research use only
PP2 (AGL 1879) Src inhibitor
Cat.No.S7008
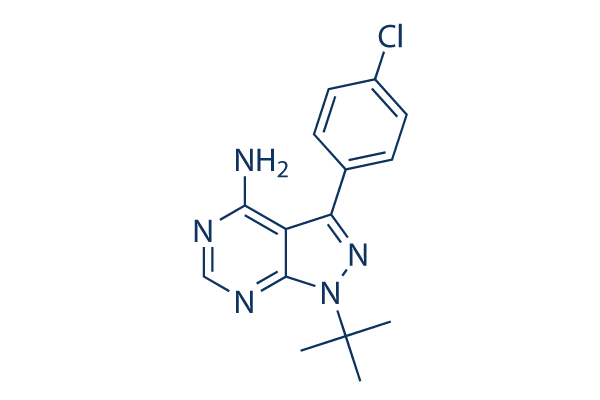
Chemical Structure
Molecular Weight: 301.77
Quality Control
| Related Targets | EGFR VEGFR JAK PDGFR FGFR HIF FLT FLT3 HER2 Bcr-Abl |
|---|---|
| Other Src Inhibitors | WH-4-023 Saracatinib (AZD0530) SU6656 PP1 UM-164 Src Inhibitor 1 RK 24466 Myristic Acid eCF506 Tolimidone (MLR-1023) |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| A549 | Growth inhibition assay | Growth inhibition of human A549 cells, IC50 = 0.01 μM. | 28814374 | |||
| T-cells | Function assay | Inhibition of adhesion kinase in human T cells, IC50 = 0.6 μM. | 18077363 | |||
| T-cells | Function assay | Inhibition of tyrosine phosphorylation in human T cells, IC50 = 0.6 μM. | 18077363 | |||
| SH-SY5Y | Antiproliferative assay | 72 hrs | Antiproliferative activity against human SH-SY5Y cells assessed as cell viability after 72 hrs by XTT assay, IC50 = 6.1 μM. | 21856155 | ||
| Saos2 | Cytotoxicity assay | 48 hrs | Cytotoxicity against human Saos2 cells after 48 hrs by MTT assay, IC50 = 8.07 μM. | 23932070 | ||
| SaOS2 | Antiproliferative assay | Antiproliferative activity against human SaOS2 cells assessed as cellular viability, IC50 = 8.1 μM. | 17929792 | |||
| A431 | Function assay | Inhibitory effect on phospho-Src/nonphospho after EGF (100 uM) stimulation of A431 cells (21), IC50 = 17 μM. | 15109642 | |||
| MEG01 | Antiproliferative assay | Antiproliferative activity against human MEG01 cells, IC50 = 17 μM. | 18257513 | |||
| A431 | Function assay | Inhibitory effect on phospho-Src (Tyr416) after EGF (100 uM) stimulation of A431 cells (38), IC50 = 22 μM. | 15109642 | |||
| K562 | Antiproliferative assay | Antiproliferative activity against human K562 cells, IC50 = 25 μM. | 18257513 | |||
| A431 | Antiproliferative assay | Tested for antiproliferative activity against human A431 cells, IC50 = 32 μM. | 15109642 | |||
| A431 | Antiproliferative assay | Antiproliferative activity against A431 cells, IC50 = 32.2 μM. | 16509573 | |||
| KU812 | Antiproliferative assay | Antiproliferative activity against human KU812 cells, IC50 = 45 μM. | 18257513 | |||
| A431 | Function assay | 10 uM | Inhibition of Src autophosphorylation of Y419 in A431 cells at 10 uM | 16509573 | ||
| 8701-BC | Proapoptotic assay | 10 uM | Proapoptotic activity against 8701-BC cells at 10 uM by PARP assay | 16509573 | ||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 9 mg/mL
(29.82 mM)
Ethanol : 4 mg/mL Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 301.77 | Formula | C15H16ClN5 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 172889-27-9 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | AG 1879,AGL 1879 | Smiles | CC(C)(C)N1C2=NC=NC(=C2C(=N1)C3=CC=C(C=C3)Cl)N | ||
Mechanism of Action
| Targets/IC50/Ki |
LCK
(Cell-free assay) 4 nM
Fyn
(Cell-free assay) 5 nM
|
|---|---|
| In vitro |
PP2 inhibits Src by binding to an area of the molecule that does not overlap with the ATP binding domain. This compound (20 μM) induces 40-50% growth inhibition of HT29 cells, this concentration reduces the Src activity as early as 1 hour and maintains a 35% inhibition of Src activity for 2 days. It (100 mM) decreases the Src activity of HT29 cells in a dose-dependent manner. This chemical (1 mM-100 mM) causes a dose-dependent growth inhibition of human colon cancer cells (HT29, SW480, and PMCO1), liver cancer cells (PLC/PRF/5, KYN-2, Li7, and HepG2), and breast cancer cells (MCF-7, MDA-MB-468, and BT-474). It (20 μM) significantly increases aggregation in most of the cancer cells (HT29, SW480, PMCO1, PLC/PRF/5, KYN-2, Li7, MCF-7, and MDA-MB-468) in E-cadherin dependent manner. This compound (20 μM) enhances E-cadherin expression and also strongly increases E-cadherin’s association with the actin cytoskeleton in cancer cells. It (20 μM) increases the expression of α-catenin, β-catenin, and γ-catenin in HT29 cells, whereas in PLC/PRF/5 and MCF-7 cells, the total protein level of α-catenin does not change, but the levels of β- catenin and γ-catenin increases slightly. This inhibitor inhibits proliferation of two cervical cancer cells (HeLa and SiHa) in a time- and dose-dependent manner. It (10 μM) down-regulates pSrc-Y416, pEGFR-Y845, and -Y1173 expression levels in HeLa and SiHa cells. This chemical (10 μM) could modulate cell cycle arrest by up-regulating p21(Cip1) and p27(Kip1) in both HeLa and SiHa cells and down-regulating expression of cyclin A, and cyclin dependent kinase-2, -4 (Cdk-2, -4) in HeLa and of cyclin B and Cdk-2 in SiHa. |
