- Bioactive Compounds
- By Signaling Pathways
- PI3K/Akt/mTOR
- Epigenetics
- Methylation
- Immunology & Inflammation
- Protein Tyrosine Kinase
- Angiogenesis
- Apoptosis
- Autophagy
- ER stress & UPR
- JAK/STAT
- MAPK
- Cytoskeletal Signaling
- Cell Cycle
- TGF-beta/Smad
- DNA Damage/DNA Repair
- Compound Libraries
- Popular Compound Libraries
- Customize Library
- Clinical and FDA-approved Related
- Bioactive Compound Libraries
- Inhibitor Related
- Natural Product Related
- Metabolism Related
- Cell Death Related
- By Signaling Pathway
- By Disease
- Anti-infection and Antiviral Related
- Neuronal and Immunology Related
- Fragment and Covalent Related
- FDA-approved Drug Library
- FDA-approved & Passed Phase I Drug Library
- Preclinical/Clinical Compound Library
- Bioactive Compound Library-I
- Bioactive Compound Library-Ⅱ
- Kinase Inhibitor Library
- Express-Pick Library
- Natural Product Library
- Human Endogenous Metabolite Compound Library
- Alkaloid Compound LibraryNew
- Angiogenesis Related compound Library
- Anti-Aging Compound Library
- Anti-alzheimer Disease Compound Library
- Antibiotics compound Library
- Anti-cancer Compound Library
- Anti-cancer Compound Library-Ⅱ
- Anti-cancer Metabolism Compound Library
- Anti-Cardiovascular Disease Compound Library
- Anti-diabetic Compound Library
- Anti-infection Compound Library
- Antioxidant Compound Library
- Anti-parasitic Compound Library
- Antiviral Compound Library
- Apoptosis Compound Library
- Autophagy Compound Library
- Calcium Channel Blocker LibraryNew
- Cambridge Cancer Compound Library
- Carbohydrate Metabolism Compound LibraryNew
- Cell Cycle compound library
- CNS-Penetrant Compound Library
- Covalent Inhibitor Library
- Cytokine Inhibitor LibraryNew
- Cytoskeletal Signaling Pathway Compound Library
- DNA Damage/DNA Repair compound Library
- Drug-like Compound Library
- Endoplasmic Reticulum Stress Compound Library
- Epigenetics Compound Library
- Exosome Secretion Related Compound LibraryNew
- FDA-approved Anticancer Drug LibraryNew
- Ferroptosis Compound Library
- Flavonoid Compound Library
- Fragment Library
- Glutamine Metabolism Compound Library
- Glycolysis Compound Library
- GPCR Compound Library
- Gut Microbial Metabolite Library
- HIF-1 Signaling Pathway Compound Library
- Highly Selective Inhibitor Library
- Histone modification compound library
- HTS Library for Drug Discovery
- Human Hormone Related Compound LibraryNew
- Human Transcription Factor Compound LibraryNew
- Immunology/Inflammation Compound Library
- Inhibitor Library
- Ion Channel Ligand Library
- JAK/STAT compound library
- Lipid Metabolism Compound LibraryNew
- Macrocyclic Compound Library
- MAPK Inhibitor Library
- Medicine Food Homology Compound Library
- Metabolism Compound Library
- Methylation Compound Library
- Mouse Metabolite Compound LibraryNew
- Natural Organic Compound Library
- Neuronal Signaling Compound Library
- NF-κB Signaling Compound Library
- Nucleoside Analogue Library
- Obesity Compound Library
- Oxidative Stress Compound LibraryNew
- Plant Extract Library
- Phenotypic Screening Library
- PI3K/Akt Inhibitor Library
- Protease Inhibitor Library
- Protein-protein Interaction Inhibitor Library
- Pyroptosis Compound Library
- Small Molecule Immuno-Oncology Compound Library
- Mitochondria-Targeted Compound LibraryNew
- Stem Cell Differentiation Compound LibraryNew
- Stem Cell Signaling Compound Library
- Natural Phenol Compound LibraryNew
- Natural Terpenoid Compound LibraryNew
- TGF-beta/Smad compound library
- Traditional Chinese Medicine Library
- Tyrosine Kinase Inhibitor Library
- Ubiquitination Compound Library
-
Cherry Picking
You can personalize your library with chemicals from within Selleck's inventory. Build the right library for your research endeavors by choosing from compounds in all of our available libraries.
