research use only
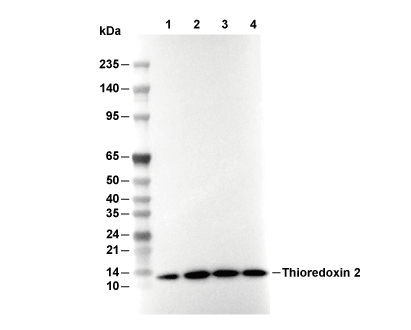
Thioredoxin 2 Antibody [B17P21]
Cat.No.: F5029
Application:
Reactivity:
Experiment Essentials
WB
Recommended SDS-PAGE separating gel concentration: 20%.
Recommended wet transfer conditions: 200 mA, 60 min,Recommended to use 0.22 μm PVDF membrane.
Recommended SDS-PAGE separating gel concentration: 20%.
Recommended wet transfer conditions: 200 mA, 60 min,Recommended to use 0.22 μm PVDF membrane.
Usage Information
| Dilution |
|---|
|
| Application |
|---|
| WB |
| Reactivity |
|---|
| Human, Mouse, Rat, Hamster, Monkey |
| Source |
|---|
| Rabbit Monoclonal Antibody |
| Storage Buffer |
|---|
| PBS, pH 7.2+50% Glycerol+0.05% BSA+0.01% NaN3 |
| Storage (from the date of receipt) |
|---|
| -20°C (avoid freeze-thaw cycles), 2 years |
| Predicted MW |
|---|
| 13 kDa |
| Positive Control | Hep G2 cells; Hep38 cells; HeLa cells; MOLT-4 cells; HL-60 cells; Daudi cells; Neuro-2a cells; C2C12 cells; NBT-II cells; KNRK cells; COS-7 cells |
|---|---|
| Negative Control |
Experimental Methods
| WB |
|---|
Experimental Protocol:
Sample preparation
1. Tissue: Lyse the tissue sample by adding an appropriate volume of ice-cold RIPA Lysis Buffer (containing Protease Inhibitor Cocktail),and homogenize the tissue at a low temperature. 2. Adherent cell: Aspirate the culture medium and wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 3. Suspension cell: Transfer the culture medium to a pre-cooled centrifuge tube. Centrifuge and aspirate the supernatant. Wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 4. Place the lysate into a pre-cooled microcentrifuge tube. Centrifuge at 4°C for 15 min. Collect the supernatant;
5. Remove a small volume of lysate to determine the protein concentration;
6. Combine the lysate with protein loading buffer. Boil 20 µL sample under 95-100°C for 5 min. Centrifuge for 5 min after cool down on ice.
Electrophoretic separation
1. According to the concentration of extracted protein, load appropriate amount of protein sample and marker onto SDS-PAGE gels for electrophoresis. Recommended separating gel (lower gel) concentration: 20%. Reference Table for Selecting SDS-PAGE Separation Gel Concentrations 2. Power up 80V for 30 minutes. Then the power supply is adjusted (110 V~150 V), the Marker is observed, and the electrophoresis can be stopped when the indicator band of the predyed protein Marker where the protein is located is properly separated. (Note that the current should not be too large when electrophoresis, too large current (more than 150 mA) will cause the temperature to rise, affecting the result of running glue. If high currents cannot be avoided, an ice bath can be used to cool the bath.)
Transfer membrane
1. Take out the converter, soak the clip and consumables in the pre-cooled converter;
2. Activate PVDF membrane with methanol for 1 min and rinse with transfer buffer;
3. Install it in the order of "black edge of clip - sponge - filter paper - filter paper - glue -PVDF membrane - filter paper - filter paper - sponge - white edge of clip"; 4. The protein was electrotransferred to PVDF membrane. ( 0.22 µm PVDF membrane is recommended )) Reference Table for Selecting PVDF Membrane Pore Size Specifications Recommended conditions for wet transfer: 200 mA, 60 min. ( Note that the transfer conditions can be adjusted according to the protein size. For high-molecular-weight proteins, a higher current and longer transfer time are recommended. However, ensure that the transfer tank remains at a low temperature to prevent gel melting.)
