research use only
SPT6 Antibody [P6D2]
Cat.No.: F7745
Application:
Reactivity:
Experiment Essentials
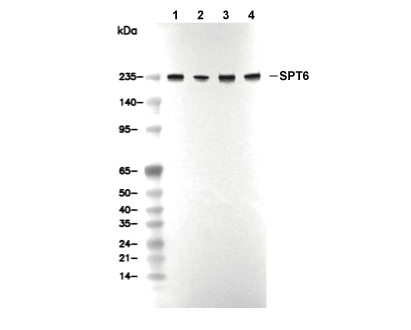
WB
Recommended SDS-PAGE separating gel concentration: 5%.
Recommended wet transfer conditions: 250 mA, 180 min.
Recommended SDS-PAGE separating gel concentration: 5%.
Recommended wet transfer conditions: 250 mA, 180 min.
Usage Information
| Dilution |
|---|
|
| Application |
|---|
| WB, IP, ChIP |
| Reactivity |
|---|
| Human, Mouse, Rat, Monkey |
| Source |
|---|
| Rabbit Monoclonal Antibody |
| Storage Buffer |
|---|
| PBS, pH 7.2+50% Glycerol+0.05% BSA+0.01% NaN3 |
| Storage (from the date of receipt) |
|---|
| -20°C (avoid freeze-thaw cycles), 2 years |
| Predicted MW |
|---|
| 210 kDa |
| Positive Control | Human breast carcinoma; HeLa cells; NIH/3T3 cells; H-4-II-E cells; COS-7 cells |
|---|---|
| Negative Control |
Experimental Methods
| WB |
|---|
Experimental Protocol:
Sample preparation
1. Tissue: Lyse the tissue sample by adding an appropriate volume of ice-cold RIPA/Nuclear Lysis Buffer (containing Protease Inhibitor Cocktail),and homogenize the tissue at a low temperature. 2. Adherent cell: Aspirate the culture medium and wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/Nuclear Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 3. Suspension cell: Transfer the culture medium to a pre-cooled centrifuge tube. Centrifuge and aspirate the supernatant. Wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/Nuclear Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 4. Place the lysate into a pre-cooled microcentrifuge tube. Centrifuge at 4°C for 15 min. Collect the supernatant;
5. Remove a small volume of lysate to determine the protein concentration;
6. Combine the lysate with protein loading buffer. Boil 20 µL sample under 95-100°C for 5 min. Centrifuge for 5 min after cool down on ice.
Electrophoretic separation
1. According to the concentration of extracted protein, load appropriate amount of protein sample and marker onto SDS-PAGE gels for electrophoresis. Recommended separating gel (lower gel) concentration: 5%. Reference Table for Selecting SDS-PAGE Separation Gel Concentrations 2. Power up 80V for 30 minutes. Then the power supply is adjusted (110 V~150 V), the Marker is observed, and the electrophoresis can be stopped when the indicator band of the predyed protein Marker where the protein is located is properly separated. (Note that the current should not be too large when electrophoresis, too large current (more than 150 mA) will cause the temperature to rise, affecting the result of running glue. If high currents cannot be avoided, an ice bath can be used to cool the bath.)
Transfer membrane
1. Take out the converter, soak the clip and consumables in the pre-cooled converter;
2. Activate PVDF membrane with methanol for 1 min and rinse with transfer buffer;
3. Install it in the order of "black edge of clip - sponge - filter paper - filter paper - glue -PVDF membrane - filter paper - filter paper - sponge - white edge of clip"; 4. The protein was electrotransferred to PVDF membrane. ( 0.45 µm PVDF membrane is recommended ) Reference Table for Selecting PVDF Membrane Pore Size Specifications Recommended conditions for wet transfer: 250 mA, 180 min. ( Note that the transfer conditions can be adjusted according to the protein size. For high-molecular-weight proteins, a higher current and longer transfer time are recommended. However, ensure that the transfer tank remains at a low temperature to prevent gel melting.)
Block
1. After electrotransfer, wash the film with TBST at room temperature for 5 minutes;
2. Incubate the film in the blocking solution for 1 hour at room temperature;
3. Wash the film with TBST for 3 times, 5 minutes each time.
Antibody incubation
1. Use 5% skim milk powder to prepare the primary antibody working liquid (recommended dilution ratio for primary antibody 1:1000), gently shake and incubate with the film at 4°C overnight; 2. Wash the film with TBST 3 times, 5 minutes each time;
3. Add the secondary antibody to the blocking solution and incubate with the film gently at room temperature for 1 hour;
4. After incubation, wash the film with TBST 3 times for 5 minutes each time.
Antibody staining
1. Add the prepared ECL luminescent substrate (or select other color developing substrate according to the second antibody) and mix evenly;
2. Incubate with the film for 1 minute, remove excess substrate (keep the film moist), wrap with plastic film, and expose in the imaging system. |
Biological Description
| Specificity |
|---|
| SPT6 Antibody [P6D2] detects endogenous levels of total SPT6 protein. |
| Subcellular Location |
|---|
| Nucleus |
| Uniprot ID |
|---|
| Q7KZ85 |
| Clone |
|---|
| P6D2 |
| Synonym(s) |
|---|
| Transcription elongation factor SPT6; SUPT6H |
| Background |
|---|
| SPT6 serves as a conserved histone chaperone and transcription elongation factor that couples RNA polymerase II activity with chromatin dynamics across eukaryotes. It organizes tandem tetratricopeptide repeats for binding phosphorylated Ser2 CTD of Rpb1 alongside an acidic N-terminal domain and SH2-like region that recruit histone H3 and processing factors like SPN1 for coordinated mRNP packaging. During transcriptional elongation, SPT6 travels with transcribing Pol II to deposit H3-H4 dimers onto newly synthesized DNA ahead of the polymerase, simultaneously retaining post-translationally modified H3 molecules through direct binding that prevents their displacement and maintains K4 methylation and K36 trimethylation marks essential for preventing cryptic initiation within gene bodies. Interaction with the Paf1 complex and Spt5 enhances Pol II processivity by facilitating nucleosome reassembly that alleviates backtracking while coordinating H3K36me3 deposition via direct recruitment of Set2 methyltransferase, thereby coupling elongation rate with downstream m6A modification and 3' end processing. SPT6 also cooperates with HDACs like Clr3 to suppress histone turnover and enforce silencing at pericentromeric repeats through retention of deacetylated H3 tails. SPT6 sustains high-fidelity gene expression during heat shock response and developmental transitions, with ubiquitous expression reflecting roles in maintaining genome stability across proliferating and differentiated states. Depletion triggers bidirectional cryptic transcription from intergenic regions and gene bodies due to nucleosome depletion, compromising mRNA quality control and viability. |
| References |
|---|
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.