Please contact us at info@selleckchem.com to customize your library.
You could select:
- Antibodies
- Bioreagents
- qPCR
- 2x SYBR Green qPCR Master Mix
- 2x SYBR Green qPCR Master Mix(Low ROX)
- 2x SYBR Green qPCR Master Mix(High ROX)
- Protein Assay
- Protein A/G Magnetic Beads for IP
- Anti-Flag magnetic beads
- Anti-Flag Affinity Gel
- Anti-Myc magnetic beads
- Anti-HA magnetic beads
- Poly DYKDDDDK Tag Peptide lyophilized powder
- Protease Inhibitor Cocktail
- Protease Inhibitor Cocktail (EDTA-Free, 100X in DMSO)
- Phosphatase Inhibitor Cocktail (2 Tubes, 100X)
- Cell Biology
- Cell Counting Kit-8 (CCK-8)
- Animal Experiment
- Mouse Direct PCR Kit (For Genotyping)
- New Products
- Contact Us
research use only
Barasertib-HQPA (AZD2811) Aurora Kinase inhibitor
Defosbarasertib (AZD1152-HQPA, AZD2811, INH-34, Barasertib-HQPA) is a highly selective Aurora B inhibitor with IC50 of 0.37 nM in a cell-free assay, ~3700 fold more selective for Aurora B over Aurora A. Phase 1.
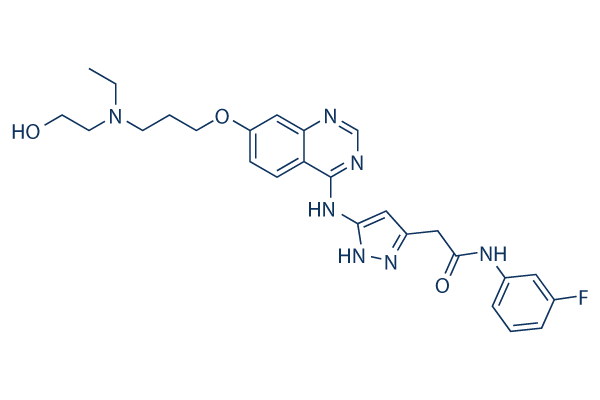
Chemical Structure
Molecular Weight: 507.56
Purity & Quality Control
Batch:
Purity:
99.97%
99.97
Related Products
| Related Targets | Aurora A Aurora B Aurora C Aurora B | Click to Expand |
|---|---|---|
| Related Products | Alisertib (MLN8237) Tozasertib (VX-680) ZM 447439 MLN8054 Hesperadin Danusertib (PHA-739358) MK-5108 TCS7010 (Aurora A Inhibitor I) AMG-900 PHA-680632 SNS-314 CCT137690 GSK1070916 CYC116 TAK-901 AZD1152(Barasertib) CCT129202 SNS-314 Mesylate LY3295668 SP-96 | Click to Expand |
| Related Compound Libraries | Kinase Inhibitor Library PI3K/Akt Inhibitor Library MAPK Inhibitor Library DNA Damage/DNA Repair compound Library Cell Cycle compound library | Click to Expand |
Signaling Pathway
Cell Culture and Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| LNCaP | Growth Inhibition Assay | 0-500 nM | 48 h | IC50=25 nM | 25277659 | |
| LNCaP | Apoptosis Assay | 0-500 nM | 48 h | induces apoptotic cell death through caspase-3 upregulation | 25277659 | |
| LNCaP | Function Assay | 50 nM | 48 h | induces micronuclei with aneugenic mechanism | 25277659 | |
| Ramos | Function Assay | 500 nM | 0-72 h | inhibits Aurora B kinase | 21371446 | |
| Daudi | Function Assay | 500 nM | 0-72 h | inhibits Aurora B kinase | 21371446 | |
| L540 | Function Assay | 500 nM | 0-72 h | inhibits Aurora B kinase | 21371446 | |
| BJAJ | Growth Inhibition Assay | 500 nM | 0-72 h | inhibits cell growth significantly | 21371446 | |
