research use only
SH-4-54 STAT3/STAT5 Inhibitor
Cat.No.S7337
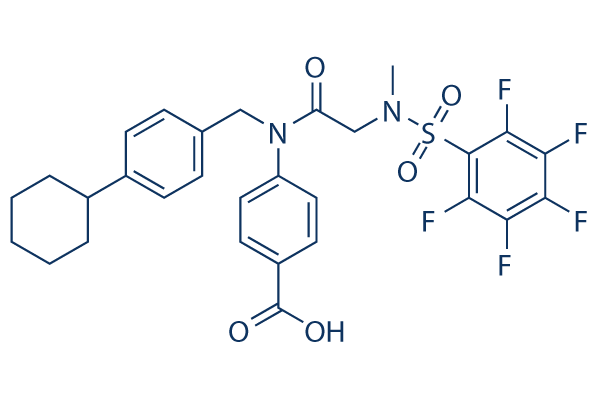
Chemical Structure
Molecular Weight: 610.59
Quality Control
| Related Targets | EGFR JAK Pim |
|---|---|
| Other STAT Inhibitors | Napabucasin (BBI608) Stattic NSC 74859 (S3I-201) Cryptotanshinone (Tanshinone C) C188-9 (TTI-101) BP-1-102 AS1517499 HO-3867 Nifuroxazide Homoharringtonine (HHT) |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| 127EF | Cytotoxicity assay | 3 days | Cytotoxicity against human 127EF cells assessed as cell viability after 3 days by Alamar Blue assay, IC50=0.066μM. | 24900612 | ||
| 30M | Cytotoxicity assay | 3 days | Cytotoxicity against human 30M cells assessed as cell viability after 3 days by Alamar Blue assay, IC50=0.1μM. | 24900612 | ||
| 84EF | Cytotoxicity assay | 3 days | Cytotoxicity against human 84EF cells assessed as cell viability after 3 days by Alamar Blue assay, IC50=0.102μM. | 24900612 | ||
| 67EF | Cytotoxicity assay | 3 days | Cytotoxicity against human 67EF cells assessed as cell viability after 3 days by Alamar Blue assay, IC50=0.106μM. | 24900612 | ||
| 73EF | Cytotoxicity assay | 3 days | Cytotoxicity against human 73EF cells assessed as cell viability after 3 days by Alamar Blue assay, IC50=0.162μM. | 24900612 | ||
| 25EF | Cytotoxicity assay | 3 days | Cytotoxicity against human 25EF cells assessed as cell viability after 3 days by Alamar Blue assay, IC50=0.234μM. | 24900612 | ||
| 30M | Cytotoxicity assay | 8 days | Cytotoxicity against human 30M cells assessed as cell viability after 8 days by Alamar Blue assay, IC50=0.43μM. | 24900612 | ||
| 73M | Cytotoxicity assay | 8 days | Cytotoxicity against human 73M cells assessed as cell viability after 8 days by Alamar Blue assay, IC50=1.03μM. | 24900612 | ||
| 147EF | Apoptosis assay | 0.1 to 1 uM | 3 hrs | Induction of apoptosis in human 147EF cells assessed as PARP cleavage at 0.1 to 1 uM after 3 hrs by Western blotting analysis | 24900612 | |
| 147EF | Function assay | 0.1 to 1 uM | 3 hrs | Inhibition of AKT phosphorylation in human 147EF cells at 0.1 to 1 uM after 3 hrs by Western blotting analysis | 24900612 | |
| 147EF | Function assay | 0.1 to 1 uM | 3 hrs | Inhibition of STAT3 phosphorylation at Y705 in human 147EF cells at 0.1 to 1 uM after 3 hrs by Western blotting analysis | 24900612 | |
| 73M | Function assay | 1 uM | Inhibition of STAT3 phosphorylation at Y705 in human 73M cells at 1 uM | 24900612 | ||
| 147EF | Function assay | 2 to 72 hrs | Inhibition of STAT3 in human 147EF cells assessed as reduction of phosphorylated Bcl-xL level after 2 to 72 hrs by Western blotting analysis | 24900612 | ||
| 147EF | Function assay | 2 to 72 hrs | Inhibition of STAT3 in human 147EF cells assessed as reduction of phosphorylated cyclin D1 level after 2 to 72 hrs by Western blotting analysis | 24900612 | ||
| BT73 | Function assay | 10 mg/kg | 3 days | In vivo inhibition of STAT3 phosphorylation in tumor of NOD-SCID mouse xenografted with human BT73 cells at 10 mg/kg, ip administered as 4 days on/3 days off schedule measured 2 hrs post last dose | 24900612 | |
| BT73 | Antitumor assay | 10 mg/kg | 3 days | Antitumor activity against human BT73 cells xenografted in NOD-SCID mouse assessed as decrease in tumor size at 10 mg/kg, ip administered as 4 days on/3 days off schedule measured 2 hrs post last dose by hematoxylin/eosin staining | 24900612 | |
