- Bioactive Compounds
- By Signaling Pathways
- PI3K/Akt/mTOR
- Epigenetics
- Methylation
- Immunology & Inflammation
- Protein Tyrosine Kinase
- Angiogenesis
- Apoptosis
- Autophagy
- ER stress & UPR
- JAK/STAT
- MAPK
- Cytoskeletal Signaling
- Cell Cycle
- TGF-beta/Smad
- Compound Libraries
- Popular Compound Libraries
- Customize Library
- Clinical and FDA-approved Related
- Bioactive Compound Libraries
- Inhibitor Related
- Natural Product Related
- Metabolism Related
- Cell Death Related
- By Signaling Pathway
- By Disease
- Anti-infection and Antiviral Related
- Neuronal and Immunology Related
- Fragment and Covalent Related
- FDA-approved Drug Library
- FDA-approved & Passed Phase I Drug Library
- Preclinical/Clinical Compound Library
- Bioactive Compound Library-I
- Bioactive Compound Library-Ⅱ
- Kinase Inhibitor Library
- Express-Pick Library
- Natural Product Library
- Human Endogenous Metabolite Compound Library
- Alkaloid Compound LibraryNew
- Angiogenesis Related compound Library
- Anti-Aging Compound Library
- Anti-alzheimer Disease Compound Library
- Antibiotics compound Library
- Anti-cancer Compound Library
- Anti-cancer Compound Library-Ⅱ
- Anti-cancer Metabolism Compound Library
- Anti-Cardiovascular Disease Compound Library
- Anti-diabetic Compound Library
- Anti-infection Compound Library
- Antioxidant Compound Library
- Anti-parasitic Compound Library
- Antiviral Compound Library
- Apoptosis Compound Library
- Autophagy Compound Library
- Calcium Channel Blocker LibraryNew
- Cambridge Cancer Compound Library
- Carbohydrate Metabolism Compound LibraryNew
- Cell Cycle compound library
- CNS-Penetrant Compound Library
- Covalent Inhibitor Library
- Cytokine Inhibitor LibraryNew
- Cytoskeletal Signaling Pathway Compound Library
- DNA Damage/DNA Repair compound Library
- Drug-like Compound Library
- Endoplasmic Reticulum Stress Compound Library
- Epigenetics Compound Library
- Exosome Secretion Related Compound LibraryNew
- FDA-approved Anticancer Drug LibraryNew
- Ferroptosis Compound Library
- Flavonoid Compound Library
- Fragment Library
- Glutamine Metabolism Compound Library
- Glycolysis Compound Library
- GPCR Compound Library
- Gut Microbial Metabolite Library
- HIF-1 Signaling Pathway Compound Library
- Highly Selective Inhibitor Library
- Histone modification compound library
- HTS Library for Drug Discovery
- Human Hormone Related Compound LibraryNew
- Human Transcription Factor Compound LibraryNew
- Immunology/Inflammation Compound Library
- Inhibitor Library
- Ion Channel Ligand Library
- JAK/STAT compound library
- Lipid Metabolism Compound LibraryNew
- Macrocyclic Compound Library
- MAPK Inhibitor Library
- Medicine Food Homology Compound Library
- Metabolism Compound Library
- Methylation Compound Library
- Mouse Metabolite Compound LibraryNew
- Natural Organic Compound Library
- Neuronal Signaling Compound Library
- NF-κB Signaling Compound Library
- Nucleoside Analogue Library
- Obesity Compound Library
- Oxidative Stress Compound LibraryNew
- Plant Extract Library
- Phenotypic Screening Library
- PI3K/Akt Inhibitor Library
- Protease Inhibitor Library
- Protein-protein Interaction Inhibitor Library
- Pyroptosis Compound Library
- Small Molecule Immuno-Oncology Compound Library
- Mitochondria-Targeted Compound LibraryNew
- Stem Cell Differentiation Compound LibraryNew
- Stem Cell Signaling Compound Library
- Natural Phenol Compound LibraryNew
- Natural Terpenoid Compound LibraryNew
- TGF-beta/Smad compound library
- Traditional Chinese Medicine Library
- Tyrosine Kinase Inhibitor Library
- Ubiquitination Compound Library
-
Cherry Picking
You can personalize your library with chemicals from within Selleck's inventory. Build the right library for your research endeavors by choosing from compounds in all of our available libraries.
Please contact us at [email protected] to customize your library.