| Ramos | Growth Inhibition Assay | 500 nM | 0-72 h | inhibits cell growth significantly | 21371446 | |
| Raji | Growth Inhibition Assay | 500 nM | 0-72 h | inhibits cell growth significantly | 21371446 | |
| Daudi | Growth Inhibition Assay | 500 nM | 0-72 h | inhibits cell growth significantly | 21371446 | |
| L428 | Growth Inhibition Assay | 500 nM | 0-72 h | inhibits cell growth | 21371446 | |
| KM-H2 | Growth Inhibition Assay | 500 nM | 0-72 h | inhibits cell growth | 21371446 | |
| HDLM-2 | Growth Inhibition Assay | 500 nM | 0-72 h | inhibits cell growth | 21371446 | |
| L450 | Growth Inhibition Assay | 500 nM | 0-72 h | inhibits cell growth | 21371446 | |
| BJAJ | Apoptosis Assay | 500 nM | 0-72 h | induces apoptosis in a time-dependent manner | 21371446 | |
| Ramos | Apoptosis Assay | 500 nM | 0-72 h | induces apoptosis in a time-dependent manner | 21371446 | |
| Raji | Apoptosis Assay | 500 nM | 0-72 h | induces apoptosis in a time-dependent manner | 21371446 | |
| Daudi | Apoptosis Assay | 500 nM | 0-72 h | induces apoptosis in a time-dependent manner | 21371446 | |
| L428 | Apoptosis Assay | 500 nM | 0-72 h | induces apoptosis in a time-dependent manner | 21371446 | |
| KM-H2 | Apoptosis Assay | 500 nM | 0-72 h | induces apoptosis in a time-dependent manner | 21371446 | |
| HDLM-2 | Apoptosis Assay | 500 nM | 0-72 h | induces apoptosis in a time-dependent manner | 21371446 | |
| L450 | Apoptosis Assay | 500 nM | 0-72 h | induces apoptosis in a time-dependent manner | 21371446 | |
| SW620 | Growth Inhibition Assay | EC50=10±2.1 nM | 21245090 | |||
| HCT116 | Growth Inhibition Assay | EC50=11±3.3 nM | 21245090 | |||
| MDA-MB-435 | Growth Inhibition Assay | 0-10000 nM | 2-5 d | DMSO | IC50=125 nM | 20175926 |
| MDA-MB-468 | Growth Inhibition Assay | 0-10000 nM | 2-5 d | DMSO | IC50=14 nM | 20175926 |
| MDA-MB-231 | Growth Inhibition Assay | 0-10000 nM | 2-5 d | DMSO | IC50=105 nM | 20175926 |
| BT474 | Growth Inhibition Assay | 0-10000 nM | 2-5 d | DMSO | IC50=8 nM | 20175926 |
| MDA-MB-361 | Growth Inhibition Assay | 0-10000 nM | 2-5 d | DMSO | IC50=70 nM | 20175926 |
| HER18 | Growth Inhibition Assay | 0-10000 nM | 2-5 d | DMSO | IC50=20 nM | 20175926 |
| HER18 | Apoptosis Assay | 100 nM | 0/24/48 h | DMSO | induces apoptosis and reduces clonogenic potential | 20175926 |
| MDA-MB-231 | Apoptosis Assay | 105 nM | 0/24/48 h | DMSO | induces apoptosis and reduces clonogenic potential | 20175926 |
