 100-μL volumes in 96-well half-area microtiter plates, using a microplate reader with the incubation temperature set at 37 °C.
|
|
| In vivo |
SB-743921 is greater efficacy in vivo against P388 leukemia. SB-743921 has significant efficacy in a broad spectrum of tumor models differing from that of taxanes. SB-743921 is shown to have activity against advanced human tumor xenografts Colo205 (complete regressions), MCF-7, SK-MES, H69, OVCAR-3 (complete and partial regressions), and HT-29, MX-1, MDA-MB-231, A2780 (tumor growth delay). SB-743921 doesn't cause the neuropathy often associated with the tubulin agents.
|
References |
|
Clinical Trial Information
(data from https://clinicaltrials.gov, updated on 2024-05-22)
| NCT Number | Recruitment | Conditions | Sponsor/Collaborators | Start Date | Phases |
|---|---|---|---|---|---|
| NCT00343564 | Completed | Non-Hodgkin''s Lymphoma|Hodgkin''s Disease |
Cytokinetics |
April 2006 | Phase 1|Phase 2 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































