research use only
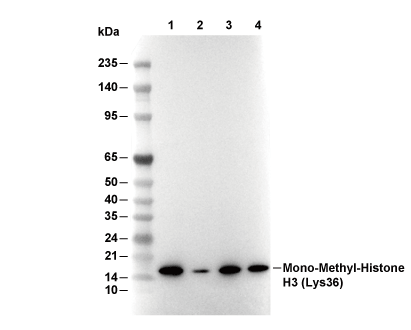
Histone H3 (mono methyl Lys36) Antibody [D2L24]
Cat.No.: F4594
Application:
Reactivity:
Experiment Essentials
WB
Recommended wet transfer conditions: 200 mA, 60 min,Recommended to use 0.22 μm PVDF membrane.
Recommended wet transfer conditions: 200 mA, 60 min,Recommended to use 0.22 μm PVDF membrane.
Usage Information
| Dilution |
|---|
|
| Application |
|---|
| WB,IP, IF, FCM, ChIP |
| Reactivity |
|---|
| Human, Mouse, Rat, Monkey |
| Source |
|---|
| Rabbit Monoclonal Antibody |
| Storage Buffer |
|---|
| PBS, pH 7.2+50% Glycerol+0.05% BSA+0.01% NaN3 |
| Storage (from the date of receipt) |
|---|
| -20°C (avoid freeze-thaw cycles), 2 years |
| Predicted MW |
|---|
| 17 kDa |
| Positive Control | HeLa cells; NIH/3T3 cells; H-4-II-E cells; COS-7 cells |
|---|---|
| Negative Control |
Experimental Methods
| WB |
|---|
Experimental Protocol:
Sample preparation
1. Tissue: Lyse the tissue sample by adding an appropriate volume of ice-cold RIPA/Nuclear Lysis Buffer (containing Protease Inhibitor Cocktail),and homogenize the tissue at a low temperature. 2. Adherent cell: Aspirate the culture medium and wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/Nuclear Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 3. Suspension cell: Transfer the culture medium to a pre-cooled centrifuge tube. Centrifuge and aspirate the supernatant. Wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/Nuclear Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 4. Place the lysate into a pre-cooled microcentrifuge tube. Centrifuge at 4°C for 15 min. Collect the supernatant;
5. Remove a small volume of lysate to determine the protein concentration;
6. Combine the lysate with protein loading buffer. Boil 20 µL sample under 95-100°C for 5 min. Centrifuge for 5 min after cool down on ice.
Electrophoretic separation
1. According to the concentration of extracted protein, load appropriate amount of protein sample and marker onto SDS-PAGE gels for electrophoresis. Recommended separating gel (lower gel) concentration: 10%. Reference Table for Selecting SDS-PAGE Separation Gel Concentrations 2. Power up 80V for 30 minutes. Then the power supply is adjusted (110 V~150 V), the Marker is observed, and the electrophoresis can be stopped when the indicator band of the predyed protein Marker where the protein is located is properly separated. (Note that the current should not be too large when electrophoresis, too large current (more than 150 mA) will cause the temperature to rise, affecting the result of running glue. If high currents cannot be avoided, an ice bath can be used to cool the bath.)
Transfer membrane
1. Take out the converter, soak the clip and consumables in the pre-cooled converter;
2. Activate PVDF membrane with methanol for 1 min and rinse with transfer buffer;
3. Install it in the order of "black edge of clip - sponge - filter paper - filter paper - glue -PVDF membrane - filter paper - filter paper - sponge - white edge of clip"; 4. The protein was electrotransferred to PVDF membrane. ( 0.22 µm PVDF membrane is recommended )) Reference Table for Selecting PVDF Membrane Pore Size Specifications Recommended conditions for wet transfer: 200 mA, 60 min. ( Note that the transfer conditions can be adjusted according to the protein size. For high-molecular-weight proteins, a higher current and longer transfer time are recommended. However, ensure that the transfer tank remains at a low temperature to prevent gel melting.)
Block
1. After electrotransfer, wash the film with TBST at room temperature for 5 minutes;
2. Incubate the film in the blocking solution for 1 hour at room temperature;
3. Wash the film with TBST for 3 times, 5 minutes each time.
Antibody incubation
1. Use 5% skim milk powder to prepare the primary antibody working liquid (recommended dilution ratio for primary antibody 1:1000), gently shake and incubate with the film at 4°C overnight; 2. Wash the film with TBST 3 times, 5 minutes each time;
3. Add the secondary antibody to the blocking solution and incubate with the film gently at room temperature for 1 hour;
4. After incubation, wash the film with TBST 3 times for 5 minutes each time.
Antibody staining
1. Add the prepared ECL luminescent substrate (or select other color developing substrate according to the second antibody) and mix evenly;
2. Incubate with the film for 1 minute, remove excess substrate (keep the film moist), wrap with plastic film, and expose in the imaging system. |
| IF |
|---|
Experimental Protocol:
Sample Preparation
1. Adherent Cells: Place a clean, sterile coverslip in a culture dish. Once the cells grow to near confluence as a monolayer, remove the coverslip for further use.