| JHH-1 | Growth Inhibition Assay | 0.3–1000 nM | 72 h | EC50=17.4±1.0 nM | 19913935 | |
| JHH-2 | Growth Inhibition Assay | 0.3–1000 nM | 72 h | EC50=218.0±10.8 nM | 19913935 | |
| JHH-4 | Growth Inhibition Assay | 0.3–1000 nM | 72 h | EC50=155.6±16.8 nM | 19913935 | |
| HuH-1 | Growth Inhibition Assay | 0.3–1000 nM | 72 h | EC50=27.3±5.0 nM | 19913935 | |
| HuH-6 | Growth Inhibition Assay | 0.3–1000 nM | 72 h | EC50=3.7±0.6 nM | 19913935 | |
| HuH-7 | Growth Inhibition Assay | 0.3–1000 nM | 72 h | EC50=6.8±0.3 nM | 19913935 | |
| HLE | Growth Inhibition Assay | 0.3–1000 nM | 72 h | EC50=45.9±6.4 nM | 19913935 | |
| HLF | Growth Inhibition Assay | 0.3–1000 nM | 72 h | EC50=126.1±12.2 nM | 19913935 | |
| PLC/PRF/5 | Growth Inhibition Assay | 0.3–1000 nM | 72 h | EC50=76.9±9.9 nM | 19913935 | |
| SK-Hep1 | Growth Inhibition Assay | 0.3–1000 nM | 72 h | EC50=21.9±1.2 nM | 19913935 | |
| Hep3B | Growth Inhibition Assay | 0.3–1000 nM | 72 h | EC50=7.6±1.2 nM | 19913935 | |
| HepG2 | Growth Inhibition Assay | 0.3–1000 nM | 72 h | EC50=14.7±1.7 nM | 19913935 | |
| Ramos | Apoptosis Assay | 25/50/100 nM | 48 h | increases the levels of the cleaved forms of PARP and caspase 3 | 19823168 | |
| Daudi | Apoptosis Assay | 25/50/100 nM | 48 h | increases the levels of the cleaved forms of PARP and caspase 3 | 19823168 | |
| BALM-14 | Apoptosis Assay | 12.5/25/50 nM | 48 h | increases the levels of the cleaved forms of PARP and caspase 3 | 19823168 | |
| BALM-27 | Apoptosis Assay | 12.5/25/50 nM | 48 h | increases the levels of the cleaved forms of PARP and caspase 3 | 19823168 | |
| NB4 | Growth Inhibition Assay | 0.01/0.1/1 μM | 48 h | inhibits cell growth significantly | 18367484 | |
| HeLa | Function Assay | 1 uM | 24 hrs | Inhibition of aurora B autophosphorylation in human HeLa cells at 1 uM after 24 hrs by Western blotting | 20684549 | |
| Click to View More Cell Line Experimental Data | ||||||
Mechanism of Action
| Targets |
|
|---|
In vitro |
||||
| In vitro | Barasertib (AZD1152-HQPA), a highly selective Aurora B inhibitor, causes polyploidy and apoptosis in many cancer cell lines.[1] |
|||
|---|---|---|---|---|
| Cell Research | Cell lines | NCI-H82 cells, SCLC and NSCLC cell lines | ||
| Concentrations | -- | |||
| Incubation Time | -- | |||
| Method | -- |
|||
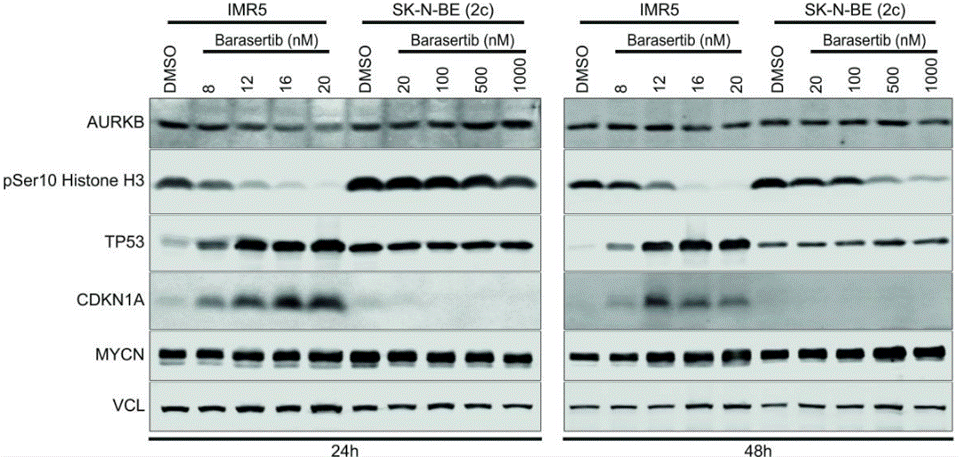
| Experimental Result Images | Methods | Biomarkers | Images | PMID |
