research use only
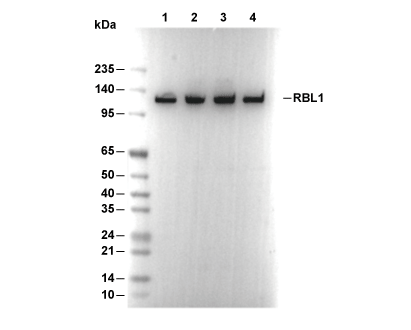
RBL1 Antibody [M18D3]
Cat.No.: F8644
Application:
Reactivity:
Experiment Essentials
WB
Recommended SDS-PAGE separating gel concentration: 5%.
Recommended SDS-PAGE separating gel concentration: 5%.
Usage Information
| Dilution |
|---|
|
| Application |
|---|
| WB, IP |
| Reactivity |
|---|
| Human, Monkey |
| Source |
|---|
| Rabbit Monoclonal Antibody |
| Storage Buffer |
|---|
| PBS, pH 7.2+50% Glycerol+0.05% BSA+0.01% NaN3 |
| Storage (from the date of receipt) |
|---|
| -20°C (avoid freeze-thaw cycles), 2 years |
| Predicted MW |
|---|
| 121 kDa |
| Positive Control | MCF7 cells; HT-29 cells; THP-1 cells; COS-7 cells |
|---|---|
| Negative Control |
Experimental Methods
| WB |
|---|
Experimental Protocol:
Sample preparation
1. Tissue: Lyse the tissue sample by adding an appropriate volume of ice-cold RIPA/Nuclear Lysis Buffer (containing Protease Inhibitor Cocktail),and homogenize the tissue at a low temperature. 2. Adherent cell: Aspirate the culture medium and wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/Nuclear Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 3. Suspension cell: Transfer the culture medium to a pre-cooled centrifuge tube. Centrifuge and aspirate the supernatant. Wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/Nuclear Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 4. Place the lysate into a pre-cooled microcentrifuge tube. Centrifuge at 4°C for 15 min. Collect the supernatant;
5. Remove a small volume of lysate to determine the protein concentration;
6. Combine the lysate with protein loading buffer. Boil 20 µL sample under 95-100°C for 5 min. Centrifuge for 5 min after cool down on ice.
Electrophoretic separation
1. According to the concentration of extracted protein, load appropriate amount of protein sample and marker onto SDS-PAGE gels for electrophoresis. Recommended separating gel (lower gel) concentration: 5%. Reference Table for Selecting SDS-PAGE Separation Gel Concentrations 2. Power up 80V for 30 minutes. Then the power supply is adjusted (110 V~150 V), the Marker is observed, and the electrophoresis can be stopped when the indicator band of the predyed protein Marker where the protein is located is properly separated. (Note that the current should not be too large when electrophoresis, too large current (more than 150 mA) will cause the temperature to rise, affecting the result of running glue. If high currents cannot be avoided, an ice bath can be used to cool the bath.)
Transfer membrane
1. Take out the converter, soak the clip and consumables in the pre-cooled converter;
2. Activate PVDF membrane with methanol for 1 min and rinse with transfer buffer;
3. Install it in the order of "black edge of clip - sponge - filter paper - filter paper - glue -PVDF membrane - filter paper - filter paper - sponge - white edge of clip"; 4. The protein was electrotransferred to PVDF membrane. ( 0.45 µm PVDF membrane is recommended ) Reference Table for Selecting PVDF Membrane Pore Size Specifications Recommended conditions for wet transfer: 200 mA, 120 min. ( Note that the transfer conditions can be adjusted according to the protein size. For high-molecular-weight proteins, a higher current and longer transfer time are recommended. However, ensure that the transfer tank remains at a low temperature to prevent gel melting.)
Block
1. After electrotransfer, wash the film with TBST at room temperature for 5 minutes;
2. Incubate the film in the blocking solution for 1 hour at room temperature;
3. Wash the film with TBST for 3 times, 5 minutes each time.
Antibody incubation
1. Use 5% skim milk powder to prepare the primary antibody working liquid (recommended dilution ratio for primary antibody 1:1000), gently shake and incubate with the film at 4°C overnight; 2. Wash the film with TBST 3 times, 5 minutes each time;
3. Add the secondary antibody to the blocking solution and incubate with the film gently at room temperature for 1 hour;
4. After incubation, wash the film with TBST 3 times for 5 minutes each time.
Antibody staining
1. Add the prepared ECL luminescent substrate (or select other color developing substrate according to the second antibody) and mix evenly;
2. Incubate with the film for 1 minute, remove excess substrate (keep the film moist), wrap with plastic film, and expose in the imaging system. |
Biological Description
| Specificity |
|---|
| RBL1 Antibody [M18D3] detects endogenous levels of total RBL1 protein. |
| Subcellular Location |
|---|
| Nucleus |
| Uniprot ID |
|---|
| P28749 |
| Clone |
|---|
| M18D3 |
| Synonym(s) |
|---|
| Retinoblastoma-like protein 1; p107; RBL1 |
| Background |
|---|
| RBL1, known as p107, belongs to the retinoblastoma (Rb) family of pocket proteins alongside Rb and RBL2/p130, where it serves as a central regulator of cell cycle progression through transcriptional repression. RBL1 contains a conserved pocket domain flanked by C-terminal and N-terminal regions that facilitate binding to E2F4/5 transcription factors and LXCXE motif-containing proteins, enabling recruitment of repressive complexes to promoter elements. Hypophosphorylated RBL1 anchors the DREAM complex, comprising E2F4/5, DP1, MuvB core (LIN proteins, RBBP4), and CHK2 kinase, to cell cycle gene promoters bearing CHR elements during G0/G1 quiescence, where it enforces silencing of S-phase entry genes like CCNA2, CDC2, and TK1 alongside G2/M regulators such as PLK1 and AURKB through HDAC-dependent chromatin compaction. Cyclin D-CDK4/6 and cyclin E-CDK2 sequentially phosphorylate RBL1 at multiple sites upon mitogenic stimulation, disrupting pocket interactions and releasing E2F transactivators to drive E2F-responsive transcription while permitting MuvB redeployment to B-MYB-FOXM1 for mitotic gene activation. RBL1 further integrates with p53 signaling via direct interaction that stabilizes p21-mediated CDK inhibition, amplifying G1 arrest during DNA damage, and cooperates with BMI1 to modulate Polycomb repressive complex 1 activity at lineage-specific loci during differentiation. RBL1 maintains quiescence in stem cells and enforces senescence-associated gene repression in post-mitotic tissues, with dynamic phosphorylation cycling calibrated by PP1/PP2A phosphatases to tissue-specific demands in liver regeneration and neuronal maturation. Loss of RBL1 elevates E2F activity and deregulates DREAM targets in osteosarcoma and breast carcinoma, promoting unrestrained proliferation through G1/S checkpoint collapse. |
| References |
|---|
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.