Block
1. After electrotransfer, wash the film with TBST at room temperature for 5 minutes;
2. Incubate the film in the blocking solution for 1 hour at room temperature;
3. Wash the film with TBST for 3 times, 5 minutes each time.
Antibody incubation
1. Use 5% skim milk powder to prepare the primary antibody working liquid (recommended dilution ratio for primary antibody 1:1000), gently shake and incubate with the film at 4°C overnight; 2. Wash the film with TBST 3 times, 5 minutes each time;
3. Add the secondary antibody to the blocking solution and incubate with the film gently at room temperature for 1 hour;
4. After incubation, wash the film with TBST 3 times for 5 minutes each time.
Antibody staining
1. Add the prepared ECL luminescent substrate (or select other color developing substrate according to the second antibody) and mix evenly;
2. Incubate with the film for 1 minute, remove excess substrate (keep the film moist), wrap with plastic film, and expose in the imaging system. |
Biological Description
| Specificity |
|---|
| Thioredoxin 2 Antibody [B17P21] detects endogenous levels of total Thioredoxin 2 protein. |
| Subcellular Location |
|---|
| Mitochondrion |
| Uniprot ID |
|---|
| Q99757 |
| Clone |
|---|
| B17P21 |
| Synonym(s) |
|---|
| Thioredoxin, mitochondrial; MTRX; Mt-Trx; Thioredoxin-2; TXN2; TRX2 |
| Background |
|---|
| Thioredoxin 2 (TRX2) belongs to the thioredoxin family of small redox proteins and localizes primarily to the mitochondrial matrix. This 166-residue protein adopts a classic thioredoxin fold with a central five-stranded beta-sheet (β1–β5) flanked by four alpha-helices (α1–α4), where the active site -Cys69-Pro72-Cys73- motif sits at the N-terminal end of α2 helix exposed for substrate access. The disulfide between Cys69–Cys73 reduces oxidized protein targets via nucleophilic attack, regenerating reduced TRX2 through thioredoxin reductase 2 (TRXR2) and NADPH. Dimerization at the N-terminus via hydrophobic contacts around Cys73 modulates activity, while an N-terminal mitochondrial targeting sequence (residues 1–55) directs import and cleavage. TRX2 detoxifies mitochondrial ROS by reducing peroxiredoxins PRX3/PRX5, preventing protein carbonylation and lipid peroxidation during oxidative stress from respiration or ischemia. It sustains mitochondrial integrity by preserving electron transport chain complexes I–IV and aconitase, averting cytochrome c release. TRX2 binds apoptosis signal-regulating kinase 1 (ASK1) via the active site thiols in reduced form, inhibiting its oligomerization and MAPK/JNK/p38 activation to block intrinsic apoptosis. Under H2O2 or hypoxia, oxidation dissociates TRX2-ASK1, unleashing caspase cascades. In protein folding, TRX2 assists protein disulfide isomerase PDIA4 in resolving misfolded interchain disulfides. Cardiolipin oxidation repair occurs via TRX2-PRXR3, protecting cristae structure. TRX2 knockout causes embryonic lethality at E9 with dilated cardiomyopathy, neural tube defects, and widespread apoptosis from ROS overload. Heart-specific deletion triggers failure under pressure overload via impaired bioenergetics. Cancer cells upregulate TRX2 to evade oxidative stress-induced death, fueling proliferation; inhibition synergizes with ROS inducers like doxorubicin. Friedreich ataxia features TRX2 deficits exacerbating frataxin loss and iron-sulfur cluster damage. Neurodegeneration links emerge from impaired ASK1 restraint in ALS and Parkinson's. Aging accelerates via chronic TRX2 decline, heightening mtDNA mutations. |
| References |
|---|
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.