research use only
ATRX Antibody [B2B21]
Cat.No.: F5060
Application:
Reactivity:
Experiment Essentials
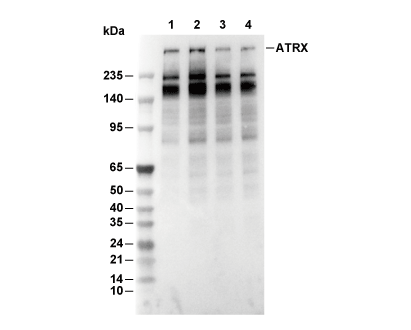
WB
Recommended SDS-PAGE separating gel concentration: 5%.
Recommended wet transfer conditions: 250 mA, 180 min.
Recommended SDS-PAGE separating gel concentration: 5%.
Recommended wet transfer conditions: 250 mA, 180 min.
Usage Information
| Dilution |
|---|
|
| Application |
|---|
| WB, IP, IHC |
| Reactivity |
|---|
| Human, Mouse, Rat, Monkey |
| Source |
|---|
| Rabbit Monoclonal Antibody |
| Storage Buffer |
|---|
| PBS, pH 7.2+50% Glycerol+0.05% BSA+0.01% NaN3 |
| Storage (from the date of receipt) |
|---|
| -20°C (avoid freeze-thaw cycles), 2 years |
| Predicted MW |
|---|
| 280 kDa |
| Positive Control | Mouse colon; Human colon carcinoma; Human esophageal carcinoma; Human parathyroid; Human prostate carcinoma; Human gastric carcinoma; Human uterus ; SH-SY5Y cells; LN-18 cells; SK-N-MC cells; MOLT-4 cells; NIH/3T3 cells; H-4-II-E cells; COS-7 cells |
|---|---|
| Negative Control | Saos-2 cells; U-2-02 cells |
Experimental Methods
| WB |
|---|
Experimental Protocol:
Sample preparation
1. Tissue: Lyse the tissue sample by adding an appropriate volume of ice-cold RIPA/NP-40 Lysis Buffer (containing Protease Inhibitor Cocktail),and homogenize the tissue at a low temperature. 2. Adherent cell: Aspirate the culture medium and wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/NP-40 Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 3. Suspension cell: Transfer the culture medium to a pre-cooled centrifuge tube. Centrifuge and aspirate the supernatant. Wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/NP-40 Lysis Buffer (containing Protease Inhibitor Cocktail) and put the sample on ice for 5 min. 4. Place the lysate into a pre-cooled microcentrifuge tube. Centrifuge at 4°C for 15 min. Collect the supernatant;
5. Remove a small volume of lysate to determine the protein concentration;
6. Combine the lysate with protein loading buffer. Boil 20 µL sample under 95-100°C for 5 min. Centrifuge for 5 min after cool down on ice.
Electrophoretic separation
1. According to the concentration of extracted protein, load appropriate amount of protein sample and marker onto SDS-PAGE gels for electrophoresis. Recommended separating gel (lower gel) concentration: 5%. Reference Table for Selecting SDS-PAGE Separation Gel Concentrations 2. Power up 80V for 30 minutes. Then the power supply is adjusted (110 V~150 V), the Marker is observed, and the electrophoresis can be stopped when the indicator band of the predyed protein Marker where the protein is located is properly separated. (Note that the current should not be too large when electrophoresis, too large current (more than 150 mA) will cause the temperature to rise, affecting the result of running glue. If high currents cannot be avoided, an ice bath can be used to cool the bath.)
Transfer membrane
1. Take out the converter, soak the clip and consumables in the pre-cooled converter;
2. Activate PVDF membrane with methanol for 1 min and rinse with transfer buffer;
3. Install it in the order of "black edge of clip - sponge - filter paper - filter paper - glue -PVDF membrane - filter paper - filter paper - sponge - white edge of clip"; 4. The protein was electrotransferred to PVDF membrane. ( 0.45 µm PVDF membrane is recommended ) Reference Table for Selecting PVDF Membrane Pore Size Specifications Recommended conditions for wet transfer: 250 mA, 180 min. ( Note that the transfer conditions can be adjusted according to the protein size. For high-molecular-weight proteins, a higher current and longer transfer time are recommended. However, ensure that the transfer tank remains at a low temperature to prevent gel melting.)
