research use only
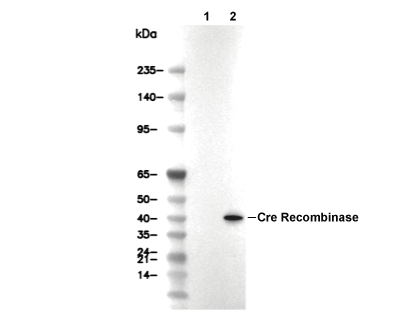
Cre Recombinase Antibody [E18L5]
Cat.No.: F4665
Application:
Reactivity:
Experiment Essentials
WB
Recommended wet transfer conditions: 200 mA, 60 min.
Recommended wet transfer conditions: 200 mA, 60 min.
Usage Information
| Dilution |
|---|
|
| Application |
|---|
| WB, FCM |
| Reactivity |
|---|
| All Species Expected |
| Source |
|---|
| Rabbit Monoclonal Antibody |
| Storage Buffer |
|---|
| PBS, pH 7.2+50% Glycerol+0.05% BSA+0.01% NaN3 |
| Storage (from the date of receipt) |
|---|
| -20°C (avoid freeze-thaw cycles), 2 years |
| Predicted MW |
|---|
| 37 kDa |
Experimental Methods
| WB |
|---|
Experimental Protocol:
Sample preparation
1. Tissue: Lyse the tissue sample by adding an appropriate volume of ice-cold RIPA/NP-40 Lysis Buffer (containing Protease Inhibitor Cocktail),and homogenize the tissue at a low temperature. 2. Adherent cell: Aspirate the culture medium and wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/NP-40 Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 3. Suspension cell: Transfer the culture medium to a pre-cooled centrifuge tube. Centrifuge and aspirate the supernatant. Wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/NP-40 Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 4. Place the lysate into a pre-cooled microcentrifuge tube. Centrifuge at 4°C for 15 min. Collect the supernatant;
5. Remove a small volume of lysate to determine the protein concentration;
6. Combine the lysate with protein loading buffer. Boil 20 µL sample under 95-100°C for 5 min. Centrifuge for 5 min after cool down on ice.
Electrophoretic separation
1. According to the concentration of extracted protein, load appropriate amount of protein sample and marker onto SDS-PAGE gels for electrophoresis. Recommended separating gel (lower gel) concentration: 10%. Reference Table for Selecting SDS-PAGE Separation Gel Concentrations 2. Power up 80V for 30 minutes. Then the power supply is adjusted (110 V~150 V), the Marker is observed, and the electrophoresis can be stopped when the indicator band of the predyed protein Marker where the protein is located is properly separated. (Note that the current should not be too large when electrophoresis, too large current (more than 150 mA) will cause the temperature to rise, affecting the result of running glue. If high currents cannot be avoided, an ice bath can be used to cool the bath.)
Transfer membrane
1. Take out the converter, soak the clip and consumables in the pre-cooled converter;
2. Activate PVDF membrane with methanol for 1 min and rinse with transfer buffer;
3. Install it in the order of "black edge of clip - sponge - filter paper - filter paper - glue -PVDF membrane - filter paper - filter paper - sponge - white edge of clip"; 4. The protein was electrotransferred to PVDF membrane. ( 0.45 µm PVDF membrane is recommended ) Reference Table for Selecting PVDF Membrane Pore Size Specifications Recommended conditions for wet transfer: 200 mA, 60 min. ( Note that the transfer conditions can be adjusted according to the protein size. For high-molecular-weight proteins, a higher current and longer transfer time are recommended. However, ensure that the transfer tank remains at a low temperature to prevent gel melting.)
Block
1. After electrotransfer, wash the film with TBST at room temperature for 5 minutes;
2. Incubate the film in the blocking solution for 1 hour at room temperature;
3. Wash the film with TBST for 3 times, 5 minutes each time.
Antibody incubation
1. Use 5% skim milk powder to prepare the primary antibody working liquid (recommended dilution ratio for primary antibody 1:1000), gently shake and incubate with the film at 4°C overnight; 2. Wash the film with TBST 3 times, 5 minutes each time;
3. Add the secondary antibody to the blocking solution and incubate with the film gently at room temperature for 1 hour;
4. After incubation, wash the film with TBST 3 times for 5 minutes each time.
Antibody staining
1. Add the prepared ECL luminescent substrate (or select other color developing substrate according to the second antibody) and mix evenly;
2. Incubate with the film for 1 minute, remove excess substrate (keep the film moist), wrap with plastic film, and expose in the imaging system. |
Biological Description
| Specificity |
|---|
| Cre Recombinase Antibody [E18L5] detects transfected levels of total Cre recombinase protein. |
| Uniprot ID |
|---|
| P06956 |
| Clone |
|---|
| E18L5 |
| Synonym(s) |
|---|
| cre; cre recombinase; cyclization recombinase; Recombinase cre |
| Background |
|---|
| Cre recombinase is a bacteriophage P1–derived tyrosine type recombinase that catalyzes precise, site specific recombination between two 34 base pair loxP recognition elements, making it a cornerstone of modern genetic engineering and conditional gene control systems. The enzyme binds the symmetric inverted repeats flanking the asymmetric spacer region of each loxP site and induces sharp DNA bending, assembling a tetrameric synaptic complex in which two Cre dimers align two loxP sites to position the scissile phosphodiester bonds for coordinated cleavage and strand exchange. Within the active site, a conserved tyrosine residue acts as a nucleophile, forming a 3′ phosphotyrosine covalent intermediate with the DNA backbone, while the 5′ hydroxyl ends attack the corresponding phosphotyrosine complex on the partner strand, generating a Holliday junction like intermediate that is resolved to produce recombinant DNA with a defined outcome, deletion, inversion, or translocation, depending on the orientation and topology of the loxP sites. Cre loxP systems are used to generate tissue specific knockouts, inducible alleles, and lineage tracing models, with Cre expression driven by cell type specific promoters or tamoxifen regulated fusion constructs that link developmental, metabolic, and immune signaling pathways to targeted genetic perturbation in defined cell populations. Although naturally evolved to maintain the P1 phage–plasmid copy number, this tight, accessory factor independent mechanism has been adapted as a reliable, genome editable switch in mammalian cells, and its extensive structural and biophysical characterization underpins the rational design of improved variants and orthogonal recombinase systems. |
| References |
|---|
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.