- Inhibitors
- Antibodies
- Compound Libraries
- New Products
- Contact Us
research use only
SU6656 Src inhibitor
Cat.No.S7774
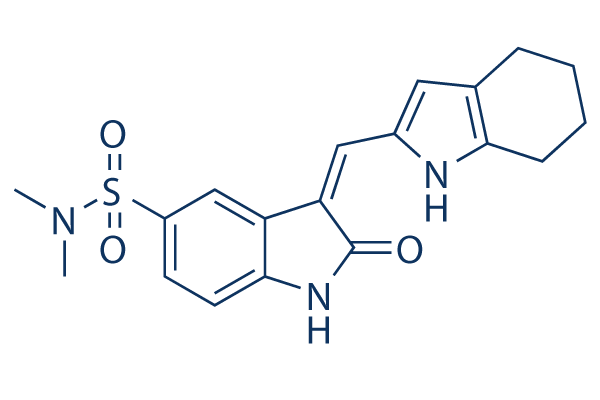
Chemical Structure
Molecular Weight: 371.45
Jump to
Quality Control
Batch:
Purity:
99.84%
99.84
| Related Targets | EGFR VEGFR JAK PDGFR FGFR HIF FLT FLT3 HER2 Bcr-Abl |
|---|---|
| Other Src Products | WH-4-023 Saracatinib (AZD0530) PP2 (AGL 1879) PP1 Src Inhibitor 1 RK 24466 Myristic Acid eCF506 Tolimidone (MLR-1023) UM-164 |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| Sf9 insect cells | Function assay | Inhibition of human recombinant His-tagged RET expressed in Sf9 insect cells, IC50=0.77 μM | ||||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 74 mg/mL
(199.21 mM)
Water : Insoluble Ethanol : Insoluble |
Molarity Calculator
Dilution Calculator
Molecular Weight Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
mg/kg
g
μL
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
%
DMSO
%
%
Tween 80
%
ddH2O
%
DMSO
+
%
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 371.45 | Formula | C19H21N3O3S |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 330161-87-0 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N/A | Smiles | CN(C)S(=O)(=O)C1=CC2=C(C=C1)NC(=O)C2=CC3=CC4=C(N3)CCCC4 | ||
Mechanism of Action
| Targets/IC50/Ki |
YES
20 nM
Lyn
130 nM
Fyn
170 nM
Src
280 nM
|
|---|---|
| In vitro |
In NIH 3T3 cells, SU 6656 inhibits the PDGF-stimulated S-phase induction with IC50 of 0.3-0.4 μM. SU 6656 also inhibits PDGF- and serum-mediated NIH 3T3 cell proliferation, as well as epidermal growth factor and colony-stimulating factor 1-stimulated DNA synthesis in normal and colony-stimulating factor 1 receptor transfected NIH 3T3 cells. SU6656 inhibits PDGF-stimulated c-Myc induction and ERK2 activation. Pretreating Jurkat T-cells with SU 6656 leads to increased VSV-G luciferase activity. SU 6656 impairs TGF-β-mediated upregulation of CTGF mRNA and protein in proximal epithelial HKC-8 cells, and also reduces CTGF expression in cells exposed to autocrine growth factors. SU 6656 interferes with Aurora kinase activity resulting in inhibition of cell division and formation multilobular nuclei. |
| Kinase Assay |
Biochemical kinase assays for IC50 determination and kinetic studies
|
|
IC50 measurements are made using poly-Glu–Tyr (4:1), or, in the case of Lck, poly-Lys–Tyr (4:1) as a peptide substrate. The divalent cation is 20 mM MgCl2 (in the case of Src, Fyn, Yes, Lyn, Csk, Frk, or Abl) or 10 mM MnCl2 (in the case of FGFR1, IGF1R, Lck, or Met). The final ATP concentrations are as follows: Src, 10 μM; Fyn, 6 μM; Yes, 100 μM; Lyn, 2 μM; Csk, 10 μM; Frk, 10 μM; Abl, 4 μM; FGFR1, 10 μM; IGF1R, 2 μM; Lck, 2 μM; Met, 5 μM; PDGFR, 6 μM. IC50 measurements of PDGFRb autophosphorylation are determined on immunoprecipitated PDGFRb. Km values are calculated using the Eadie-Hofstee method.
|
|
| In vivo |
SU 6656 markedly and dose-dependently attenuates -induced experimental nicotine withdrawal syndrome in mice measured in terms of WSS and anxiety score. |
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | phospho-c-Abl / c-Abl / phospho-c-Src / c-Src p-p38 / p38 / p-CREB / CREB p-STAT3 / STAT3 |
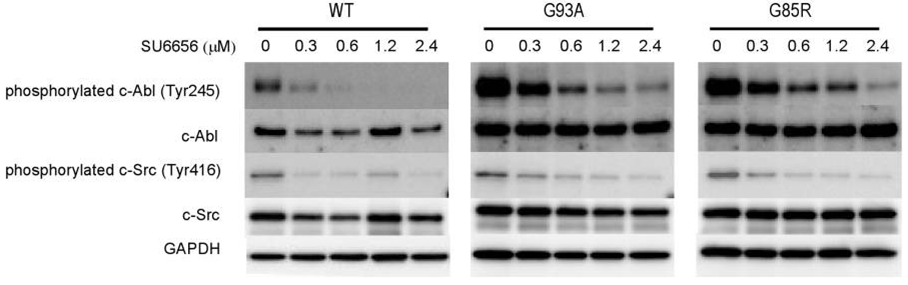
|
23049975 |
| Immunofluorescence | E-cadherin / Fibronectin p-p38 |
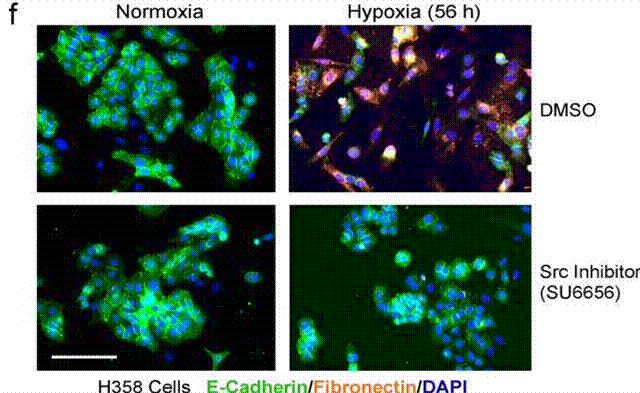
|
23246962 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