Block
1. After electrotransfer, wash the film with TBST at room temperature for 5 minutes;
2. Incubate the film in the blocking solution for 1 hour at room temperature;
3. Wash the film with TBST for 3 times, 5 minutes each time.
Antibody incubation
1. Use 5% skim milk powder to prepare the primary antibody working liquid (recommended dilution ratio for primary antibody 1:1000), gently shake and incubate with the film at 4°C overnight; 2. Wash the film with TBST 3 times, 5 minutes each time;
3. Add the secondary antibody to the blocking solution and incubate with the film gently at room temperature for 1 hour;
4. After incubation, wash the film with TBST 3 times for 5 minutes each time.
Antibody staining
1. Add the prepared ECL luminescent substrate (or select other color developing substrate according to the second antibody) and mix evenly;
2. Incubate with the film for 1 minute, remove excess substrate (keep the film moist), wrap with plastic film, and expose in the imaging system. |
| IHC |
|---|
Experimental Protocol:
Deparaffinization/Rehydration
1. Deparaffinize/hydrate sections:
2. Incubate sections in three washes of xylene for 5 min each.
3. Incubate sections in two washes of 100% ethanol for 10 min each.
4. Incubate sections in two washes of 95% ethanol for 10 min each.
5. Wash sections two times in dH2O for 5 min each.
6.Antigen retrieval: For Citrate: Heat slides in a microwave submersed in 1X citrate unmasking solution until boiling is initiated; continue with 10 min at a sub-boiling temperature (95°-98°C). Cool slides on bench top for 30 min.
Staining
1. Wash sections in dH2O three times for 5 min each.
2. Incubate sections in 3% hydrogen peroxide for 10 min.
3. Wash sections in dH2O two times for 5 min each.
4. Wash sections in wash buffer for 5 min.
5. Block each section with 100–400 µl of blocking solution for 1 hr at room temperature.
6. Remove blocking solution and add 100–400 µl primary antibody diluent in to each section. Incubate overnight at 4°C.
7. Remove antibody solution and wash sections with wash buffer three times for 5 min each.
8. Cover section with 1–3 drops HRPas needed. Incubate in a humidified chamber for 30 min at room temperature.
9. Wash sections three times with wash buffer for 5 min each.
10. Add DAB Chromogen Concentrate to DAB Diluent and mix well before use.
11. Apply 100–400 µl DAB to each section and monitor closely. 1–10 min generally provides an acceptable staining intensity.
12. Immerse slides in dH2O.
13. If desired, counterstain sections with hematoxylin.
14. Wash sections in dH2O two times for 5 min each.
15. Dehydrate sections: Incubate sections in 95% ethanol two times for 10 sec each; Repeat in 100% ethanol, incubating sections two times for 10 sec each; Repeat in xylene, incubating sections two times for 10 sec each.
16. Mount sections with coverslips and mounting medium.
|
Biological Description
| Specificity |
|---|
| ATRX Antibody [B2B21] detects endogenous levels of total ATRX protein. |
| Subcellular Location |
|---|
| Chromosome, Nucleus, Telomere |
| Uniprot ID |
|---|
| P46100 |
| Clone |
|---|
| B2B21 |
| Synonym(s) |
|---|
| Transcriptional regulator ATRX; ATP-dependent helicase ATRX; X-linked helicase II; X-linked nuclear protein (XNP); Znf-HX; ATRX; RAD54L; XH2 |
| Background |
|---|
| ATRX, or alpha-thalassemia/mental retardation syndrome X-linked, is a SWI/SNF family chromatin remodeler and ATPase/helicase that partners with DAXX to deposit the histone variant H3.3 at heterochromatic regions such as telomeres, pericentromeres, and ribosomal DNA repeats, as well as at euchromatic regulatory elements, thereby ensuring genomic stability, replication fidelity, and precise control of gene expression. ATRX features an N-terminal ADD domain that binds H3K9 trimethylation through a plant homeodomain finger, a central SNF2-like helicase core with seven motifs for ATP hydrolysis and DNA translocation, a C-terminal SWI/SNF ATPase domain, and zinc fingers including those that recognize G-quadruplex DNA; key residues enable specific deposition of H3.3 without nucleosome assembly activity. The primary function of ATRX involves binding G-quadruplex structures, putative quadruplex sequences, and CpG islands to maintain chromatin accessibility at active promoters and enhancers, increasing marks such as H3K4 trimethylation and H3K27 acetylation and promoting transcription factor occupancy by factors like RUNX3 and GATA1. ATRX deposits H3.3 to facilitate replication-coupled repair and elongation, indicated by H3K36 trimethylation, represses telomeric RNA, and coordinates sister chromatid cohesion and meiosis through stochastic single-cell effects at alpha-globin and zinc finger gene loci. ATRX supports S-phase progression through MRN complex recruitment and replication fork restart, enables homologous recombination repair subpathways, and contributes to developmental gene silencing, while its loss results in heterogeneous chromatin closure, replication stress, and alternative lengthening of telomeres in cancer cells. ATRX mutations cause ATR-X syndrome, characterized by intellectual disability and alpha-thalassemia due to globin dysregulation, and ATRX loss in cancers such as pancreatic neuroendocrine tumors and glioblastomas enables telomerase-independent survival. |
| References |
|---|
|
Tech Support
Tel: +1-832-582-8158 Ext:3
If you have any other enquiries, please leave a message.