| Kinase Assay |
Immune complex enzyme assays
|
|
The acid-treated enolase is diluted 1:20 with 1× PBS before aliquoting 100 mL/well into a Nunc 96-well high protein binding assay plate. Assay wells are then aspirated; blocked with 0.5% bovine serum, 1× PBS for 1 h at 37 ℃;and then washed five times with 300 mL of 1× PBS/well. The source of Lck is either LSTRA cells or Lck expressed in HeLa cells using a vaccinia expression system. FynT is expressed in HeLa cells using the vaccinia system. Cells (12.5×106/mL) are lysed in lysis buffer, and the lysates are clarified by centrifugation at 14,000 cpm for 15 min at 4 ℃ in an Eppendorf tube. The clarified lysates are then incubated with the appropriate anti-kinase antibody at 10 μg/mL for 2 h at 4 ℃. Protein A-Sepharose beads are added to the antibody/lysate mixture at 250 μL/mL and allowed to incubate for 30 min at 4 ℃. The beads are then washed twice in 1 mL of lysis buffer and twice in 1 mL of kinase buffer (25 mM HEPES, 3 mM MnCl2, 5mMMgCl2, and 100 μM sodium orthovanadate) and resuspended to 50% (w/v) in kinase buffer. Twenty-five microliters of the bead suspension is added to each well of the enolase-coated 96-well high protein binding plate together with an appropriate concentration of this compound and [γ-32P]ATP (25 μL/well of a 200 μCi/mL solution in kinase buffer). After incubation for 20 min at 20 ℃, 60 μLl of boiling 2× solubilization buffer containing 10 mM ATP is added to the assay wells to terminate the reactions. Thirty microliters of the samples is removed from the wells, boiled for 5 min, and run on a 7.5% SDS-polyacrylamide gel. The gels are subsequently dried and exposed to Kodak X-AR film. For quantitation, films are scanned using a Molecular Dynamics laser scanner, and the optical density of the major substrate band, enolase p46, is determined. In companion experiments for measuring the activity of this chemical against Lck, the assay plate is washed with two wash cycles on a Skatron harvester using 50 mM EDTA, 1 mM ATP. Scintillation fluid (100 μL) is then added to the wells, and 32P incorporation is measured using a micro-β-counter.
|
|
| In vivo |
PP2 (5 mg/kg/day) induces some slowing in the growth rate of the primary tumors relative to the control treated with vehicle in SCID mice inoculated HT29 cells in the spleen. This compound induces some slowing in the growth rate of the primary tumors relative to the control treated with vehicle in SCID mice inoculated HT29 cells in the spleen. This chemical significantly reduces the relative liver weight and liver metastasis volume compared with the controls in SCID mice inoculated HT29 cells in the spleen. This compound (1.5 mg/kg i.p.) treated rats show approximately 50% reduction of infarct size on T2-weighted MRI and in TTC staining compared with controls in rats with focal ischemic brain injury. This chemical results in better the neurological score than controls in rats with focal ischemic brain injury. |
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | p-PI3K / PI3K / p-AKT / AKT / Bcl-2 / Caspase-3 p-Src / Src / p-MAPK / MAPK |
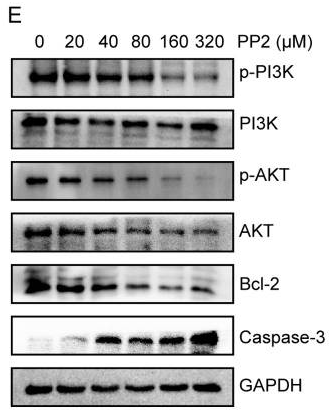
|
30250573 |
| Immunofluorescence | β-catenin FAK / p-FAK |
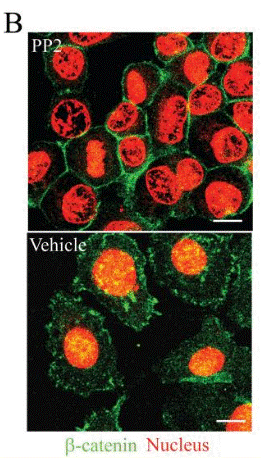
|
18566211 |
Clinical Trial Information
(data from https://clinicaltrials.gov, updated on 2024-05-22)
| NCT Number | Recruitment | Conditions | Sponsor/Collaborators | Start Date | Phases |
|---|---|---|---|---|---|
| NCT03842371 | Unknown status | Sepsis Syndrome |
West China Hospital |
February 11 2019 | -- |
| NCT02407626 | Terminated | Myocardial Ischemia |
Triemli Hospital|University of Alberta |
September 2015 | Not Applicable |
| NCT02315287 | Unknown status | Type 2 Diabetes |
Seoul National University Bundang Hospital |
September 2014 | Phase 4 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.
Frequently Asked Questions
Question 1:
Could you please suggest me the in vivo details about the dilution to reduce the amount of DMSO to 1 to 5% for it?
Answer:
For in vivo study, we recommend to use 4% DMSO +Corn oil up to 2.5 mg/ml for it.






