You could select:
- Antibodies
- Bioreagents
- qPCR
- 2x SYBR Green qPCR Master Mix
- 2x SYBR Green qPCR Master Mix(Low ROX)
- 2x SYBR Green qPCR Master Mix(High ROX)
- Protein Assay
- Protein A/G Magnetic Beads for IP
- Anti-Flag magnetic beads
- Anti-Flag Affinity Gel
- Anti-Myc magnetic beads
- Anti-HA magnetic beads
- Poly FLAG Peptide lyophilized powder
- Protease Inhibitor Cocktail
- Protease Inhibitor Cocktail (EDTA-Free, 100X in DMSO)
- Phosphatase Inhibitor Cocktail (2 Tubes, 100X)
- Cell Biology
- Cell Counting Kit-8 (CCK-8)
- Animal Experiment
- Mouse Direct PCR Kit (For Genotyping)
- New Products
- Contact Us
Hesperadin
Hesperadin potently inhibits Aurora B with IC50 of 250 nM in a cell-free assay. It markedly reduces the activity of AMPK, Lck, MKK1, MAPKAP-K1, CHK1 and PHK while it does not inhibit MKK1 activity in vivo.
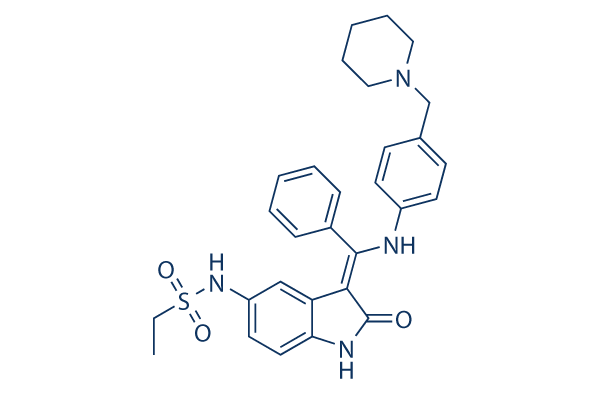
Hesperadin Chemical Structure
CAS: 422513-13-1
Selleck's Hesperadin has been cited by 43 publications
Purity & Quality Control
Batch:
Purity:
99.09%
99.09
Hesperadin Related Products
| Related Targets | Aurora A Aurora B Aurora C Aurora B | Click to Expand |
|---|---|---|
| Related Compound Libraries | Kinase Inhibitor Library PI3K/Akt Inhibitor Library MAPK Inhibitor Library DNA Damage/DNA Repair compound Library Cell Cycle compound library | Click to Expand |
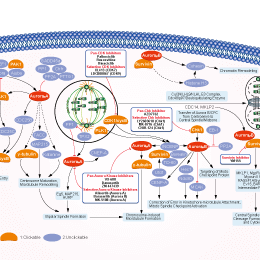
Signaling Pathway
Choose Selective Aurora Kinase Inhibitors
Cell Data
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| HepG2 cells | Cytotoxicity assay | 48 h | Cytotoxicity against human HepG2 cells after 48 hrs by MTT assay, TC50=0.2 μM | 24910766 | ||
| Click to View More Cell Line Experimental Data | ||||||
Biological Activity
| Description | Hesperadin potently inhibits Aurora B with IC50 of 250 nM in a cell-free assay. It markedly reduces the activity of AMPK, Lck, MKK1, MAPKAP-K1, CHK1 and PHK while it does not inhibit MKK1 activity in vivo. | ||||
|---|---|---|---|---|---|
| Targets |
|
| In vitro | ||||
| In vitro | Hesperadin inhibits the ability of immunoprecipitated Aurora B to phosphorylate histone H3 with IC50 of 250 nM and markedly reduces the activity of other kinases (AMPK, Lck, MKK1, MAPKAP-K1, CHK1, and PHK) at a concentration of 1 μM. In contrast, only 20-100 nM of Hesperadin is sufficient to induce the loss of mitotic histone H3-Ser10 phosphorylation in HeLa cells. Hesperadin treatment causes defects in mitosis and cytokinesis, leading to stoppage of proliferation of HeLa cells and polyploidization, which can be specifically ascribed to the inhibition of Aurora B function during the process of chromosome attachment. Hesperadin (100 nM) quickly overrides the mitotic arrest induced by taxol or monastrol but not by nocodazole. Hesperadin and nocodazole treatment in HeLa cells abolishes kinetochore localization of BubR1 and diminishes the intensity of Bub1 at kinetochores, suggesting that Aurora B function is required for efficient kinetochore recruitment of BubR1 and Bub1, which in turn might be necessary for prolonged checkpoint signaling. [1] Hesperadin prevents the phosphorylation of recombinant trypanosome histone H3 by the T. brucei Aurora kinase-1 (TbAUK1) from pathogenic Trypanosoma brucei with IC50 of 40 nM in vitro kinase assays. Hesperadin significantly inhibits cell growth of cultured infectious bloodstream forms (BF) with IC50 of 48 nM, and only weakly inhibits cell growth of insect stage procyclic forms (PF) with IC50 of 550 nM. [2] | |||
