research use only
UNC1999 Histone Methyltransferase inhibitor
Cat.No.S7165
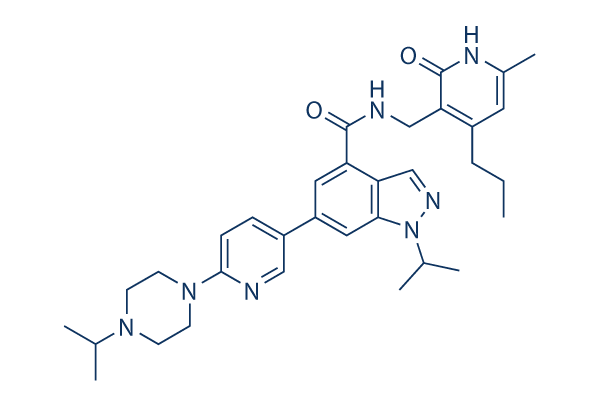
Chemical Structure
Molecular Weight: 569.74
Quality Control
| Related Targets | HDAC JAK BET PKC PARP HIF PRMT EZH2 AMPK Histone Acetyltransferase |
|---|---|
| Other Histone Methyltransferase Inhibitors | Pinometostat (EPZ5676) 3-Deazaneplanocin A (DZNep) Hydrochloride BIX-01294 Trihydrochloride EPZ015666 (GSK3235025) EPZ004777 MM-102 (HMTase Inhibitor IX) Chaetocin SGC 0946 EPZ005687 UNC0638 |
Cell Culture, Treatment & Working Concentration
| Cell Lines | Assay Type | Concentration | Incubation Time | Formulation | Activity Description | PMID |
|---|---|---|---|---|---|---|
| human MCF10A cells | Function assay | 72 h | Inhibition of EZH2 in human MCF10A cells assessed as reduction of H3K27me3 level after 72 hrs by Western blot analysis, IC50=0.124 μM | 25406853 | ||
| human MCF10A cells | Cytotoxic assay | Cytotoxicity against human MCF10A cells assessed as cell viability by Alamar Blue assay, EC50=19.2 μM | 25406853 | |||
| Click to View More Cell Line Experimental Data | ||||||
Solubility
|
In vitro |
DMSO
: 100 mg/mL
(175.51 mM)
Ethanol : 100 mg/mL Water : Insoluble |
Molarity Calculator
|
In vivo |
|||||
In vivo Formulation Calculator (Clear solution)
Step 1: Enter information below (Recommended: An additional animal making an allowance for loss during the experiment)
Step 2: Enter the in vivo formulation (This is only the calculator, not formulation. Please contact us first if there is no in vivo formulation at the solubility Section.)
Calculation results:
Working concentration: mg/ml;
Method for preparing DMSO master liquid: mg drug pre-dissolved in μL DMSO ( Master liquid concentration mg/mL, Please contact us first if the concentration exceeds the DMSO solubility of the batch of drug. )
Method for preparing in vivo formulation: Take μL DMSO master liquid, next addμL PEG300, mix and clarify, next addμL Tween 80, mix and clarify, next add μL ddH2O, mix and clarify.
Method for preparing in vivo formulation: Take μL DMSO master liquid, next add μL Corn oil, mix and clarify.
Note: 1. Please make sure the liquid is clear before adding the next solvent.
2. Be sure to add the solvent(s) in order. You must ensure that the solution obtained, in the previous addition, is a clear solution before proceeding to add the next solvent. Physical methods such
as vortex, ultrasound or hot water bath can be used to aid dissolving.
Chemical Information, Storage & Stability
| Molecular Weight | 569.74 | Formula | C33H43N7O2 |
Storage (From the date of receipt) | |
|---|---|---|---|---|---|
| CAS No. | 1431612-23-5 | Download SDF | Storage of Stock Solutions |
|
|
| Synonyms | N/A | Smiles | CCCC1=C(C(=O)NC(=C1)C)CNC(=O)C2=C3C=NN(C3=CC(=C2)C4=CN=C(C=C4)N5CCN(CC5)C(C)C)C(C)C | ||
Mechanism of Action
| Features |
The first orally bioavailable inhibitor against wild-type and mutant EZH2 as well as EZH1.
|
|---|---|
| Targets/IC50/Ki |
EZH2
(Cell-free assay) 2 nM
EZH1
(Cell-free assay) 45 nM
|
| In vitro |
UNC1999 is highly potent for both EZH2 Y641N and EZH2 Y641F mutants in vitro. This compound causes concentration-dependent reductions of H3K27me3 in MCF10A cells with IC50 of 124 nM , while shows low cellular toxicity. It displays potent, concentration-dependent inhibition of cell proliferation with EC50 of 633 nM in a DLBCL cell line harboring the EZH2Y641N mutant. In addition, biotinylated UNC1999 enriches EZH2 from HEK293T cell lysates, and thus may be used in chemoproteomics studies.
