research use only
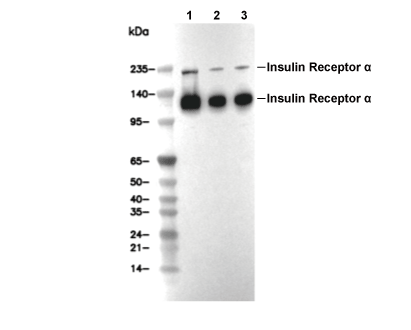
Insulin Receptor α Antibody [J24M5]
Cat.No.: F3503
Application:
Reactivity:
Experiment Essentials
WB
Recommended SDS-PAGE separating gel concentration: 5%.
Recommended SDS-PAGE separating gel concentration: 5%.
Usage Information
| Dilution |
|---|
|
| Application |
|---|
| WB |
| Reactivity |
|---|
| Human, Mouse, Rat |
| Source |
|---|
| Rabbit Monoclonal Antibody |
| Storage Buffer |
|---|
| PBS, pH 7.2+50% Glycerol+0.05% BSA+0.01% NaN3 |
| Storage (from the date of receipt) |
|---|
| -20°C (avoid freeze-thaw cycles), 2 years |
| Predicted MW |
|---|
| 135 kDa, 220 kDa |
| Positive Control | HCC1419 cells; Hep382.1-7 cells; Hepa 1-6 cells; H-4-II-E cells |
|---|---|
| Negative Control |
Experimental Methods
| WB |
|---|
Experimental Protocol:
Sample preparation
1. Tissue: Lyse the tissue sample by adding an appropriate volume of ice-cold RIPA/NP-40 Lysis Buffer (containing Protease Inhibitor Cocktail),and homogenize the tissue at a low temperature. 2. Adherent cell: Aspirate the culture medium and wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/NP-40 Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 3. Suspension cell: Transfer the culture medium to a pre-cooled centrifuge tube. Centrifuge and aspirate the supernatant. Wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/NP-40 Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 4. Place the lysate into a pre-cooled microcentrifuge tube. Centrifuge at 4°C for 15 min. Collect the supernatant;
5. Remove a small volume of lysate to determine the protein concentration;
6. Combine the lysate with protein loading buffer. Boil 20 µL sample under 95-100°C for 5 min. Centrifuge for 5 min after cool down on ice.
Electrophoretic separation
1. According to the concentration of extracted protein, load appropriate amount of protein sample and marker onto SDS-PAGE gels for electrophoresis. Recommended separating gel (lower gel) concentration: 5%. Reference Table for Selecting SDS-PAGE Separation Gel Concentrations 2. Power up 80V for 30 minutes. Then the power supply is adjusted (110 V~150 V), the Marker is observed, and the electrophoresis can be stopped when the indicator band of the predyed protein Marker where the protein is located is properly separated. (Note that the current should not be too large when electrophoresis, too large current (more than 150 mA) will cause the temperature to rise, affecting the result of running glue. If high currents cannot be avoided, an ice bath can be used to cool the bath.)
Transfer membrane
1. Take out the converter, soak the clip and consumables in the pre-cooled converter;
2. Activate PVDF membrane with methanol for 1 min and rinse with transfer buffer;
3. Install it in the order of "black edge of clip - sponge - filter paper - filter paper - glue -PVDF membrane - filter paper - filter paper - sponge - white edge of clip"; 4. The protein was electrotransferred to PVDF membrane. ( 0.45 µm PVDF membrane is recommended ) Reference Table for Selecting PVDF Membrane Pore Size Specifications Recommended conditions for wet transfer: 200 mA, 120 min. ( Note that the transfer conditions can be adjusted according to the protein size. For high-molecular-weight proteins, a higher current and longer transfer time are recommended. However, ensure that the transfer tank remains at a low temperature to prevent gel melting.)
Block
1. After electrotransfer, wash the film with TBST at room temperature for 5 minutes;
2. Incubate the film in the blocking solution for 1 hour at room temperature;
3. Wash the film with TBST for 3 times, 5 minutes each time.
Antibody incubation
1. Use 5% skim milk powder to prepare the primary antibody working liquid (recommended dilution ratio for primary antibody 1:1000), gently shake and incubate with the film at 4°C overnight; 2. Wash the film with TBST 3 times, 5 minutes each time;
3. Add the secondary antibody to the blocking solution and incubate with the film gently at room temperature for 1 hour;
4. After incubation, wash the film with TBST 3 times for 5 minutes each time.
Antibody staining
1. Add the prepared ECL luminescent substrate (or select other color developing substrate according to the second antibody) and mix evenly;
2. Incubate with the film for 1 minute, remove excess substrate (keep the film moist), wrap with plastic film, and expose in the imaging system. |
Biological Description
| Specificity |
|---|
| Insulin Receptor α Antibody [J24M5] detects endogenous levels of total Insulin Receptor α protein. |
| Subcellular Location |
|---|
| Cell membrane, Endosome, Lysosome, Membrane |
| Uniprot ID |
|---|
| P06213 |
| Clone |
|---|
| J24M5 |
| Synonym(s) |
|---|
| Insulin receptor precursor; IR; CD220; INSR |
| Background |
|---|
| The insulin receptor belongs to the receptor tyrosine kinase family and exists as a heterotetrameric complex of two extracellular α-subunits and two transmembrane β-subunits linked by disulfide bonds. The α-subunit spans residues 1-735 and binds insulin at high affinity, while the β-subunit (residues 736-1382) contains the tyrosine kinase domain spanning approximately residues 980-1255. Insulin binding induces conformational changes that activate the intrinsic kinase activity, leading to autophosphorylation on multiple tyrosine residues in the β-subunit activation loop (e.g., Tyr1158, Tyr1162, Tyr1163). Phosphorylated tyrosines serve as docking sites for adaptor proteins like IRS and Shc. IRS recruits PI3K to launch the PI3K/Akt pathway, driving glucose uptake through GLUT4 translocation, glycogen synthesis by inhibiting GSK3, and protein synthesis via mTOR. Meanwhile, Shc triggers the MAPK/ERK cascade to spur cell proliferation and gene expression through Elk-1 and other transcription factors. Together, these signals control metabolism, growth, and survival. Dysregulation underlies type 2 diabetes, where insulin resistance, hyperinsulinemia, and receptor downregulation weaken signaling. Cancer arises when PI3K/Akt/mTOR and MAPK pathways ramp up from overexpression or mutations. Hybrid receptors with IGF1R also bind IGF-1 tightly, blending metabolic control with mitogenic drive. |
| References |
|---|
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.