| BT73 | Antitumor assay | 10 mg/kg | 3 days | Antitumor activity against human BT73 cells xenografted in NOD-SCID mouse assessed as induction of apoptosis in tumor at 10 mg/kg, ip administered as 4 days on/3 days off schedule measured 2 hrs post last dose by TUNEL assay | 24900612 | |
| BT73 | Antitumor assay | 10 mg/kg | 3 days | Antitumor activity against human BT73 cells xenografted in NOD-SCID mouse assessed as tumor growth inhibition by measuring downregulation of Ki67 protein expression at 10 mg/kg, ip administered as 4 days on/3 days off schedule measured 2 hrs post last dos | 24900612 | |
| Vero | Function assay | Vero cells viability counterscreen for qRT-PCR qHTS assay of selected Zika virus inhibitors | 33229545 | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 100 mg/mL
(163.77 mM)
Ethanol : 100 mg/mL Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 610.59 | Formula | C29H27F5N2O5S |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 1456632-40-8 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N/A | Smiles | CN(CC(=O)N(CC1=CC=C(C=C1)C2CCCCC2)C3=CC=C(C=C3)C(=O)O)S(=O)(=O)C4=C(C(=C(C(=C4F)F)F)F)F | ||
Mechanism of Action
| Targets/IC50/Ki |
STAT3
(Cell-free assay) 300 nM(Kd)
STAT5
(Cell-free assay) 464 nM(Kd)
|
|---|---|
| In vitro |
SH-4-54 shows unprecedented cytotoxicity in human glioblastoma brain cancer stem cells (BTSCs), while has no toxicity in human fetal astrocytes. In addition, this compound effectively suppresses STAT3 phosphorylation and its downstream transcriptional targets.
|
| Kinase Assay |
Surface Plasmon Resonance (SPR) studies
|
|
The binding experiments are carried out on a ProteOn XPR36 biosensor at 25°C using the HTE sensor chip. The flow cells of the sensor chip are loaded with a nickel solution at 30 μL/min for 120 s to saturate the Tris–NTA surface with Ni(II) ions. Purified His-tagged STAT3 and STAT5 in PBST buffer (PBS with 0.005% (v/v) Tween-20 and 0.001% DMSO pH 7.4) is injected in the first and second channels of the chip respectively in the vertical direction at a flow rate of 25 μg/μL for 300 s, which attained, on average, ~8000 resonance unit (RU). After a wash with PBST buffer, inhibitors binding to the immobilized proteins is monitored by injecting a range of concentrations along with a blank at a flow rate of 100 μL/min for 200 s for each of these small molecules. When the injection of the small molecule inhibitor is completed, running buffer is allowed to flow over the immobilized substrates for the non-specifically bound inhibitors to dissociate for 600 s. Following dissociation of the inhibitors, the chip surface is regenerated with an injection of 1 M NaCl at a flow rate of 100 μL/ml for 18 s. Interspot channel reference is used for non-specific binding corrections and the blank channel used with each analyte injection served as a double reference to correct for possible baseline drift. Data are analyzed using ProteOn Manager Software version 3.1. The Langmuir 1:1 binding model was used to determine the KD values.
|
|
| In vivo |
In mice orthotopically xenografted with BT73, SH-4-54 (10 mg/kg i.p.) exhibits BBB permeability, potently suppresses glioma tumor growth, and inhibits pSTAT3.
|
References |
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | LMP1 / p-STAT3 / STAT3 |
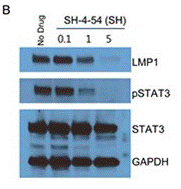
|
28445949 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































