research use only
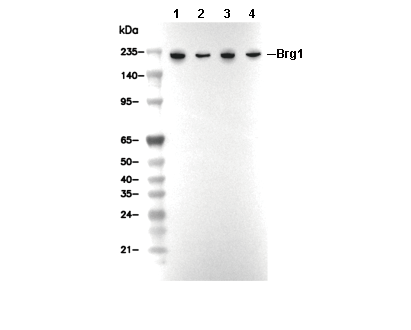
Brg1 Antibody [D24D16]
Cat.No.: F4669
Application:
Reactivity:
Experiment Essentials
WB
Recommended SDS-PAGE separating gel concentration: 5%.
Recommended wet transfer conditions: 250 mA, 180 min.
Recommended SDS-PAGE separating gel concentration: 5%.
Recommended wet transfer conditions: 250 mA, 180 min.
Usage Information
| Dilution |
|---|
|
| Application |
|---|
| WB, IP, ChIP |
| Reactivity |
|---|
| Human, Mouse, Rat, Monkey |
| Source |
|---|
| Rabbit Monoclonal Antibody |
| Storage Buffer |
|---|
| PBS, pH 7.2+50% Glycerol+0.05% BSA+0.01% NaN3 |
| Storage (from the date of receipt) |
|---|
| -20°C (avoid freeze-thaw cycles), 2 years |
| Predicted MW |
|---|
| 220 kDa |
| Positive Control | MCF7 cells; NCCIT cells; 3T3 cells; KNRK cells; OS-7 cells; 293T cells; RPMI 8226 cells |
|---|---|
| Negative Control | A549 cells |
Experimental Methods
| WB |
|---|
Experimental Protocol:
Sample preparation
1. Tissue: Lyse the tissue sample by adding an appropriate volume of ice-cold RIPA/Nuclear Lysis Buffer (containing Protease Inhibitor Cocktail),and homogenize the tissue at a low temperature. 2. Adherent cell: Aspirate the culture medium and wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/Nuclear Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 3. Suspension cell: Transfer the culture medium to a pre-cooled centrifuge tube. Centrifuge and aspirate the supernatant. Wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/Nuclear Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 4. Place the lysate into a pre-cooled microcentrifuge tube. Centrifuge at 4°C for 15 min. Collect the supernatant;
5. Remove a small volume of lysate to determine the protein concentration;
6. Combine the lysate with protein loading buffer. Boil 20 µL sample under 95-100°C for 5 min. Centrifuge for 5 min after cool down on ice.
Electrophoretic separation
1. According to the concentration of extracted protein, load appropriate amount of protein sample and marker onto SDS-PAGE gels for electrophoresis. Recommended separating gel (lower gel) concentration: 5%. Reference Table for Selecting SDS-PAGE Separation Gel Concentrations 2. Power up 80V for 30 minutes. Then the power supply is adjusted (110 V~150 V), the Marker is observed, and the electrophoresis can be stopped when the indicator band of the predyed protein Marker where the protein is located is properly separated. (Note that the current should not be too large when electrophoresis, too large current (more than 150 mA) will cause the temperature to rise, affecting the result of running glue. If high currents cannot be avoided, an ice bath can be used to cool the bath.)
Transfer membrane
1. Take out the converter, soak the clip and consumables in the pre-cooled converter;
2. Activate PVDF membrane with methanol for 1 min and rinse with transfer buffer;
3. Install it in the order of "black edge of clip - sponge - filter paper - filter paper - glue -PVDF membrane - filter paper - filter paper - sponge - white edge of clip"; 4. The protein was electrotransferred to PVDF membrane. ( 0.45 µm PVDF membrane is recommended ) Reference Table for Selecting PVDF Membrane Pore Size Specifications Recommended conditions for wet transfer: 250 mA, 180 min. ( Note that the transfer conditions can be adjusted according to the protein size. For high-molecular-weight proteins, a higher current and longer transfer time are recommended. However, ensure that the transfer tank remains at a low temperature to prevent gel melting.)
Block
1. After electrotransfer, wash the film with TBST at room temperature for 5 minutes;
2. Incubate the film in the blocking solution for 1 hour at room temperature;
3. Wash the film with TBST for 3 times, 5 minutes each time.
Antibody incubation
1. Use 5% skim milk powder to prepare the primary antibody working liquid (recommended dilution ratio for primary antibody 1:1000), gently shake and incubate with the film at 4°C overnight; 2. Wash the film with TBST 3 times, 5 minutes each time;
3. Add the secondary antibody to the blocking solution and incubate with the film gently at room temperature for 1 hour;
4. After incubation, wash the film with TBST 3 times for 5 minutes each time.
Antibody staining
1. Add the prepared ECL luminescent substrate (or select other color developing substrate according to the second antibody) and mix evenly;
2. Incubate with the film for 1 minute, remove excess substrate (keep the film moist), wrap with plastic film, and expose in the imaging system. |
Biological Description
| Specificity |
|---|
| Brg1 Antibody [D24D16] detects endogenous levels of total Brg1 protein. |
| Subcellular Location |
|---|
| Nucleus |
| Uniprot ID |
|---|
| P51532 |
| Clone |
|---|
| D24D16 |
| Synonym(s) |
|---|
| SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4; SMARCA4; BAF190A; Protein BRG-1; Protein brahma homolog 1; SNF2-beta; Transcription activator BRG1; SMARCA4; BAF190A; BRG1; SNF2B; SNF2L4 |
| Background |
|---|
| Brg1 is the central ATPase subunit of the mammalian SWI/SNF chromatin remodeling complexes, including BAF and PBAF, and hydrolyzes ATP to disrupt histone-DNA contacts, enabling the sliding and repositioning of nucleosomes to regulate approximately 10 to 15 percent of genes involved in development, differentiation, and proliferation. Brg1 contains N-terminal proline-rich regions that are unique compared to BRM, a bromodomain for acetyl-lysine recognition, helicase and SNF2 ATPase domains with seven motifs that generate around 10 piconewtons of force to translocate DNA along the histone octamer, and a BRK domain with a GYF-like fold for mediating protein interactions. It is recruited via an LXCXE motif to retinoblastoma protein or through glutamine-rich domains to nuclear receptors, p53, or BRCA1. The primary function of Brg1 is context-dependent transcriptional activation or repression: Brg1-BAF complexes containing BAF250a or BAF250b mobilize nucleosomes at p21 and p16INK4a promoters, cooperating with p53 and CBP acetyltransferase to destabilize p53 after DNA damage, with the PRR essential for ubiquitin-mediated degradation, while PBAF-Brg1 complexes with BAF200 repress ribosomal DNA via H2A.Z exchange. Brg1’s ATPase activity, which is about 100 times faster than the basal rate, couples to 50 base pair DNA translocation, exposing transcription factor binding sites. Brg1 is critical for heart morphogenesis, as knockout results in embryonic lethality and outflow tract defects through repression of Gata4 and Nkx2.5, regulates T-cell differentiation by balancing Th17 and Treg populations, and supports stem cell pluripotency and ES cell self-renewal via Oct4 and Sox2. BRG1 mutations, including truncations seen in about 20 percent of cancers, mimic the effects of SNF5 loss. Brg1 is inactivated in pancreatic and medullary thyroid cancers and Burkitt lymphoma due to RTK pathway activation, but can act as a paradoxical oncogene in non-small cell lung cancer and squamous cell carcinoma through cooperation with MYC. |
| References |
|---|
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.