| Western blot | AURKB / pSer10 Histone H3 / TP53 / CDKN1A / MYCN p53 / p21 / p-p38 / p38 |
|
26497213 | |
| Immunofluorescence | p21 |
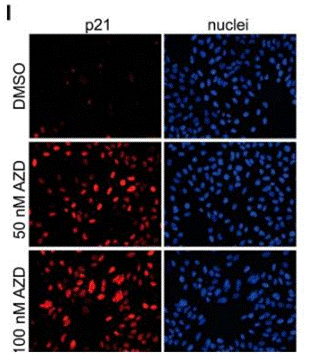
|
24782314 | |
| Growth inhibition assay | Cell viability |
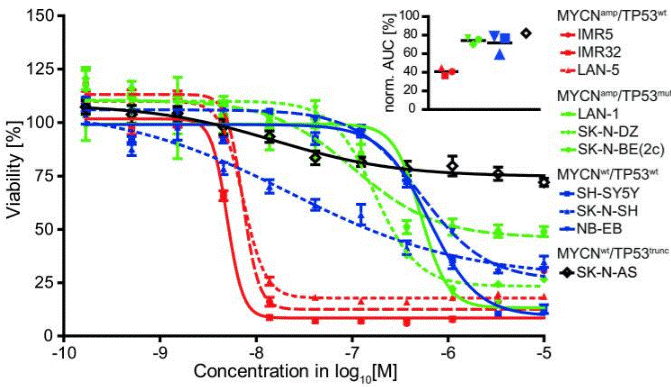
|
26497213 | |
In Vivo |
||
| In vivo | Barasertib (AZD1152-HQPA) is an aurora B kinase inhibitor, and has efficacy against RB1 / SCLC cell line xenografts, RB1 / SCLC PDXs, and autochthonous Rb1 / neuroendocrine tumors.[1] |
|
|---|---|---|
| NCT Number | Recruitment | Conditions | Sponsor/Collaborators | Start Date | Phases |
|---|---|---|---|---|---|
| NCT02579226 | Completed | Advanced Solid Tumours |
AstraZeneca |
October 28 2015 | Phase 1 |
References |
|
Chemical Information
| Molecular Weight | 507.56 | Formula | C26H30FN7O3 |
| CAS No. | 722544-51-6 | SDF | Download SDF |
| Synonyms | AZD2811, INH-34, Barasertib-HQPA , Defosbarasertib | ||
| Smiles | CCN(CCCOC1=CC2=C(C=C1)C(=NC=N2)NC3=NNC(=C3)CC(=O)NC4=CC(=CC=C4)F)CCO | ||
Storage and Stability
| Storage (From the date of receipt) | |||
|
In vitro |
DMSO : 100 mg/mL ( (197.02 mM) Moisture-absorbing DMSO reduces solubility. Please use fresh DMSO.) Water : Insoluble Ethanol : Insoluble |
Molecular Weight Calculator |
|
In vivo Add solvents to the product individually and in order. |
In vivo Formulation Calculator |
|||||
Preparing Stock Solutions
Molarity Calculator
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
mg/kg
g
μL
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
% DMSO
%
% Tween 80
% ddH2O
%DMSO
%
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Tech Support
Answers to questions you may have can be found in the inhibitor handling instructions. Topics include how to prepare stock solutions, how to store inhibitors, and issues that need special attention for cell-based assays and animal experiments.
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.
* Indicates a Required Field
Frequently Asked Questions
Question 1:
Can you let me know what solvent I can use for it, cat # S1147, for in vivo use? (IP injection in mice)
Answer:
S1147 can be dissolved in 30% PEG400/0.5% Tween80/5% Propylene glycol at 30mg/ml as a clear solution. Usually, when prepare the solution, we will add organic solvents first, then add Tween 80, then water. But this compound can not dissolve in 30% PEG400/0.5% Tween80/5% Propylene glycol clearly. After water was added, it became a clear solution.