2. Suspension Cells: Seed the cells onto a clean, sterile slide coated with poly-L-lysine.
3. Frozen Sections: Allow the slide to thaw at room temperature. Wash it with pure water or PBS for 2 times, 3 minutes each time.
4. Paraffin Sections: Deparaffinization and rehydration. Wash the slide with pure water or PBS for 3 times, 3 minutes each time. Then perform antigen retrieval.
Fixation
1. Fix the cell coverslips/spots or tissue sections at room temperature using a fixative such as 4% paraformaldehyde (4% PFA) for 10-15 minutes.
2. Wash the sample with PBS for 3 times, 3 minutes each time.
Permeabilization
1.Add a detergent such as 0.1–0.3% Triton X-100 to the sample and incubate at room temperature for 10–20 minutes.
(Note: This step is only required for intracellular antigens. For antigens expressed on the cell membrane, this step is unnecessary.)
Wash the sample with PBS for 3 times, 3 minutes each time.
Blocking
Add blocking solution and incubate at room temperature for at least 1 hour. (Common blocking solutions include: serum from the same source as the secondary antibody, BSA, or goat serum.)
Note: Ensure the sample remains moist during and after the blocking step to prevent drying, which can lead to high background.
Immunofluorescence Staining (Day 1)
1. Remove the blocking solution and add the diluted primary antibody.
2. Incubate the sample in a humidified chamber at 4°C overnight.
Immunofluorescence Staining (Day 2)
1. Remove the primary antibody and wash with PBST for 3 times, 5 minutes each time.
2. Add the diluted fluorescent secondary antibody and incubate in the dark at 4°C for 1–2 hours.
3. Remove the secondary antibody and wash with PBST for 3 times, 5 minutes each time.
4. Add diluted DAPI and incubate at room temperature in the dark for 5–10 minutes.
5. Wash with PBST for 3 times, 5 minutes each time.
Mounting
1. Mount the sample with an anti-fade mounting medium.
2. Allow the slide to dry at room temperature overnight in the dark.
3. Store the slide in a slide storage box at 4°C, protected from light.
|
Biological Description
| Specificity |
|---|
| Histone H3 (mono methyl Lys36) Antibody [D2L24] detects endogenous levels of total Histone H3 protein only when it is methylated at Lys36. |
| Subcellular Location |
|---|
| Chromosome, Nucleosome core, Nucleus |
| Uniprot ID |
|---|
| P68431 |
| Clone |
|---|
| D2L24 |
| Synonym(s) |
|---|
| H3; H3 clustered histone 1; H3 histone family, member A; H3/A; H31; H3C1; H3C10; H3C11; H3C12; H3C2; H3C3; H3C4; H3C6; H3C7; H3C8; H3FA; H3FB; H3FC; H3FC HIST1H3C; H3FD; H3FF; H3FH; H3FI; H3FJ; H3FK |
| Background |
|---|
| Histone H3 monomethyl Lys36 (H3K36me1) forms part of the core histone family that wraps DNA into nucleosomes alongside H2A, H2B, and H4 to create chromatin's basic unit. Lysine 36 lies exposed on the H3 globular domain, where methyltransferases like NSD1 add a single methyl group during active gene transcription. H3K36me1 builds up in gene bodies as RNA polymerase II elongates, recruiting PWWP proteins like LEDGF and chromodomain readers to lock in transcriptional accuracy. The mark guides the Rpd3S deacetylase complex to clear acetyl marks behind the polymerase, blocking unwanted transcription from internal gene promoters. It teams up with the Paf1 complex and Asf1 chaperone to keep polymerase moving smoothly while reassembling nucleosomes in its wake. H3K36me1 shapes splicing outcomes by drawing MRG15-PTB complexes that skip weak exons, as seen in FGFR2 regulation. On male X chromosomes, it binds MSL complex chromodomains to balance gene dosage through hypertranscription. At DNA double-strand breaks, H3K36me1 pulls in NHEJ repair factors like 53BP1 and Ku70 for faithful repair. SETD2 builds on H3K36me1 to create the trimethyl form, linking both to serine-2 phosphorylation on polymerase CTD. Without proper H3K36 methyltransferases, cells suffer faulty mRNA processing and genomic chaos. NSD1 loss causes Sotos syndrome by scrambling H3K36me1 control over growth genes. NSD2 excess from t(4;14) in multiple myeloma floods cells with H3K36me1 to fuel cancer. SETD2 defects in renal cell carcinoma shatter H3K36me1 oversight of transcription quality. |
| References |
|---|
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.