|---|---|---|---|---|
| Kinase Assay | The Aurora B kinase assay | |||
| For the Aurora B kinase assay, HeLa cells are lysed in a buffer containing 50 mM NaCl. The whole cell extract is spun at 13,000 rpm for 20 minutes at 4 °C using a table top centrifuge. The pellet obtained from 200 mg of whole cell extract is extracted again in 15 mL lysis buffer containing 250 mM NaCl in order to obtain active Aurora B kinase from mitotic chromatin. The low speed supernatant of the latter extract is used for immunoprecipitation. Monoclonal mouse anti–AIM-1, or mouse anti-HA, is coupled to GammaBind Plus Sepharose, and beads are rotated over-end in the extract for 90 minutes at 4 °C. Beads are washed, aliquoted, and washed in kinase buffer (20 mM Tris, pH 7.5, 150 mM NaCl, 10 mM MgCl2, 1 mM DTT, 10 mM NaF). The kinase assay is performed with 10 μL beads in 20 μL kinase buffer containing 5 μg histone H3, 10 μM ATP, 2.5 μCi [γ-32P]ATP, and different concentrations of Hesperadin for 20 minutes at 37 °C. SDS sample buffer is added, and samples are boiled and resolved by SDS-PAGE. The gel is dried, and the radioactive signal is detected by PhosphorImager analysis. The data is analyzed using ImageQuant software. | ||||
| Cell Research | Cell lines | HeLa cells and PtK1 cells | ||
| Concentrations | Final concentration ~500 nM | |||
| Incubation Time | 24 and 48 hours | |||
| Method | Cells are exposed to different concentrations of Hesperadin for 24 and 48 hours. At indicated time points, methanol-fixed cell samples are washed with PBS and subsequently stained in PI buffer (50 μg/mL propidium iodide, 10 mM Tris, pH 7.5, 5 mM MgCl2, 200 μg/mL RNase A) for 20-40 minutes at 37 °C. The DNA content is determined by flow cytometry. | |||
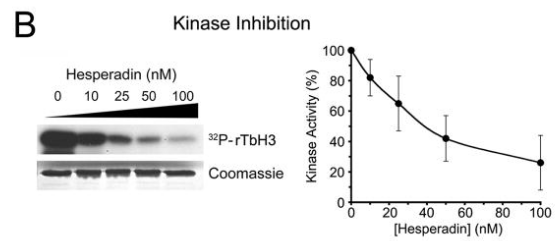
| Experimental Result Images | Methods | Biomarkers | Images | PMID |
| Western blot | trypanosome histone H3 |
|
19320832 | |
Chemical Information & Solubility
| Molecular Weight | 516.65 | Formula | C29H32N4O3S |
| CAS No. | 422513-13-1 | SDF | Download Hesperadin SDF |
| Smiles | CCS(=O)(=O)NC1=CC2=C(C=C1)NC(=C2C(=NC3=CC=C(C=C3)CN4CCCCC4)C5=CC=CC=C5)O | ||
| Storage (From the date of receipt) | |||
|
In vitro |
DMSO : 100 mg/mL ( (193.55 mM); Moisture-absorbing DMSO reduces solubility. Please use fresh DMSO.) Water : Insoluble Ethanol : Insoluble |
Molecular Weight Calculator |
|
In vivo Add solvents to the product individually and in order. |
In vivo Formulation Calculator |
||||
Preparing Stock Solutions
Molarity Calculator
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
mg/kg
g
μL
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
% DMSO
%
% Tween 80
% ddH2O
%DMSO
%
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Tech Support
Answers to questions you may have can be found in the inhibitor handling instructions. Topics include how to prepare stock solutions, how to store inhibitors, and issues that need special attention for cell-based assays and animal experiments.
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.
* Indicates a Required Field
Tags: buy Hesperadin | Hesperadin supplier | purchase Hesperadin | Hesperadin cost | Hesperadin manufacturer | order Hesperadin | Hesperadin distributor