|
| Kinase Assay |
Scintillation Proximity Assay
|
|
Methyltransferase activity assays are performed by monitoring the incorporation of tritiumlabeled methyl group from S-adenosylmethionine (3H-SAM) to biotinylated peptide substrates using Scintillation Proximity Assay (SPA) for PRC2-EZH2 trimeric complex (EZH2:EED:SUZ12), PRC2-EZH1 pentameric complex (EZH1:EED:SUZ12:RBBP4:AEBP2), SETD7, G9a, GLP, SETDB1, SETD8, SUV420H1, SUV420H2, SUV39H2, MLL1 tetrameric complex (MLL:WDR5:RbBP5:ASH2L), PRMT1, PRMT3, PRMT5-MEP50 complex and SMYD2. The reaction buffer for SMYD2 and SMYD3 is 50 mM Tris pH 9.0, 5 mM DTT, 0.01% TritonX-100; for G9a, GLP and SUV39H2 is 25 mM potassium phosphate pH 8.0, 1 mM EDTA, 2 mM MgCl2 and 0.01% Triton X-100; and for other HMTs 20 mM Tris pH 8.0, 5 mM DTT, 0.01% TritonX-100. To stop the enzymatic reactions, 10 μL of 7.5 M guanidine hydrochloride is added, followed by 180 μL of buffer, mixed and transferred to a 384-well FlashPlate. After mixing, the reaction mixtures are incubated and the CPM counts are measured using Topcount plate reader. The CPM counts in the absence of this compound for each data set are defined as 100% activity. In the absence of the enzyme, the CPM counts in each data set are defined as background (0%). IC50 values are determined using compound concentrations ranging from 100 nM to 100 μM. The IC50 values are determined using SigmaPlot software. EZH2-Y641F assays are performed using 30 nM of enzyme in 20 mM Tris pH 8, 5 mM DTT, 0.01% Triton X-100, 5 μM SAM and 1 μM of H3 (1-24) peptide (same as for the wild-type PRC2-EZH2 complex). For DNMT1, the assay is performed using hemimethylated dsDNA as a substrate. The dsDNA substrate is prepared by annealing two complementary strands (biotintlated forward strand: BGAGCCCGTAAGCCCGTTCAGGTCG and reverse strand: CGACCTGAACGGGCTTACGGGCTC), synthesized by Eurofins MWG Operon. Reaction buffer is 20 mM Tris-HCl, pH 8.0, 5mM DTT, 0.01% Triton X-100. Methyltransferase activity assays for DOT1L is performed using Filter-plates. Reaction mixtures in 20 mM Tris-HCl, pH 8.0, 5 mM DTT, 2 mM MgCl2 and 0.01% Triton X-100 are incubated at room temperature for 1h, 100 μL 10% TCA is added, mixed and transferred to filter-plate. Plates are centrifuged at 2000 rpm for 2 min followed by 2 additional 10% TCA wash and one ethanol wash (180 μL) followed by centrifugation. Plates are dried and 100 μL MicroO is added and centrifuged. 70 μL MicroO is added and CPM are measured using Topcount plate reader.
|
|
| In vivo |
Treatment of UNC1999 (150 and 50 mg/kg, i.p.) results in the plasma concentrations of this compound above its cellular IC50 over 24 hours in vivo. In addition, it is also orally bioavailable in mice, which makes chronic animal studies more practical and convenient.
|
References |
|
Applications
| Methods | Biomarkers | Images | PMID |
|---|---|---|---|
| Western blot | H3K27Me3 / Cleaved PARP / LC3B H3K27me2 / H3K27me1 / H3K27ac / H3K4me3 / H3K9me2 / H3K36me2 |
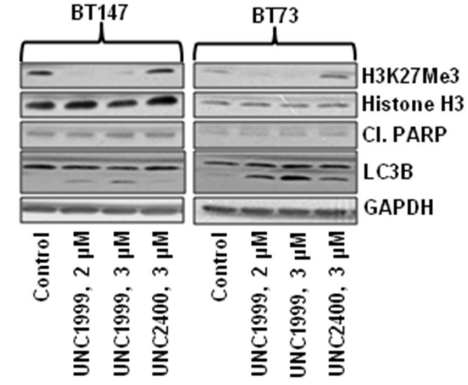
|
27449082 |
| Immunofluorescence | EZH2 / H3K27me3 NF-κB |
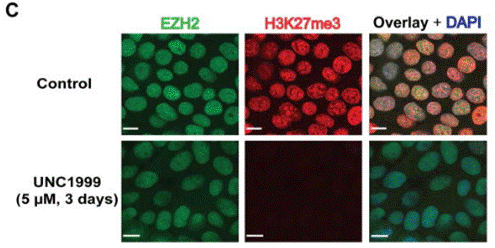
|
23614352 |
| Growth inhibition assay | Cell viability |
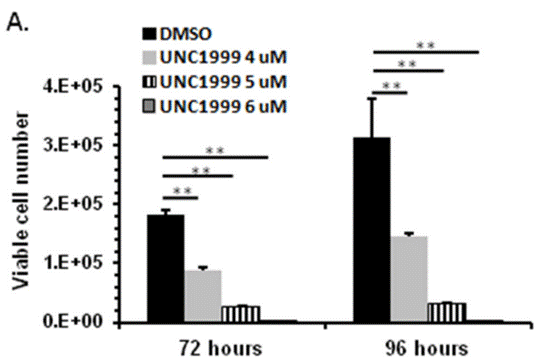
|
27449082 |
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.






































