research use only
Zosuquidar 3HCl P-gp modulator
Cat.No.S1481
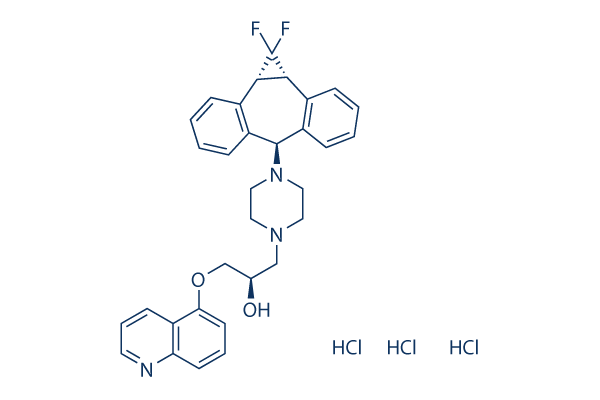
Chemical Structure
Molecular Weight: 636.99
Quality Control
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| CCRF-CEM/VCR1000 | Function assay | 240 secs | Inhibition of P-glycoprotein-mediated efflux from human CCRF-CEM/VCR1000 cells after 240 secs by FACS flow cytometric analysis, IC50=0.05888μM | 22452412 | ||
| DAOY | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for DAOY cells | 29435139 | |||
| SJ-GBM2 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SJ-GBM2 cells | 29435139 | |||
| A673 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for A673 cells | 29435139 | |||
| SK-N-MC | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SK-N-MC cells | 29435139 | |||
| NB-EBc1 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for NB-EBc1 cells | 29435139 | |||
| SK-N-SH | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for SK-N-SH cells | 29435139 | |||
| NB1643 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for NB1643 cells | 29435139 | |||
| LAN-5 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for LAN-5 cells | 29435139 | |||
| BT-12 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for BT-12 cells | 29435139 | |||
| OHS-50 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for OHS-50 cells | 29435139 | |||
| RD | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for RD cells | 29435139 | |||
| MG 63 (6-TG R) | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for MG 63 (6-TG R) cells | 29435139 | |||
| Rh30 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for Rh30 cells | 29435139 | |||
| Rh41 | qHTS assay | qHTS of pediatric cancer cell lines to identify multiple opportunities for drug repurposing: Primary screen for Rh41 cells | 29435139 | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 127 mg/mL
(199.37 mM)
Water : 23 mg/mL Ethanol : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 636.99 | Formula | C32H31F2N3O2.3HCl |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 167465-36-3 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | LY335979 3HCl, RS 33295-198 3HCl, D06387 3HCl | Smiles | C1CN(CCN1CC(COC2=CC=CC3=C2C=CC=N3)O)C4C5=CC=CC=C5C6C(C6(F)F)C7=CC=CC=C47.Cl.Cl.Cl | ||
Mechanism of Action
| Targets/IC50/Ki |
P-gp
(Cell-free assay) 60 nM(Ki)
|
|---|---|
| In vitro |
LY335979 competitively inhibits equilibrium binding of Pgp by blocking [3H]azidopine photoaffinity labeling of the Pgp in CEM/VLB100 plasma membranes. LY335979 alone shows the cytotoxicity to drug-sensitive and MDR cell lines with IC50 ranging from 6 μM-16 μM and produces its ability to completely reverse the resistance of the oncolytics to the MDR cell lines P388/ADR, MCF7/ADR, 2780AD, or UCLA-P3.003VLB at concentration of 0.1 and 0.5 μM. LY335979 significantly restores drug sensitivity in P-gp-expressing leukemia cell lines including K562/HHT40, K562/HHT90, K562/DOX and HL60/DNR, and enhances the cytotoxicity of anthracyclines and ozogamicin (Mylotarg) in primary AML blasts with active P-gp. A latest paper indicates that LY335979 completely inhibits apically directed transport of (Z)-endoxifen in the ABCB1-transduced cells. |
| Kinase Assay |
ATPase Assay
|
|
P-Glycoprotein ATPase activity is measured by the liberation of inorganic phosphate from ATP. The assay is measured in a 96-well plate for 90 min at 37 °C. Membranes (8 μg-10 μg protein) are incubated in a total volume of 100 μL of buffer A containing 5 mM sodium azide, 1 mM ouabain, 1 mM EGTA, 3 mM ATP, an ATP regenerating system composed of 5 mM phosphoenolpyruvate, and 3.6 units/mL pyruvate kinase in the presence and absence of 1 mM sodium vanadate. Pgp-ATPase activity is defined as the vanadate-sensitive portion of the total ATPase activity. Plates are read 3 minutes after the addition of the detection solution. The absorbance is measured at 690 nm by a microtiter dish reader. A phosphate standard curve is used to calculate the μmol of phosphate formed. Samples are measured in triplicate.
|
References |
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































