research use only
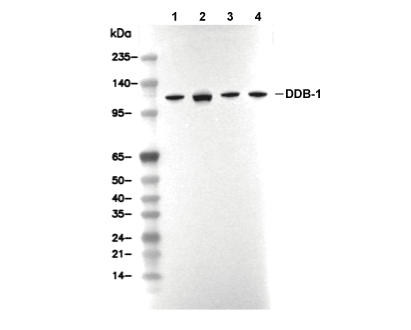
DDB1 Antibody [M17E15]
Cat.No.: F5000
Application:
Reactivity:
Usage Information
| Dilution |
|---|
|
| Application |
|---|
| WB |
| Reactivity |
|---|
| Human, Mouse, Rat, Monkey |
| Source |
|---|
| Rabbit Monoclonal Antibody |
| Storage Buffer |
|---|
| PBS, pH 7.2+50% Glycerol+0.05% BSA+0.01% NaN3 |
| Storage (from the date of receipt) |
|---|
| -20°C (avoid freeze-thaw cycles), 2 years |
| Predicted MW |
|---|
| 127 kDa |
Biological Description
| Specificity |
|---|
| DDB1 Antibody [M17E15] detects endogenous levels of total DDB1 protein. |
| Clone |
|---|
| M17E15 |
| Synonym(s) |
|---|
| DNA damage-binding protein 1; DDB p127 subunit; DNA damage-binding protein a (DDBa); XAP-1; UV-DDB 1; XPE-binding factor (XPE-BF); Xeroderma pigmentosum group E-complementing protein (XPCe); DDB1; XAP1 |
| Background |
|---|
| DDB1 (damage-specific DNA binding protein 1), the scaffold subunit of the CRL4 ubiquitin E3 ligase family, orchestrates nucleotide excision repair initiation, histone eviction, and cell cycle checkpoint enforcement by recruiting diverse substrate receptors to damaged chromatin sites. Its propeller blade architecture with a central β-sheet core accommodates the DDB2 WD40 domain for lesion-specific recognition of cyclobutane pyrimidine dimers and 6-4 photoproducts, while the lateral surface docks CUL4A-ROC1 to catalyze K48/K63-linked polyubiquitination of repair factors like XPC, DDB2 itself, and histones H3/H4. Upon UV exposure, DDB1 translocates from soluble to tightly chromatin-bound fractions in a DDB2-dependent manner, facilitating ubiquitination-mediated displacement of core histones to expose lesions for XPC loading and TFIIH recruitment; this histone code handover licenses dual incision while preventing error-prone repair. DDB1 scaffolds additional DCAFs like VprBP for HIV integration and CDT1 for replication licensing, integrating DNA damage with G1/S transition via p21/p27 degradation. DDB1 maintains genomic stability across proliferating tissues, with acute depletion triggering p53-dependent apoptosis in neural progenitors and hematopoietic stem cells through unresolved replication stress. Mutations underlie xeroderma pigmentosum E through defective UV-DDB assembly, while CUL4-DDB1 overexpression drives tumorigenesis via CDT1 hyperdegradation and replication origin firing. |
| References |
|---|
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.