research use only
DBeQ p97 inhibitor
Cat.No.S7199
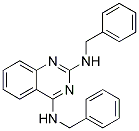
Chemical Structure
Molecular Weight: 340.42
Quality Control
| Related Targets | Proteasome E1 Activating E3 Ligase DUB SUMO E2 conjugating |
|---|---|
| Other p97 Inhibitors | NMS-873 CB-5339 NPD8733 ML240 |
Solubility
|
In vitro |
DMSO
: 68 mg/mL
(199.75 mM)
Ethanol : 5 mg/mL Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 340.42 | Formula | C22H20N4 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 177355-84-9 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | JRF 12 | Smiles | C1=CC=C(C=C1)CNC2=NC(=NC3=CC=CC=C32)NCC4=CC=CC=C4 | ||
Mechanism of Action
| Features |
Rapidly and potently induces activation of executioner caspases and cell death.
|
|---|---|
| Targets/IC50/Ki |
p97
1.5 μM
|
| In vitro |
DBeQ blocks UbG76V-GFP, ODD-Luc and Luc-ODC degradation with IC50 of 2.6 μM, 56 μM and 45 μM in HeLa cells. This compound is at least 50-fold less potent toward N-ethylmaleimide–sensitive factor (NSF) and 26S proteasome. It inhibits p97 competitively with respect to ATP, with Ki of 3.2 μM, suggesting that it binds to the active site of the D2 domain. This compound (10 μM) potently blocks degradation of TCRα-GFP in HEK293 cells. It induces CHOP within 3 hours in a concentration-dependent manner but does not increase p21 level in HEK293 cells. This compound (15 μM) induces a strong accumulation of LC3-II in the nucleus plus membrane-enriched and cytosolic fractions in Hela cells. It acts by blocking autophagic degradation of LC3-II instead of inducing autophagy in HeLa cells. This compound (10 μM) rapidly promotes activation of the “executioner” caspases-3 and -7 in HeLa cells. It activates the intrinsic caspase-9 apoptotic pathway more than the extrinsic caspase-8 pathway, whereas STS activates both pathways to a similar extent. This compound is fivefold more active against multiple myeloma (RPMI8226) cells than normal human fetal lung fibroblasts (MRC5), with HeLa and Hek293 cells showing intermediate sensitivities. It exhibits 20-fold selectivity for stabilizing p97-dependent vs. independent UPS reporter substrates in HeLa cells. This chemical impairs degradation of substrates within the ERAD and autophagy pathways. It (12 μM) inhibits intracellular neutralization in a dose-dependent manner in HeLa cells. This compound (10 μM), which completely inhibits degradation of virus and antibody in the fate-of-capsid experiment, fails to prevent degradation of IgG Fc. It (9 μM) reduces the initial gradient of neutralization as a function of antibody concentration. This compound decreases both basal and nutrient-stimulated phosphorylation of MTOR targets similar to the effects of rapamycin in U20S cells.
|
| Kinase Assay |
Manual ATPase Assay
|
|
Assay Buffer [20 μL of 2.5× concentration, where 1× = 50 mM Tris (pH 7.4), 20 mM MgCl2, 1 mM EDTA, and 0.5 mM tris(2-carboxyethyl)phosphine (TCEP)] is dispensed into each well of a 96-well plate. Purified p97 (25 μL of 50 μM) is diluted in 975 μL of 1× Assay Buffer, and 10 μL is dispensed in each well. This compound (10 μL) or 5% DMSO (10 μL) is then added to each well, and the plate is incubated at room temperature for 10 min. The ATPase assay is carried out by adding to each well 10 μL of 500 μM ATP (pH 7.5), incubating at room temperature for 60 min, and then adding 50 μL Kinase Glo Plus reagent, followed by a final 10-min incubation at room temperature in the dark. Luminescence is read on an Analyst AD. This chemical is assayed at a range of concentrations (0, 0.048, 0.24, 1.2, 6, and 30 μM) in triplicate.
|
References |
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































