research use only
MS436 BET inhibitor
Cat.No.S7305
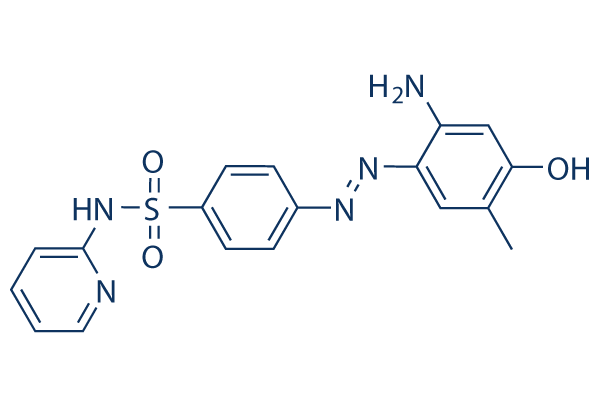
Chemical Structure
Molecular Weight: 383.42
Quality Control
Solubility
|
In vitro |
DMSO
: 55 mg/mL
(143.44 mM)
Ethanol : 1 mg/mL Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 383.42 | Formula | C18H17N5O3S |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 1395084-25-9 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N/A | Smiles | CC1=CC(=C(C=C1O)N)N=NC2=CC=C(C=C2)S(=O)(=O)NC3=CC=CC=N3 | ||
Mechanism of Action
| Targets/IC50/Ki |
BRD4 (1)
<0.085 μM(Ki)
BRD4 (2)
0.34 μM(Ki)
|
|---|---|
| In vitro |
MS436 exhibits low nanomolar affinity with preference for the first bromodomain over the second. This compound effectively inhibits BRD4 activity in NF-κB-directed production of nitric oxide and proinflammatory cytokine interleukin-6 in murine macrophages without siginificanty inhibition on cell viability. |
| Kinase Assay |
Fluorescence Anisotropy Binding Assay
|
|
Competition experiments are performed with a BrD protein (0.25–1 µM) and the fluorescent probe (80 nM), and increasing concentration of unlabeled competing ligand in a PBS buffer (pH 7.4) in total volume of 80 µL Measurements are obtained after a 1 hour incubation of the fluorescent ligand and the protein at 25°C with Safire 2 microplate reader. In a competition-binding assay, fluorescent ligand concentration is ≤ 2Kd, and protein concentration was set at which 50-80% of fluorescent ligand is bound. Dissociation constant of a competing ligand is calculated with the correction to Cheng-Prussoff equation introduced by Nicolovska-Coleska and colleagues. Assuming one-site competitive binding model, the equation used to calculate Ki’s from IC50 values recovered from fitting data using Prism.
|
References |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































